

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.6
(162 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
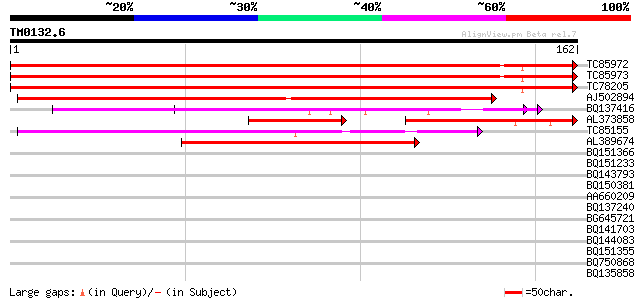
Sequences producing significant alignments: (bits) Value
TC85972 homologue to SP|O65743|RL24_CICAR 60S ribosomal protein ... 289 4e-79
TC85973 homologue to SP|O65743|RL24_CICAR 60S ribosomal protein ... 289 4e-79
TC78205 homologue to PIR|T47559|T47559 60S ribosomal protein-lik... 288 7e-79
AJ502894 weakly similar to SP|P38665|RL24 60S ribosomal protein ... 119 7e-28
BQ137416 similar to GP|10180025|gb| 60S ribosomal protein L24 {P... 109 4e-25
AL373858 SP|O65743|RL24 60S ribosomal protein L24. [Chickpea Ga... 57 4e-17
TC85155 similar to PIR|T00407|T00407 60S ribosomal protein L30 [... 80 4e-16
AL389674 similar to SP|P38665|RL24_ 60S ribosomal protein L24 (L... 55 2e-08
BQ151366 similar to GP|20146300|dbj hypothetical protein~similar... 32 0.085
BQ151233 weakly similar to PIR|T00407|T004 60S ribosomal protein... 32 0.085
BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 30 0.32
BQ150381 similar to GP|595768|gb|AA non-functional lacZ alpha pe... 30 0.32
AA660209 similar to GP|6642637|gb|A unknown protein {Arabidopsis... 30 0.55
BQ137240 similar to PIR|F75311|F753 ABC transporter ATP-binding... 29 0.94
BG645721 29 0.94
BQ141703 28 1.6
BQ144083 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 28 1.6
BQ151355 homologue to GP|12620658|gb| ID556 {Bradyrhizobium japo... 28 2.1
BQ750868 weakly similar to GP|4104436|gb|A acetolactate synthase... 28 2.1
BQ135858 similar to GP|23491723|db formin homology protein A {Di... 27 3.6
>TC85972 homologue to SP|O65743|RL24_CICAR 60S ribosomal protein L24.
[Chickpea Garbanzo] {Cicer arietinum}, partial (98%)
Length = 764
Score = 289 bits (739), Expect = 4e-79
Identities = 146/164 (89%), Positives = 156/164 (95%), Gaps = 2/164 (1%)
Frame = +1
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHN+LKPSKLTWTAM+RKQ
Sbjct: 73 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNKLKPSKLTWTAMYRKQ 252
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA +K+RR+TKKPYSRSIVGATLEVIQK+RTEKPEVRDAAREAALREIKER+K
Sbjct: 253 HKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDAAREAALREIKERIK 432
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGG--KVAAPKGPKLGGGGGKR 162
KTKDEK+AKKAEV +KAQKS GKG K A PKGPK+GGGGGKR
Sbjct: 433 KTKDEKKAKKAEVASKAQKS-GKGNVQKGAMPKGPKMGGGGGKR 561
>TC85973 homologue to SP|O65743|RL24_CICAR 60S ribosomal protein L24.
[Chickpea Garbanzo] {Cicer arietinum}, partial (98%)
Length = 725
Score = 289 bits (739), Expect = 4e-79
Identities = 146/164 (89%), Positives = 156/164 (95%), Gaps = 2/164 (1%)
Frame = +2
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHN+LKPSKLTWTAM+RKQ
Sbjct: 59 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNKLKPSKLTWTAMYRKQ 238
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA +K+RR+TKKPYSRSIVGATLEVIQK+RTEKPEVRDAAREAALREIKER+K
Sbjct: 239 HKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDAAREAALREIKERIK 418
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGG--KVAAPKGPKLGGGGGKR 162
KTKDEK+AKKAEV +KAQKS GKG K A PKGPK+GGGGGKR
Sbjct: 419 KTKDEKKAKKAEVASKAQKS-GKGNVQKGAMPKGPKMGGGGGKR 547
>TC78205 homologue to PIR|T47559|T47559 60S ribosomal protein-like -
Arabidopsis thaliana, partial (93%)
Length = 858
Score = 288 bits (737), Expect = 7e-79
Identities = 145/163 (88%), Positives = 153/163 (92%), Gaps = 1/163 (0%)
Frame = +1
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHN+LKPSKLTWTAM+RKQ
Sbjct: 55 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNKLKPSKLTWTAMYRKQ 234
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA +K+RR+TKKPYSRSIVGATLEVIQK+R+EKPEVRDAAREAALREIKERVK
Sbjct: 235 HKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRSEKPEVRDAAREAALREIKERVK 414
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGG-KVAAPKGPKLGGGGGKR 162
KTKDEK+AKKAE Q KAQKS GKG A KGPKLGGGGGKR
Sbjct: 415 KTKDEKKAKKAESQAKAQKSVGKGNVSKGASKGPKLGGGGGKR 543
>AJ502894 weakly similar to SP|P38665|RL24 60S ribosomal protein L24 (L30).
[Yeast] {Kluyveromyces lactis}, partial (84%)
Length = 501
Score = 119 bits (297), Expect = 7e-28
Identities = 59/137 (43%), Positives = 91/137 (66%)
Frame = +2
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
+K E F+G KIYPG+G ++RSD++ + FV SK + F R P K++WT ++R+ HK
Sbjct: 68 MKIESDSFTGNKIYPGKGKLYVRSDNRSYRFVGSKSESLFLQRKNPRKISWTTVYRRMHK 247
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K I +E A+K+ R K + R+IVGA+ E I+ +R +KPEVR+AAR A RE KE K
Sbjct: 248 KGITEEVAKKRTRRAVK-HQRAIVGASWEAIKAKRNQKPEVREAARAEAARESKENKKAE 424
Query: 123 KDEKRAKKAEVQTKAQK 139
+ +++A+KA++ A +
Sbjct: 425 QSKRKAEKAKLAQVASR 475
>BQ137416 similar to GP|10180025|gb| 60S ribosomal protein L24 {Prunus
avium}, partial (37%)
Length = 475
Score = 109 bits (273), Expect = 4e-25
Identities = 68/147 (46%), Positives = 83/147 (56%), Gaps = 7/147 (4%)
Frame = +1
Query: 13 AKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHKKDIAQEAARK 72
AKIYPGRGIRFIRSDSQVFL VNSKCKRYFHN+LKPSKLTWTAM+R+ ++ I + K
Sbjct: 7 AKIYPGRGIRFIRSDSQVFLXVNSKCKRYFHNKLKPSKLTWTAMYREATQEGICPRSCEK 186
Query: 73 KRRSTKKPYSRS-IVGATLEVIQKRRTEK----PEVRDAAREAALREIKER--VKKTKDE 125
+ S +K + ++ LEVI K K P + EIK K DE
Sbjct: 187 EA*SXQKALLQGPLLXLPLEVIXKEEIRKSQRVPGIAXPEEACPFXEIKGEG*KKNQXDE 366
Query: 126 KRAKKAEVQTKAQKSQGKGGKVAAPKG 152
K + + QK Q G K+ KG
Sbjct: 367 KEGE------ERQKGQX*GTKILLXKG 429
Score = 60.5 bits (145), Expect = 3e-10
Identities = 42/109 (38%), Positives = 55/109 (49%), Gaps = 8/109 (7%)
Frame = +2
Query: 48 PSKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSRSIVGATL-EVIQKRRTEK------ 100
P L KQHKK AQEA +K+RR+TKKPY + L ++ K+R+EK
Sbjct: 113 PQSLHGLPCIGKQHKKVFAQEAVKKRRRATKKPYFKVXCWXYLWKLFXKKRSEKARGFRG 292
Query: 101 -PEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVA 148
P + A LRE ER K +K K +V+ KAQKS G V+
Sbjct: 293 LPXQKKPALSXKLRERGERKTKXMRKKAKKGRKVKXKAQKSCWXKGNVS 439
>AL373858 SP|O65743|RL24 60S ribosomal protein L24. [Chickpea Garbanzo]
{Cicer arietinum}, partial (34%)
Length = 466
Score = 57.4 bits (137), Expect(2) = 4e-17
Identities = 33/53 (62%), Positives = 38/53 (71%), Gaps = 4/53 (7%)
Frame = +3
Query: 114 EIKERVKKTKDEKRAKKAEVQTKAQKSQGK---GGKVAAPKGP-KLGGGGGKR 162
EIKER+KKTKDEK+AKKAEV +KAQKS+ AP+GP GGGGKR
Sbjct: 162 EIKERIKKTKDEKKAKKAEVASKAQKSRETEMFTNGCYAPQGP*NWAGGGGKR 320
Score = 46.2 bits (108), Expect(2) = 4e-17
Identities = 21/28 (75%), Positives = 25/28 (89%)
Frame = +2
Query: 69 AARKKRRSTKKPYSRSIVGATLEVIQKR 96
A +K+RR+TKKPYSRSIVGATLEVI +
Sbjct: 2 AVKKRRRATKKPYSRSIVGATLEVISNK 85
>TC85155 similar to PIR|T00407|T00407 60S ribosomal protein L30 [imported] -
Arabidopsis thaliana, partial (62%)
Length = 1190
Score = 80.1 bits (196), Expect = 4e-16
Identities = 45/135 (33%), Positives = 73/135 (53%), Gaps = 2/135 (1%)
Frame = +1
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
++ E C F + +YPG GI+F+R+D+++F F SKC + F + P K+ WT +R+ H
Sbjct: 463 MRLEKCWFCSSTVYPGHGIQFVRNDAKIFRFCRSKCHKNFKMKRNPRKVKWTKAYRRVHG 642
Query: 63 KDIAQEAARKKRRSTKKP--YSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
KD+ Q++ + R +P Y R++ L+ I K K +V A A+R + K
Sbjct: 643 KDMTQDSTFEFERKRNRPERYDRNLTEEVLKAIPK--IAKIQVSRAESHHAIR---MKGK 807
Query: 121 KTKDEKRAKKAEVQT 135
K K +K A K Q+
Sbjct: 808 KKKVQKEAAKEYEQS 852
>AL389674 similar to SP|P38665|RL24_ 60S ribosomal protein L24 (L30). [Yeast]
{Kluyveromyces lactis}, partial (34%)
Length = 437
Score = 54.7 bits (130), Expect = 2e-08
Identities = 30/68 (44%), Positives = 41/68 (60%)
Frame = -2
Query: 50 KLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAARE 109
KL+WT ++R+ HKK I +E A + R T K S ++I+ +R PEVR+AAR
Sbjct: 421 KLSWTQVYRRMHKKGITEEVANIRTRRTVKLSKSSCRLHLGKLIRAKRIP*PEVREAARP 242
Query: 110 AALREIKE 117
ALRE KE
Sbjct: 241 TALREAKE 218
>BQ151366 similar to GP|20146300|dbj hypothetical protein~similar to Oryza
sativa OSJNBa0058E19.20, partial (9%)
Length = 1137
Score = 32.3 bits (72), Expect = 0.085
Identities = 31/101 (30%), Positives = 41/101 (39%), Gaps = 6/101 (5%)
Frame = +2
Query: 68 EAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTK--DE 125
E RKKRR+ P + G KRR EA +E +E+ KK + +
Sbjct: 152 ERKRKKRRTRPGPPRKEKSGEP-----KRRGGPGRGGAKKPEATNKEGEEKQKKDEPAQD 316
Query: 126 KRAKKAEVQTKAQKSQGKGGKVAAPKGP----KLGGGGGKR 162
+R K +E Q G + A KGP K GGG R
Sbjct: 317 RRGKASEQQPPPPAQPGAKRRTAGTKGPPRGSKKGGGNADR 439
>BQ151233 weakly similar to PIR|T00407|T004 60S ribosomal protein L30
[imported] - Arabidopsis thaliana, partial (37%)
Length = 775
Score = 32.3 bits (72), Expect = 0.085
Identities = 17/65 (26%), Positives = 33/65 (50%), Gaps = 2/65 (3%)
Frame = +2
Query: 33 FVNSKCKRYFHNRLKPSKLTWTAMFRKQHKKDIAQEAA--RKKRRSTKKPYSRSIVGATL 90
FV+ + F + P + WT + + H KD+ Q++ K++R+ + Y R++ L
Sbjct: 194 FVDPRATINFKMKRNPQ*VKWTNAYMRMHGKDLTQDSTFEFKRKRT*PERYDRNLTEEFL 373
Query: 91 EVIQK 95
+ I K
Sbjct: 374 KAIPK 388
Score = 29.6 bits (65), Expect = 0.55
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +1
Query: 16 YPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPS 49
YPG GI +D+ +F+F SK F N + S
Sbjct: 142 YPGHGIHVDPNDANIFVFCRSKSHNKFQNEEESS 243
>BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (19%)
Length = 1100
Score = 30.4 bits (67), Expect = 0.32
Identities = 17/59 (28%), Positives = 30/59 (50%)
Frame = -1
Query: 104 RDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGKR 162
R +A R+ + +K ++ KR K + + K ++ +G G + + GGGGGKR
Sbjct: 509 RGGGGQAKERKRGRKRRKGENAKREKGGKERGKKERKKGAGNGIYEGEYKGGGGGGGKR 333
Score = 27.3 bits (59), Expect = 2.7
Identities = 14/49 (28%), Positives = 27/49 (54%)
Frame = -2
Query: 114 EIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGKR 162
++ +R ++K E+ + + K + +GKGGK +G + G GG+R
Sbjct: 553 KVGKRKGESKWEQGQGEGGDKRKKGRGEGKGGKGKTQRGKRGGKKGGRR 407
>BQ150381 similar to GP|595768|gb|AA non-functional lacZ alpha peptide
{unidentified cloning vector}, partial (10%)
Length = 545
Score = 30.4 bits (67), Expect = 0.32
Identities = 19/61 (31%), Positives = 31/61 (50%)
Frame = -1
Query: 94 QKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGP 153
Q++R + E + R+ + E +KKTK++ K + Q + Q GKG K +GP
Sbjct: 218 QRKRKDNRETKAHKRKKTI----ETIKKTKNDAST*KTKRQKRMQHPDGKGEK----RGP 63
Query: 154 K 154
K
Sbjct: 62 K 60
>AA660209 similar to GP|6642637|gb|A unknown protein {Arabidopsis thaliana},
partial (4%)
Length = 544
Score = 29.6 bits (65), Expect = 0.55
Identities = 16/50 (32%), Positives = 28/50 (56%)
Frame = +1
Query: 94 QKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGK 143
+K+ + K +V++ E KE++KK KD+KR +K + +K GK
Sbjct: 1 EKKDSRKEKVKEKKEEG-----KEKIKKNKDDKRGEK-RXNKEEKKGHGK 132
>BQ137240 similar to PIR|F75311|F753 ABC transporter ATP-binding protein -
Deinococcus radiodurans (strain R1), partial (4%)
Length = 1094
Score = 28.9 bits (63), Expect = 0.94
Identities = 23/88 (26%), Positives = 41/88 (46%)
Frame = +3
Query: 75 RSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQ 134
R+ KK +++ E +K RT + R+A R+ K+R +K K K A+ +
Sbjct: 126 RNRKKNDTQATKNDKEEPRKKERTRPRKPREARGPERWRKRKKRSRK----KEKKDAKTR 293
Query: 135 TKAQKSQGKGGKVAAPKGPKLGGGGGKR 162
T+ ++ +GG+ K GG +R
Sbjct: 294 TRDRQKDREGGRGRR*KQTNRESGGERR 377
>BG645721
Length = 779
Score = 28.9 bits (63), Expect = 0.94
Identities = 19/62 (30%), Positives = 31/62 (49%)
Frame = -2
Query: 95 KRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPK 154
K +K + R + RE ++ KKTK+EK+ ++ Q K + Q G +PK P+
Sbjct: 481 KSNNKKQDRRTNKKNHKTRETIKQ-KKTKEEKKNRETIQQMKKKNHQTMLGFTESPKCPQ 305
Query: 155 LG 156
G
Sbjct: 304 KG 299
>BQ141703
Length = 1323
Score = 28.1 bits (61), Expect = 1.6
Identities = 25/90 (27%), Positives = 41/90 (44%), Gaps = 16/90 (17%)
Frame = +3
Query: 58 RKQHKKDIAQEAARKKRRSTKKPY-SRSIVGATLEVIQKRRTEKPEVRDAAR-------- 108
R H+K + AR+KRR T ++ A E ++ + E VRDA +
Sbjct: 921 RWSHEKQVP*ARAREKRRHTVSTLDEQTNESAR*ENLRIQHDESDRVRDATQRHDRDAPI 1100
Query: 109 -----EAALREIKERV--KKTKDEKRAKKA 131
+ +RE E+ K K+E+R+KK+
Sbjct: 1101 *EAVADETVRETHEKTCRDKRKEERRSKKS 1190
>BQ144083 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (7%)
Length = 467
Score = 28.1 bits (61), Expect = 1.6
Identities = 15/41 (36%), Positives = 24/41 (57%), Gaps = 2/41 (4%)
Frame = -3
Query: 124 DEKRAKKAEVQTKAQKSQ--GKGGKVAAPKGPKLGGGGGKR 162
+EK + AE +++ +K + GKGG+ K GGG GK+
Sbjct: 459 EEKNREMAEDRSEGRKEETRGKGGEGGGRKSGGEGGGRGKK 337
>BQ151355 homologue to GP|12620658|gb| ID556 {Bradyrhizobium japonicum},
partial (4%)
Length = 1420
Score = 27.7 bits (60), Expect = 2.1
Identities = 20/51 (39%), Positives = 26/51 (50%), Gaps = 3/51 (5%)
Frame = +2
Query: 114 EIKERVKKTKDEKRAKKAEVQTKAQKSQGKG---GKVAAPKGPKLGGGGGK 161
E + + K+ + EKR K E TK QK Q +G G K PK GG G+
Sbjct: 203 ESRGKKKQKRREKRQNKGEKGTK-QKPQNRGPGPGDHPQRKQPKNRGGEGR 352
Score = 26.2 bits (56), Expect = 6.1
Identities = 20/84 (23%), Positives = 37/84 (43%)
Frame = +2
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKE 117
+KQ +++ Q K + +KP +R G +R+ K + + +
Sbjct: 218 KKQKRREKRQNKGEKGTK--QKPQNR---GPGPGDHPQRKQPKNRGGEGRHSPTTKAENQ 382
Query: 118 RVKKTKDEKRAKKAEVQTKAQKSQ 141
+TKDEKR +K + + K +K Q
Sbjct: 383 EPPRTKDEKRKEKEDKERKEKKKQ 454
>BQ750868 weakly similar to GP|4104436|gb|A acetolactate synthase; oxo-acid
lyase {Leuconostoc lactis}, partial (7%)
Length = 494
Score = 27.7 bits (60), Expect = 2.1
Identities = 13/53 (24%), Positives = 26/53 (48%)
Frame = +1
Query: 49 SKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKP 101
++L W F + + I QEA + ++T + R + G + +V +K + P
Sbjct: 64 NELPWCRRFFIKVRGMICQEAPHSRAQTTALSHGRKVTGQSTQVCRKHPGDSP 222
>BQ135858 similar to GP|23491723|db formin homology protein A {Dictyostelium
discoideum}, partial (4%)
Length = 1033
Score = 26.9 bits (58), Expect = 3.6
Identities = 16/35 (45%), Positives = 20/35 (56%)
Frame = +2
Query: 126 KRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGG 160
K+ KK Q K + + KGG+V P LGGGGG
Sbjct: 182 KKKKKPPPQKKKKNLKKKGGEVPPPFF-FLGGGGG 283
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,923,331
Number of Sequences: 36976
Number of extensions: 47594
Number of successful extensions: 444
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 425
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 436
length of query: 162
length of database: 9,014,727
effective HSP length: 89
effective length of query: 73
effective length of database: 5,723,863
effective search space: 417841999
effective search space used: 417841999
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0132.6


