

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0131.12
(78 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
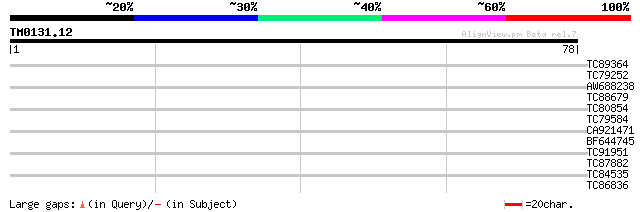
Score E
Sequences producing significant alignments: (bits) Value
TC89364 weakly similar to PIR|C96745|C96745 hypothetical protein... 29 0.38
TC79252 MtN21 28 0.85
AW688238 weakly similar to EGAD|60190|6299 putative disease-resi... 28 0.85
TC88679 similar to PIR|A86430|A86430 hypothetical protein AAF197... 26 3.2
TC80854 similar to GP|21595633|gb|AAM66119.1 unknown {Arabidopsi... 26 3.2
TC79584 similar to GP|20197980|gb|AAM15335.1 Expressed protein {... 26 3.2
CA921471 25 4.2
BF644745 similar to PIR|A85231|A85 hypothetical protein AT4g2035... 25 5.5
TC91951 similar to GP|5441893|dbj|BAA82391.1 ESTs C99174(E10437)... 25 7.2
TC87882 homologue to GP|4589852|dbj|BAA76902.1 cycloartenol synt... 25 7.2
TC84535 homologue to GP|11121216|emb|CAC14792. histone H3 {Morti... 24 9.4
TC86836 similar to GP|21553680|gb|AAM62773.1 unknown {Arabidopsi... 24 9.4
>TC89364 weakly similar to PIR|C96745|C96745 hypothetical protein T9N14.3
[imported] - Arabidopsis thaliana, partial (45%)
Length = 1903
Score = 28.9 bits (63), Expect = 0.38
Identities = 15/36 (41%), Positives = 18/36 (49%), Gaps = 3/36 (8%)
Frame = -2
Query: 8 RRLFASLLITNIMNNPDHVWNITWKL---LANDIQH 40
R L A I +NN H W +WKL L D+QH
Sbjct: 1707 RFLLARFRINKHLNNISHSWTQSWKLSSTLNTDLQH 1600
>TC79252 MtN21
Length = 1872
Score = 27.7 bits (60), Expect = 0.85
Identities = 19/61 (31%), Positives = 28/61 (45%), Gaps = 11/61 (18%)
Frame = +3
Query: 17 TNIMNNPDHVWNITWKLLANDIQHEYRGTLTQ------PGMI-----PTFSLSSKPILLK 65
T IM N D VW I W + N + Y G +T G++ P F+ S P+++
Sbjct: 942 TLIMENKDSVWTIGWDM--NLLAAAYAGIVTSSISYYIQGLVIKKKGPVFATSFSPLMMI 1115
Query: 66 I 66
I
Sbjct: 1116 I 1118
>AW688238 weakly similar to EGAD|60190|6299 putative disease-resistance clone
pH359-3 {Oryza sativa}, partial (18%)
Length = 690
Score = 27.7 bits (60), Expect = 0.85
Identities = 19/58 (32%), Positives = 24/58 (40%), Gaps = 1/58 (1%)
Frame = -3
Query: 2 HSGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLT-QPGMIPTFSLS 58
H LRR+F SL + N P H + + W YR T Q + TF LS
Sbjct: 343 HLMETLRRIFPSLFVNLQQNVPQHFFYVVWS--------RYRDPYT*QSKFLTTFWLS 194
>TC88679 similar to PIR|A86430|A86430 hypothetical protein AAF19754.1
[imported] - Arabidopsis thaliana, partial (47%)
Length = 809
Score = 25.8 bits (55), Expect = 3.2
Identities = 9/26 (34%), Positives = 18/26 (68%)
Frame = -1
Query: 7 LRRLFASLLITNIMNNPDHVWNITWK 32
L+ L AS+++T ++ P ++W+ WK
Sbjct: 284 LQILAASVILTAAVHQPHYLWDHVWK 207
>TC80854 similar to GP|21595633|gb|AAM66119.1 unknown {Arabidopsis
thaliana}, partial (95%)
Length = 563
Score = 25.8 bits (55), Expect = 3.2
Identities = 8/23 (34%), Positives = 17/23 (73%)
Frame = -1
Query: 49 PGMIPTFSLSSKPILLKIDNHLQ 71
P + P + LS +P+L ++++HL+
Sbjct: 230 PDLSPQYHLSEQPVLEEVEDHLE 162
>TC79584 similar to GP|20197980|gb|AAM15335.1 Expressed protein {Arabidopsis
thaliana}, partial (78%)
Length = 866
Score = 25.8 bits (55), Expect = 3.2
Identities = 14/49 (28%), Positives = 23/49 (46%)
Frame = +3
Query: 27 WNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKIDNHLQSNGK 75
W++TW L +I + + L ++P LS+K I H+ S K
Sbjct: 108 WSLTWNLTFPNIMNGCKTLLFFKPLLPNPFLSNKSIRCNSHGHVISTRK 254
>CA921471
Length = 381
Score = 25.4 bits (54), Expect = 4.2
Identities = 8/16 (50%), Positives = 11/16 (68%)
Frame = -3
Query: 18 NIMNNPDHVWNITWKL 33
N+ DH+WN +WKL
Sbjct: 307 NLTQKIDHLWNKSWKL 260
>BF644745 similar to PIR|A85231|A85 hypothetical protein AT4g20350 [imported]
- Arabidopsis thaliana, partial (52%)
Length = 684
Score = 25.0 bits (53), Expect = 5.5
Identities = 13/41 (31%), Positives = 20/41 (48%)
Frame = +1
Query: 13 SLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIP 53
+LL+ NI P WKLL N + G + + G++P
Sbjct: 238 TLLLNNIYGAPSS----KWKLLKNRRLQNWGGVVHEKGLLP 348
>TC91951 similar to GP|5441893|dbj|BAA82391.1 ESTs C99174(E10437)
D22295(C10709) correspond to a region of the predicted
gene.~Similar to, partial (23%)
Length = 526
Score = 24.6 bits (52), Expect = 7.2
Identities = 15/38 (39%), Positives = 20/38 (52%), Gaps = 7/38 (18%)
Frame = -2
Query: 9 RLFASLLITNIMNNPD-------HVWNITWKLLANDIQ 39
RL LLI N N+PD H+ + ++LLA IQ
Sbjct: 393 RLIIILLINNSSNSPD*QKKKKVHLTKVNFRLLAFAIQ 280
>TC87882 homologue to GP|4589852|dbj|BAA76902.1 cycloartenol synthase
{Glycyrrhiza glabra}, partial (51%)
Length = 1408
Score = 24.6 bits (52), Expect = 7.2
Identities = 10/38 (26%), Positives = 20/38 (52%)
Frame = +1
Query: 25 HVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
H+W++ W A+ E++ L P + F++ + PI
Sbjct: 745 HLWHLVWGKRADCCWEEFQ*LLKHP*SL*LFAVEAAPI 858
>TC84535 homologue to GP|11121216|emb|CAC14792. histone H3 {Mortierella
alpina}, partial (98%)
Length = 654
Score = 24.3 bits (51), Expect = 9.4
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = -3
Query: 11 FASLLITNIMNNPDHVWNITWKLL 34
F+ LLI + + D W+IT +LL
Sbjct: 217 FSGLLIASNFSKSDGTWSITMRLL 146
>TC86836 similar to GP|21553680|gb|AAM62773.1 unknown {Arabidopsis
thaliana}, partial (61%)
Length = 1387
Score = 24.3 bits (51), Expect = 9.4
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = -2
Query: 45 TLTQPGMIPTFSLSSKPILLKIDNHLQSNGKSLK 78
TL Q M P S+S P+L D+ + +SLK
Sbjct: 477 TLKQEQMEPMISVSKFPLLSV*DSRTSQHNRSLK 376
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,959,635
Number of Sequences: 36976
Number of extensions: 40290
Number of successful extensions: 206
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 206
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 206
length of query: 78
length of database: 9,014,727
effective HSP length: 54
effective length of query: 24
effective length of database: 7,018,023
effective search space: 168432552
effective search space used: 168432552
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0131.12


