

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
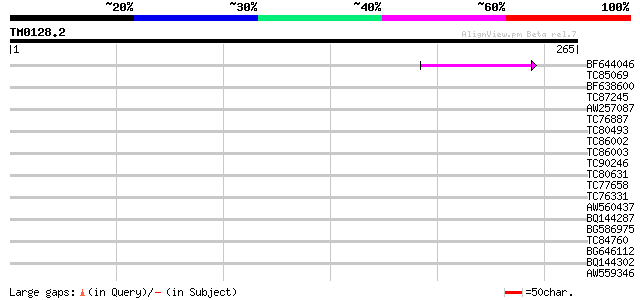
Query= TM0128.2
(265 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF644046 52 2e-07
TC85069 similar to GP|1354468|gb|AAD12774.1| U1 snRNP 70K trunca... 31 0.56
BF638600 similar to SP|O14405|GUN4_ Endoglucanase IV precursor (... 30 0.96
TC87245 similar to GP|10334991|gb|AAG15839.2 NADPH-dependent man... 30 0.96
AW257087 homologue to GP|6683126|dbj KIAA0339 protein {Homo sapi... 30 1.3
TC76887 similar to GP|12802327|gb|AAK07827.1 mitochondrial proce... 29 1.6
TC80493 similar to GP|21618131|gb|AAM67181.1 unknown {Arabidopsi... 29 1.6
TC86002 weakly similar to PIR|S10205|S10205 pathogenesis-related... 29 2.1
TC86003 weakly similar to GP|7407641|gb|AAF62171.1| pathogenesis... 29 2.1
TC90246 similar to PIR|T00872|T00872 probable protein kinase At2... 29 2.1
TC80631 similar to GP|21555659|gb|AAM63908.1 unknown {Arabidopsi... 29 2.1
TC77658 similar to PIR|T46895|T46895 acyl-CoA dehydrogenase (EC ... 28 2.8
TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding pro... 28 2.8
AW560437 similar to GP|9827551|emb|C hypothetical protein LM26.2... 28 3.6
BQ144287 similar to PIR|G86292|G86 hypothetical protein AAF82153... 28 3.6
BG586975 28 3.6
TC84760 similar to GP|8567777|gb|AAF76349.1| unknown protein {Ar... 28 4.8
BG646112 weakly similar to GP|10177368|dbj contains similarity t... 28 4.8
BQ144302 similar to GP|6815109|dbj| hypothetical protein {Oryza ... 25 5.3
AW559346 similar to GP|12802327|gb mitochondrial processing pept... 27 8.1
>BF644046
Length = 597
Score = 52.0 bits (123), Expect = 2e-07
Identities = 23/54 (42%), Positives = 32/54 (58%)
Frame = +3
Query: 193 YLHGYIFNELFMKLPFSTFVCDVLTFLNVAPCQLLPNAWGFTRCFELLCEPLNF 246
+++ ++F ++ K PF+ F CD L LNVA QL PN F FE+ CE L F
Sbjct: 9 HMYSFVFEDIGFKFPFTNFECDFLKALNVASSQLHPNCCAFMCGFEISCESLGF 170
>TC85069 similar to GP|1354468|gb|AAD12774.1| U1 snRNP 70K truncated protein
{Arabidopsis thaliana}, partial (79%)
Length = 675
Score = 30.8 bits (68), Expect = 0.56
Identities = 15/40 (37%), Positives = 19/40 (47%)
Frame = +1
Query: 88 HNSPSSSDPTQKRTVSITSHLSTGNPSTSEDWRTSDKCID 127
H SS+ P ++R+ S S PST DW S ID
Sbjct: 106 HGRSSSAQPKRRRSSSYQSSEPRQRPSTQTDWTESSNWID 225
>BF638600 similar to SP|O14405|GUN4_ Endoglucanase IV precursor (EC
3.2.1.4) (Endo-1 4-beta-glucanase IV) (Cellulase IV)
(EGIV)., partial (10%)
Length = 690
Score = 30.0 bits (66), Expect = 0.96
Identities = 20/58 (34%), Positives = 30/58 (51%), Gaps = 8/58 (13%)
Frame = +2
Query: 4 IPVTFQCSDHHCS--------LHSYYFTHPRLVPLFSLYFSCFSIHLLSMKLPKMSAK 53
IP TF CS H S +HS+ T R++PLF+ + + FS K+P ++K
Sbjct: 17 IPFTF-CSLHILSRFFDRSRFIHSFTSTASRILPLFTNHKTNFSYTTTFFKMPSFTSK 187
>TC87245 similar to GP|10334991|gb|AAG15839.2 NADPH-dependent mannose
6-phosphate reductase {Orobanche ramosa}, partial (98%)
Length = 1196
Score = 30.0 bits (66), Expect = 0.96
Identities = 9/34 (26%), Positives = 21/34 (61%)
Frame = +3
Query: 98 QKRTVSITSHLSTGNPSTSEDWRTSDKCIDDKCL 131
QK + +T+H G + +++W ++ C+D++ L
Sbjct: 621 QKHGICVTAHTPLGGAAANKEWFGTESCLDEQIL 722
>AW257087 homologue to GP|6683126|dbj KIAA0339 protein {Homo sapiens},
partial (2%)
Length = 466
Score = 29.6 bits (65), Expect = 1.3
Identities = 21/74 (28%), Positives = 34/74 (45%), Gaps = 18/74 (24%)
Frame = +1
Query: 63 NSDSDDGDKTTVSQEVITV------------------DSDSESHNSPSSSDPTQKRTVSI 104
+SDSD ++ S V +V DSDS+S +S SSS + + S
Sbjct: 169 DSDSDSSSSSSTSSSVSSVKKPAVSAAKATPAKKEESDSDSDSDSSSSSSSSSSSSSSSS 348
Query: 105 TSHLSTGNPSTSED 118
+S S+ + S++ D
Sbjct: 349 SSSSSSSSSSSNSD 390
>TC76887 similar to GP|12802327|gb|AAK07827.1 mitochondrial processing
peptidase beta subunit {Cucumis melo}, partial (92%)
Length = 1974
Score = 29.3 bits (64), Expect = 1.6
Identities = 26/77 (33%), Positives = 36/77 (45%)
Frame = -1
Query: 15 CSLHSYYFTHPRLVPLFSLYFSCFSIHLLSMKLPKMSAK*QSYYSLSLNSDSDDGDKTTV 74
CSL Y F P P+FS+ S LLS+ L + +K + LS S+ D T
Sbjct: 657 CSLEVYAFKCP---PIFSISSSKSLAFLLSVPLKIICSKKCAVPLLSSVSNLDPASIQTP 487
Query: 75 SQEVITVDSDSESHNSP 91
+ VI + DS + SP
Sbjct: 486 TVAVIALRLDSVATRSP 436
>TC80493 similar to GP|21618131|gb|AAM67181.1 unknown {Arabidopsis
thaliana}, partial (18%)
Length = 847
Score = 29.3 bits (64), Expect = 1.6
Identities = 15/44 (34%), Positives = 21/44 (47%), Gaps = 5/44 (11%)
Frame = +3
Query: 5 PVTFQCS-----DHHCSLHSYYFTHPRLVPLFSLYFSCFSIHLL 43
P +F+CS D HC+ S + TH + L S F+ H L
Sbjct: 351 PFSFKCSSCTSCDQHCTGGSCFMTHTHELQLVHFIISWFACHFL 482
>TC86002 weakly similar to PIR|S10205|S10205 pathogenesis-related protein 1
- common tobacco, partial (72%)
Length = 830
Score = 28.9 bits (63), Expect = 2.1
Identities = 15/42 (35%), Positives = 21/42 (49%), Gaps = 2/42 (4%)
Frame = -2
Query: 11 SDHHCSLHSYYFTHPRLVPLFSLYFSCFS--IHLLSMKLPKM 50
S HHCS+ S+ P VP SL C +H + + PK+
Sbjct: 307 SCHHCSMLSFLHMDPWSVPFCSLLLICLHRFLHRMQLLHPKI 182
>TC86003 weakly similar to GP|7407641|gb|AAF62171.1| pathogenesis-related
protein 1 {Betula pendula}, partial (95%)
Length = 356
Score = 28.9 bits (63), Expect = 2.1
Identities = 15/42 (35%), Positives = 21/42 (49%), Gaps = 2/42 (4%)
Frame = -2
Query: 11 SDHHCSLHSYYFTHPRLVPLFSLYFSCFS--IHLLSMKLPKM 50
S HHCS+ S+ P VP SL C +H + + PK+
Sbjct: 235 SCHHCSMLSFLHMDPWSVPFCSLLLICLHRFLHRMQLLHPKI 110
>TC90246 similar to PIR|T00872|T00872 probable protein kinase At2g45590
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1451
Score = 28.9 bits (63), Expect = 2.1
Identities = 30/105 (28%), Positives = 41/105 (38%), Gaps = 3/105 (2%)
Frame = -1
Query: 13 HHCSLHSYYFTHPRLVPLFSLYFSCFSIHLLSMKLPKMSAK*QSYYSLSLNSDSDDGDKT 72
HHC H ++F H LFS +S FS P + ++ SLS +S
Sbjct: 485 HHC*YHHHHFLH-----LFSFSYSSFSPLQPLHNAPPSTIPSPTFSSLSSSSHQQ----- 336
Query: 73 TVSQEVITVDSDSESHNSPSSSDPTQKRTVSITSHLS---TGNPS 114
+ I S S +SP SS + +SH S NPS
Sbjct: 335 -FPRPSIPTIPISASSSSPFSSHSQSYHPLIHSSHFSLPFQSNPS 204
>TC80631 similar to GP|21555659|gb|AAM63908.1 unknown {Arabidopsis
thaliana}, partial (60%)
Length = 1296
Score = 28.9 bits (63), Expect = 2.1
Identities = 19/64 (29%), Positives = 31/64 (47%), Gaps = 10/64 (15%)
Frame = +2
Query: 208 FSTFVCDVLTFLN------VAPCQLLPNAWGFTRCFELLCEPLN----FGPPYPLFLFFY 257
FS ++C V +N + PC+ + WG C + CE + FG + +F+ FY
Sbjct: 926 FSQWLCHVHNVVNRSISKPIFPCERVDARWGKLDCEQNACEIIGSISIFGKIW*IFI-FY 1102
Query: 258 KISI 261
KI +
Sbjct: 1103KIIV 1114
>TC77658 similar to PIR|T46895|T46895 acyl-CoA dehydrogenase (EC 1.3.99.3)
[validated] - Arabidopsis thaliana, partial (94%)
Length = 2886
Score = 28.5 bits (62), Expect = 2.8
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = +2
Query: 8 FQCSDHHCSLHSYYFTHPRLVPLFSLYFS 36
F S HHC LH+ +F + P SL+F+
Sbjct: 35 FSFSSHHCFLHNQHFHTSQHKPTQSLHFT 121
>TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding protein
{Nicotiana tabacum}, partial (97%)
Length = 2581
Score = 28.5 bits (62), Expect = 2.8
Identities = 25/81 (30%), Positives = 39/81 (47%), Gaps = 16/81 (19%)
Frame = +1
Query: 201 ELFMKLPFSTFVCDVLTFLN--VAPCQ--------------LLPNAWGFTRCFELLCEPL 244
++FM L F++ V D L+F + ++P Q LLP G + C L + L
Sbjct: 2308 KIFMLLGFTSLVVDYLSFGHWFLSPFQIELFH*TRYCRFLDLLPWVEGMSDC*YLFID-L 2484
Query: 245 NFGPPYPLFLFFYKISIAKPS 265
N P F+FFY++ + K S
Sbjct: 2485 NLEP*NCPFVFFYRLGVEKIS 2547
>AW560437 similar to GP|9827551|emb|C hypothetical protein LM26.22
{Leishmania major}, partial (37%)
Length = 575
Score = 28.1 bits (61), Expect = 3.6
Identities = 17/61 (27%), Positives = 28/61 (45%), Gaps = 2/61 (3%)
Frame = +3
Query: 65 DSDDGDKTTVSQEVITVDSDSESHNSPSSSDPTQK--RTVSITSHLSTGNPSTSEDWRTS 122
+ G TT + E V S S + P + ++ + VS + S G+PS + +WR
Sbjct: 360 EPSSGTLTTSANEEDFVPSTSIAFQCPQGENLIEEINKVVSFAQNKSIGSPSRNYNWRLE 539
Query: 123 D 123
D
Sbjct: 540 D 542
>BQ144287 similar to PIR|G86292|G86 hypothetical protein AAF82153.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 1176
Score = 28.1 bits (61), Expect = 3.6
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = -3
Query: 244 LNFGPPYPLFLFFY 257
LN GPP P+FLFF+
Sbjct: 103 LNVGPPRPIFLFFF 62
>BG586975
Length = 589
Score = 28.1 bits (61), Expect = 3.6
Identities = 12/35 (34%), Positives = 21/35 (59%)
Frame = -2
Query: 197 YIFNELFMKLPFSTFVCDVLTFLNVAPCQLLPNAW 231
+IF + FM+LPFS ++C V +++ C + W
Sbjct: 219 FIFFDAFMELPFSFYMCSV----DLSECNFAIDFW 127
>TC84760 similar to GP|8567777|gb|AAF76349.1| unknown protein {Arabidopsis
thaliana}, partial (36%)
Length = 781
Score = 27.7 bits (60), Expect = 4.8
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = +3
Query: 20 YYFTHPRLVPLFSLYFSCFSIHLL 43
+YF HP L+P +L+ S F H L
Sbjct: 114 FYFLHPHLLPQSTLHHSQFQYHRL 185
>BG646112 weakly similar to GP|10177368|dbj contains similarity to heat shock
transcription factor HSF30~gene_id:K16L22.13, partial
(5%)
Length = 742
Score = 27.7 bits (60), Expect = 4.8
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = +2
Query: 161 HNDDLSLNIAPCDDNEVVTLKPS 183
H +DLSLNI C +++V L PS
Sbjct: 341 HLEDLSLNITNCHGDDMVYLPPS 409
>BQ144302 similar to GP|6815109|dbj| hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 1047
Score = 25.0 bits (53), Expect(2) = 5.3
Identities = 9/22 (40%), Positives = 13/22 (58%)
Frame = +3
Query: 243 PLNFGPPYPLFLFFYKISIAKP 264
P F PP+P FF+ +S+ P
Sbjct: 528 PFFFLPPFPPLFFFFPVSLPFP 593
Score = 20.8 bits (42), Expect(2) = 5.3
Identities = 8/18 (44%), Positives = 10/18 (55%)
Frame = +1
Query: 207 PFSTFVCDVLTFLNVAPC 224
PF FV ++ F V PC
Sbjct: 373 PFFLFVSPLIFFHTVCPC 426
>AW559346 similar to GP|12802327|gb mitochondrial processing peptidase beta
subunit {Cucumis melo}, partial (22%)
Length = 432
Score = 26.9 bits (58), Expect = 8.1
Identities = 25/77 (32%), Positives = 34/77 (43%)
Frame = -2
Query: 15 CSLHSYYFTHPRLVPLFSLYFSCFSIHLLSMKLPKMSAK*QSYYSLSLNSDSDDGDKTTV 74
CSL Y F P P+FS+ S LLS+ L + +K + LS + D T
Sbjct: 416 CSLEVYAFKCP---PIFSISSSKSLAFLLSVPLKIICSKKCAVPLLSSVXNLDPASIQTP 246
Query: 75 SQEVITVDSDSESHNSP 91
VI + DS + SP
Sbjct: 245 PVAVIALRLDSVATRSP 195
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,969,714
Number of Sequences: 36976
Number of extensions: 187003
Number of successful extensions: 1469
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 1448
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1464
length of query: 265
length of database: 9,014,727
effective HSP length: 94
effective length of query: 171
effective length of database: 5,538,983
effective search space: 947166093
effective search space used: 947166093
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0128.2


