

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0126.7
(542 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
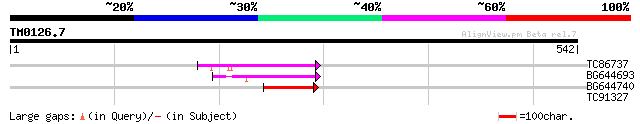
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 58 1e-08
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 58 1e-08
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 57 2e-08
TC91327 similar to GP|22597168|gb|AAN03471.1 unknown protein {Gl... 28 6.9
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 57.8 bits (138), Expect = 1e-08
Identities = 38/126 (30%), Positives = 71/126 (56%), Gaps = 8/126 (6%)
Frame = +1
Query: 180 GLMGDYSELFST--IWDLPPSR-VQDHAIHL---QEG--AAIPNIRPYRCPHYQKSEIEK 231
G++ ++ +LF+ + +P SR + DHAI L ++G +P Y + ++K
Sbjct: 340 GVLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLYGMSRQELLVLKK 519
Query: 232 LVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLY 291
++++LD G + S S + V+ V+K G RFC+DYRA + IT +++P+ +I E L
Sbjct: 520 TLEDLLDKGFIKASGSAAGAPVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLR 699
Query: 292 KLGEAK 297
++ A+
Sbjct: 700 RVAGAR 717
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 57.8 bits (138), Expect = 1e-08
Identities = 38/108 (35%), Positives = 58/108 (53%), Gaps = 5/108 (4%)
Frame = +2
Query: 195 LPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQ----KSEIEKL-VKEMLDVGIFRPSISQF 249
+PP D I L +PN+ P P Y+ K ++ KL +K++L+ G +PSI
Sbjct: 17 VPPEWKIDFGIDL-----LPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP* 181
Query: 250 SSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
+V+ +KKKDG R IDY + + I K+P+ +IDEL L +K
Sbjct: 182 GVVVLFLKKKDGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSK 325
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 57.0 bits (136), Expect = 2e-08
Identities = 26/53 (49%), Positives = 37/53 (69%)
Frame = -1
Query: 243 RPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGE 295
+PSIS + ++ V+KKDG +R CIDYR F+K+T NK+P+ ID L K+ E
Sbjct: 262 QPSISP*GAALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQE 104
>TC91327 similar to GP|22597168|gb|AAN03471.1 unknown protein {Glycine max},
partial (70%)
Length = 738
Score = 28.5 bits (62), Expect = 6.9
Identities = 15/25 (60%), Positives = 15/25 (60%)
Frame = -1
Query: 88 FFLFNLGGLDVVLGL*WFASLGHIR 112
F LF L GLD LGL WF LG R
Sbjct: 357 FGLFRLFGLDGFLGLFWFFRLGRWR 283
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.349 0.158 0.543
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,614,158
Number of Sequences: 36976
Number of extensions: 248754
Number of successful extensions: 1754
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1282
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 461
Number of HSP's gapped (non-prelim): 1349
length of query: 542
length of database: 9,014,727
effective HSP length: 101
effective length of query: 441
effective length of database: 5,280,151
effective search space: 2328546591
effective search space used: 2328546591
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0126.7


