

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
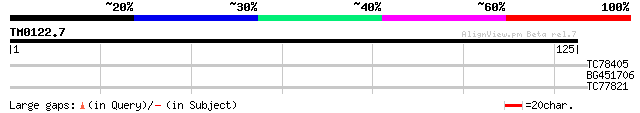
Query= TM0122.7
(125 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78405 peroxidase precursor 25 6.0
BG451706 weakly similar to GP|15722495|emb macrofage activating ... 25 7.9
TC77821 similar to GP|21618197|gb|AAM67247.1 putative aminotrans... 25 7.9
>TC78405 peroxidase precursor
Length = 988
Score = 25.4 bits (54), Expect = 6.0
Identities = 11/25 (44%), Positives = 15/25 (60%)
Frame = -2
Query: 91 SRSFSPLSLDVVTKAEKLSLSDGRF 115
S +F P L + EKL++ DGRF
Sbjct: 582 SFTFKPCDLKLEISCEKLNVGDGRF 508
>BG451706 weakly similar to GP|15722495|emb macrofage activating
glycoprotein {Filobasidiella neoformans}, partial (20%)
Length = 644
Score = 25.0 bits (53), Expect = 7.9
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = -3
Query: 53 MNCKEELYLLQIDAKKLGD 71
+NC+E+ LL ID K+GD
Sbjct: 99 LNCREKSQLLTIDRIKVGD 43
>TC77821 similar to GP|21618197|gb|AAM67247.1 putative aminotransferase
{Arabidopsis thaliana}, partial (93%)
Length = 1631
Score = 25.0 bits (53), Expect = 7.9
Identities = 11/28 (39%), Positives = 18/28 (64%), Gaps = 1/28 (3%)
Frame = -1
Query: 1 MSGIFVYRISTAKEWEEL-QANGSTFGG 27
++G F+Y++ T K W+ L + N S F G
Sbjct: 419 LNGCFLYKVWTIKIWKTLTKINSSMFDG 336
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,936,767
Number of Sequences: 36976
Number of extensions: 47480
Number of successful extensions: 226
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 226
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 226
length of query: 125
length of database: 9,014,727
effective HSP length: 84
effective length of query: 41
effective length of database: 5,908,743
effective search space: 242258463
effective search space used: 242258463
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0122.7


