

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.6
(378 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
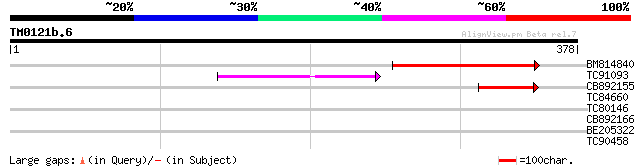
Sequences producing significant alignments: (bits) Value
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 121 4e-28
TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein... 75 4e-14
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 64 7e-11
TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG282... 37 0.013
TC80146 36 0.028
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 33 0.14
BE205322 similar to PIR|F86218|F86 protein F22O13.8 [imported] -... 28 7.6
TC90458 similar to GP|19343726|gb|AAH25494.1 Similar to N-acylam... 27 9.9
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 121 bits (304), Expect = 4e-28
Identities = 60/98 (61%), Positives = 77/98 (78%)
Frame = +3
Query: 256 NGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAM 315
+GTR+I+ +L K +I A V+ GE ++IPRM+L PS A++ F+R QFP++L FAM
Sbjct: 27 HGTRLIIVSLGKNVICARVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPLVLSFAM 206
Query: 316 TINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSRN 353
TINKSQGQ+ T VGLYLPRPVFTH QLYVA+SRVKSR+
Sbjct: 207 TINKSQGQTLTSVGLYLPRPVFTHGQLYVAVSRVKSRS 320
>TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein T24M8.10 -
Arabidopsis thaliana, partial (11%)
Length = 701
Score = 75.1 bits (183), Expect = 4e-14
Identities = 42/109 (38%), Positives = 63/109 (57%)
Frame = +3
Query: 139 PCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEIEYLN 198
P DP+ IV +YP L+ F Q RAIL T E V++IN+++L +I GEE +
Sbjct: 381 PPGDPIDAIVQSTYPNLVSQYNNEQFLQSRAILTSTDEVVDQINDYVLKLIPGEERVIYS 560
Query: 199 CDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRN 247
+ +S+ + + + EFL K S +PNH++ LKVG PIML+R+
Sbjct: 561 AN---RSEVNDVQAFDAIPPEFLQSLKTSDLPNHKLTLKVGTPIMLLRD 698
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 64.3 bits (155), Expect = 7e-11
Identities = 31/40 (77%), Positives = 34/40 (84%)
Frame = -2
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
FAMTINKSQGQS H+G+YLP VF+H QLYVALSRV SR
Sbjct: 358 FAMTINKSQGQSLKHIGVYLPSSVFSHGQLYVALSRVTSR 239
>TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 1009
Score = 37.0 bits (84), Expect = 0.013
Identities = 17/24 (70%), Positives = 20/24 (82%)
Frame = +2
Query: 329 GLYLPRPVFTHVQLYVALSRVKSR 352
G+YLP+P+F H LYVALSRV SR
Sbjct: 83 GMYLPQPIF*HG*LYVALSRVTSR 154
>TC80146
Length = 476
Score = 35.8 bits (81), Expect = 0.028
Identities = 14/28 (50%), Positives = 23/28 (82%)
Frame = -3
Query: 316 TINKSQGQSSTHVGLYLPRPVFTHVQLY 343
TINKS+ QS +++ +YL RP+F+H ++Y
Sbjct: 450 TINKSR*QSLSYMKIYLSRPIFSHEEMY 367
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 33.5 bits (75), Expect = 0.14
Identities = 15/23 (65%), Positives = 18/23 (78%)
Frame = -1
Query: 331 YLPRPVFTHVQLYVALSRVKSRN 353
+ R VF+H QLYVA+SRV SRN
Sbjct: 289 FYDREVFSHGQLYVAISRVSSRN 221
>BE205322 similar to PIR|F86218|F86 protein F22O13.8 [imported] - Arabidopsis
thaliana, partial (5%)
Length = 456
Score = 27.7 bits (60), Expect = 7.6
Identities = 20/72 (27%), Positives = 34/72 (46%), Gaps = 4/72 (5%)
Frame = +3
Query: 62 KEAVRRLW----VLLSILHIYGSIAR*WN*LLI*DCKMLTHILHQQK*KSLLIGCFKWET 117
+E +R W VLL +L + R L++ +L ++ ++K*K+L + C KW
Sbjct: 102 REYLRMKW*ILRVLLVVLQMTTHTMRDLRLLILMTNCILRNLYRRRK*KTLFLSC*KWRV 281
Query: 118 EQLQPLMKLNHS 129
+P L S
Sbjct: 282 RLRRPKKLLKKS 317
>TC90458 similar to GP|19343726|gb|AAH25494.1 Similar to
N-acylaminoacyl-peptide hydrolase {Mus musculus},
partial (6%)
Length = 987
Score = 27.3 bits (59), Expect = 9.9
Identities = 18/66 (27%), Positives = 30/66 (45%), Gaps = 11/66 (16%)
Frame = -2
Query: 126 LNHSFRFLLIFI-------EPC----KDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPT 174
L H LL+F+ E C DPLL+++ SYP ++ S QE + P+
Sbjct: 749 LGHLGETLLVFLPSRGVR*ETCYKIESDPLLKVLQLSYPFRWYHSSPQSTNQEYQLYQPS 570
Query: 175 LESVEE 180
+ ++
Sbjct: 569 AQGYKQ 552
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.343 0.151 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,344,960
Number of Sequences: 36976
Number of extensions: 181219
Number of successful extensions: 1583
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1566
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1582
length of query: 378
length of database: 9,014,727
effective HSP length: 98
effective length of query: 280
effective length of database: 5,391,079
effective search space: 1509502120
effective search space used: 1509502120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0121b.6


