

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.2
(253 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
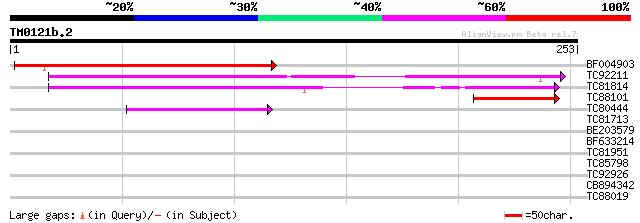
BF004903 similar to PIR|B96789|B96 protein T23E18.4 [imported] -... 182 1e-46
TC92211 similar to PIR|B86182|B86182 hypothetical protein [impor... 159 7e-40
TC81814 weakly similar to GP|9294541|dbj|BAB02804.1 high mobilit... 140 4e-34
TC88101 similar to PIR|B86182|B86182 hypothetical protein [impor... 41 4e-04
TC80444 similar to GP|15450515|gb|AAK96550.1 At2g17400 {Arabidop... 41 4e-04
TC81713 29 2.0
BE203579 similar to PIR|F96585|F96 hypothetical protein F20D21.2... 28 2.6
BF633214 similar to GP|14495224|db contains EST C72594(E1889)~si... 28 4.4
TC81951 28 4.4
TC85798 similar to PIR|T51979|T51979 multicatalytic endopeptidas... 27 7.5
TC92926 similar to GP|11139266|gb|AAG31651.1 PRLI-interacting fa... 27 9.9
CB894342 similar to GP|20197988|gb expressed protein {Arabidopsi... 27 9.9
TC88019 similar to GP|21539559|gb|AAM53332.1 unknown protein {Ar... 27 9.9
>BF004903 similar to PIR|B96789|B96 protein T23E18.4 [imported] - Arabidopsis
thaliana, partial (30%)
Length = 709
Score = 182 bits (462), Expect = 1e-46
Identities = 89/118 (75%), Positives = 101/118 (85%), Gaps = 1/118 (0%)
Frame = +2
Query: 3 SAARTIRSGGDD-GKFYPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLD 61
SA R+ SG DD K YPPPLASH +VND LF DTLR+FHF M +K+MIPVIGG++LD
Sbjct: 356 SAVRSKNSGNDDEAKHYPPPLASHNDVVNDPTLFWDTLRRFHFLMATKFMIPVIGGKELD 535
Query: 62 LHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVH 119
LH LYVEVTRRSGYEKVVAEKKWREVGSVF FS+TTTSAS+VL+KHY NLLY++EQVH
Sbjct: 536 LHVLYVEVTRRSGYEKVVAEKKWREVGSVFRFSSTTTSASFVLRKHYLNLLYHYEQVH 709
>TC92211 similar to PIR|B86182|B86182 hypothetical protein [imported] -
Arabidopsis thaliana, partial (31%)
Length = 798
Score = 159 bits (403), Expect = 7e-40
Identities = 89/232 (38%), Positives = 135/232 (57%), Gaps = 1/232 (0%)
Frame = +3
Query: 18 YPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEK 77
YPPP+A++E +V++ KLF+ L + H MG+K+MIPVIGGR+LDLH L+VEVT R G+EK
Sbjct: 126 YPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIGGRELDLHRLFVEVTSRGGFEK 305
Query: 78 VVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNS 137
++ ++KW+EV VFNF +T T+AS+VL+K+Y +LLY++EQ+++FK + T VL S
Sbjct: 306 IIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYEQIYYFKAR-DWTNTTSDVLQS 482
Query: 138 FTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNV 197
+ +Q +P+ P + + N+ P +
Sbjct: 483 QSSIPAPAPKMQFSHPS----------------------PQVQPAVFQQLKVNSAPPEAM 596
Query: 198 EAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFH-PEETVPPPSSRVV 248
+ G I+ KFD GYLV+V +GSE L+GV++ P+ V P S V
Sbjct: 597 GSSSAGSQVVGVIDGKFDSGYLVTVTIGSEKLKGVLYQAPQNPVLPASHHSV 752
>TC81814 weakly similar to GP|9294541|dbj|BAB02804.1 high mobility group
protein-like {Arabidopsis thaliana}, partial (33%)
Length = 867
Score = 140 bits (354), Expect = 4e-34
Identities = 87/231 (37%), Positives = 119/231 (50%), Gaps = 3/231 (1%)
Frame = +1
Query: 18 YPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEK 77
YPPP A + +V D+ LF L+ FH +G+K IP IGG+ LDLH L+VEVT R G EK
Sbjct: 181 YPPPTAPYSDLVRDSNLFQQKLQSFHDSLGTKLKIPTIGGKPLDLHHLFVEVTSRGGIEK 360
Query: 78 VVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPP---AGTV 134
V+ ++KW+EV FNF T TS S++++K Y +LLY+FEQ ++F Q P + P +G V
Sbjct: 361 VIVDRKWKEVIMSFNFRDTITSGSFMVRKTYLSLLYHFEQAYYFCKQVPPSTPDALSGNV 540
Query: 135 LNSFTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPE 194
NSFT D A + +P + P
Sbjct: 541 ANSFTT-----------------------------------NTDGAAINDSPVQVS--PI 609
Query: 195 SNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSS 245
S + G G I+ KFD GY+V+V LGSE L+GV++H SS
Sbjct: 610 SPAQTLG--SSVRGTIDMKFDDGYIVTVDLGSEQLKGVLYHVSSNASKGSS 756
>TC88101 similar to PIR|B86182|B86182 hypothetical protein [imported] -
Arabidopsis thaliana, partial (7%)
Length = 1362
Score = 41.2 bits (95), Expect = 4e-04
Identities = 19/38 (50%), Positives = 25/38 (65%)
Frame = +3
Query: 208 GRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSS 245
G I+ KFD GY+V+V LGSE L+GV++H SS
Sbjct: 960 GTIDGKFDDGYIVTVDLGSEQLKGVLYHVSSNASKDSS 1073
>TC80444 similar to GP|15450515|gb|AAK96550.1 At2g17400 {Arabidopsis
thaliana}, partial (31%)
Length = 1118
Score = 41.2 bits (95), Expect = 4e-04
Identities = 23/65 (35%), Positives = 32/65 (48%)
Frame = +1
Query: 53 PVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLL 112
P G L+ L+ V R GYEKV + K WR VG F T T+ S+ + Y L
Sbjct: 43 PKFYGEWLNCLKLWRAVMRLGGYEKVTSCKLWRSVGESFKPPKTCTTVSWTFRGFYEKAL 222
Query: 113 YNFEQ 117
++E+
Sbjct: 223 LDYER 237
>TC81713
Length = 736
Score = 28.9 bits (63), Expect = 2.0
Identities = 13/27 (48%), Positives = 19/27 (70%)
Frame = +1
Query: 105 KKHYWNLLYNFEQVHFFKVQGPITPPA 131
KKHY + +Y+++ FKVQGPI P+
Sbjct: 151 KKHYLSFIYHYQLR--FKVQGPIYTPS 225
>BE203579 similar to PIR|F96585|F96 hypothetical protein F20D21.21 [imported]
- Arabidopsis thaliana, partial (20%)
Length = 596
Score = 28.5 bits (62), Expect = 2.6
Identities = 27/96 (28%), Positives = 40/96 (41%)
Frame = +2
Query: 85 REVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCFME 144
RE+ ++ + + SA LK + +L +F + F Q T GTV FT+ E
Sbjct: 62 RELENLDKDADSRKSAMRALKSYVKDL--DFRAIPMFLAQVSETKETGTVSGEFTISLYE 235
Query: 145 MDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLA 180
+ L P SIC LS S +S L+
Sbjct: 236 V--LDPTMPEDKKKHIIHSICKPLSDSLTSSLDSLS 337
>BF633214 similar to GP|14495224|db contains EST C72594(E1889)~similar to
Arabidopsis thaliana chromosome 1 F25I16.5~unknown
protein, partial (16%)
Length = 502
Score = 27.7 bits (60), Expect = 4.4
Identities = 14/55 (25%), Positives = 25/55 (45%), Gaps = 15/55 (27%)
Frame = +2
Query: 39 LRQFHFHMGSKYMIPVIGGRKL---------------DLHTLYVEVTRRSGYEKV 78
++ +H H G ++P++GG +L DLH L + R S + +V
Sbjct: 38 VKGYHLHQGKVTLLPLVGGERLVIFGGRGEGDANYLNDLHILDLRTMRWSSFPEV 202
>TC81951
Length = 653
Score = 27.7 bits (60), Expect = 4.4
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = -1
Query: 91 FNFSTTTTSASYVLKKHYWNLLYNFEQVHFFK 122
F+FS++ TS KK W F +H FK
Sbjct: 587 FSFSSSATSGGAXQKKLRWPSAKKFSHIHLFK 492
>TC85798 similar to PIR|T51979|T51979 multicatalytic endopeptidase complex
(EC 3.4.99.46) chain PBB2 [imported] - Arabidopsis
thaliana, partial (94%)
Length = 1161
Score = 26.9 bits (58), Expect = 7.5
Identities = 12/36 (33%), Positives = 17/36 (46%)
Frame = -2
Query: 53 PVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVG 88
P +GGR DL L + + + K+ E W E G
Sbjct: 146 PPLGGRSTDLVILEIRINPENE*RKMEIELSWNEAG 39
>TC92926 similar to GP|11139266|gb|AAG31651.1 PRLI-interacting factor K
{Arabidopsis thaliana}, partial (35%)
Length = 1032
Score = 26.6 bits (57), Expect = 9.9
Identities = 9/43 (20%), Positives = 22/43 (50%)
Frame = +3
Query: 73 SGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNF 115
+G K+ + + W+ + F + + Y+ K+ +WN+ + F
Sbjct: 393 AGVVKLSSRQNWKSI*KFFTSHFSALAE*YLRKRKWWNIKHQF 521
>CB894342 similar to GP|20197988|gb expressed protein {Arabidopsis thaliana},
partial (64%)
Length = 838
Score = 26.6 bits (57), Expect = 9.9
Identities = 15/39 (38%), Positives = 21/39 (53%), Gaps = 4/39 (10%)
Frame = +3
Query: 148 LQGMYPNIHAVTEFE----SICDVLSGSSSSGKPDLALV 182
+QG P IH T + + DVLSGSS+ DL ++
Sbjct: 198 VQGYSPTIHPTTSTDLSSIATLDVLSGSSARQHTDLTML 314
>TC88019 similar to GP|21539559|gb|AAM53332.1 unknown protein {Arabidopsis
thaliana}, partial (66%)
Length = 1706
Score = 26.6 bits (57), Expect = 9.9
Identities = 11/36 (30%), Positives = 22/36 (60%)
Frame = -1
Query: 67 VEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASY 102
+E+++RSG+E + E++WR V + T +A +
Sbjct: 293 LEISQRSGHESLQEERRWRSNCCVLSAKIQT*TARW 186
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.137 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,683,659
Number of Sequences: 36976
Number of extensions: 122900
Number of successful extensions: 652
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 647
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 651
length of query: 253
length of database: 9,014,727
effective HSP length: 94
effective length of query: 159
effective length of database: 5,538,983
effective search space: 880698297
effective search space used: 880698297
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0121b.2


