

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
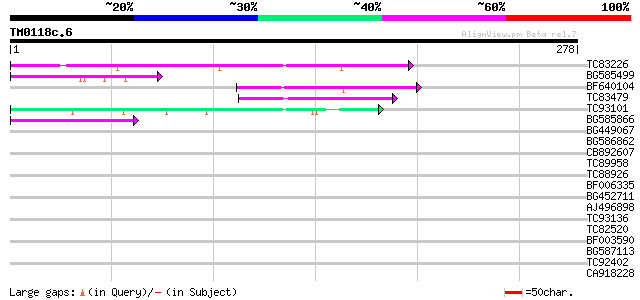
Query= TM0118c.6
(278 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 63 1e-10
BG585499 55 3e-08
BF640104 45 4e-05
TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protei... 43 1e-04
TC93101 42 2e-04
BG585866 41 4e-04
BG449067 39 0.002
BG586862 38 0.004
CB892607 similar to GP|8927657|gb| EST gb|N38213 comes from this... 37 0.006
TC89958 similar to GP|12483623|dbj|BAB21445. NADH dehydrogenase ... 36 0.014
TC88926 similar to GP|7290507|gb|AAF45960.1| peb gene product {D... 35 0.024
BF006335 similar to GP|14334538|gb| unknown protein {Arabidopsis... 31 0.46
BG452711 31 0.60
AJ496898 similar to GP|23171295|gb CG3731-PA {Drosophila melanog... 31 0.60
TC93136 31 0.60
TC82520 30 1.0
BF003590 29 1.7
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 29 1.7
TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like ser... 28 5.1
CA918228 weakly similar to GP|18478606|gb| repetitive proline ri... 28 5.1
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 63.2 bits (152), Expect = 1e-10
Identities = 54/216 (25%), Positives = 94/216 (43%), Gaps = 18/216 (8%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRSQARS------RNS 54
CPRC + E I H CP S+ +W G+ NF + ++I + +
Sbjct: 243 CPRCHSKTETITHLFMSCPLSKRVW--FGSNLCINFDNLPNPNFIN*LYEAIL*KDECIT 416
Query: 55 VRFLAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGV-----------AG 103
+ A ++ +W RN V ED L + +R + ++ +
Sbjct: 417 I*IAAIIYNLWHARNLSVLEDQTILEMDIIQRASNCISDYKQANTQAPPSMARTGYDPRS 596
Query: 104 DHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGN-A 162
H +++W+ P G+VK+N D + + G G +IRD+ G+ + G + A
Sbjct: 597 QHRPAKNTKWKRPNLGLVKVNTDANLQNHGK-WGLGIIIRDEVGLVMAASTWETDGNDRA 773
Query: 163 LIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
L AEA ALL G+ D G+ +V E D +L++++
Sbjct: 774 LEAEAYALLTGMRFAKDCGFXKVXFEGDNEKLMKMV 881
>BG585499
Length = 792
Score = 55.1 bits (131), Expect = 3e-08
Identities = 28/85 (32%), Positives = 43/85 (49%), Gaps = 10/85 (11%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAW-TN-FGVGDHGSWI-RSQARSRNSV-- 55
CP C +E +LH L DC + ++W+RL W TN F D W+ ++ ++ N V
Sbjct: 348 CPCCGNADETVLHVLCDCRPASQVWIRLVPSDWITNFFSFDDCRDWVFKNLSKRSNGVSK 527
Query: 56 -----RFLAGVWGVWKWRNNMVFED 75
F+ W +W WRN +FE+
Sbjct: 528 FKWQPTFMTTCWHMWTWRNKAIFEE 602
>BF640104
Length = 344
Score = 44.7 bits (104), Expect = 4e-05
Identities = 28/92 (30%), Positives = 47/92 (50%), Gaps = 1/92 (1%)
Frame = -1
Query: 112 RWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNA-LIAEATAL 170
+W P QGV+K+N D + +D G G + R+D GI + +R G + AEA +
Sbjct: 341 KWIKPHQGVIKINCDANLTSED-VWGIGVITRNDNGIVMASGTWNRPGFMCPITAEAWGV 165
Query: 171 LLGLELVWDLGYRQVMVEVDCGELLQVMADEE 202
D G++ V+ E D +L+ +++ EE
Sbjct: 164 YQAALFALDQGFQNVLFENDNEKLISMLSREE 69
>TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 1222
Score = 43.1 bits (100), Expect = 1e-04
Identities = 27/78 (34%), Positives = 40/78 (50%)
Frame = +2
Query: 113 WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLL 172
W+ P+ G KLN D S ++ G GGL+RD +G + F + G+ + E A+
Sbjct: 977 WKKPEIGWTKLNTDGSVNKETA--GFGGLLRDYRGEPICAFVSKAPQGDTFLVELWAIWR 1150
Query: 173 GLELVWDLGYRQVMVEVD 190
GL L LG + + VE D
Sbjct: 1151 GLVLSLGLGIKSIWVESD 1204
>TC93101
Length = 675
Score = 42.4 bits (98), Expect = 2e-04
Identities = 48/211 (22%), Positives = 77/211 (35%), Gaps = 28/211 (13%)
Frame = -1
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLG-ALAWTNFGVGDHGSWIRSQARSRNS---VR 56
CPRC E H C +Q +W ++ + + + W+ ++ ++
Sbjct: 612 CPRCYFNLETTNHIFMSCERTQRVWFGSQLSIRFPDNSTINFSDWLFDAISNQTEEIIIK 433
Query: 57 FLAGVWGVWKWRNNMVFED----SPWLVDEACRRICHEHDEFI-------------TFSN 99
A + +W RN +FE+ ++ A I I SN
Sbjct: 432 ISAITYSIWHARNKAIFENQFVSEDTIIQ*AQNSILAYEQATIKPQNPNIVLSSLSATSN 253
Query: 100 GVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQG------IW-LGG 152
S + SRWQ P ++K N D + Q R G G +IR+ G W + G
Sbjct: 252 TNTTRRRSNVRSRWQKPLNNILKANCDANLQVQGR-WGLGCIIRNADGEAKVTATWCING 76
Query: 153 FYAHRSGGNALIAEATALLLGLELVWDLGYR 183
F A AE A+L + L D G++
Sbjct: 75 FDC------AATAETYAILAAMYLAKDYGFK 1
>BG585866
Length = 828
Score = 41.2 bits (95), Expect = 4e-04
Identities = 19/63 (30%), Positives = 26/63 (41%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRSQARSRNSVRFLAG 60
C RC EE HC+RDC S+ IW ++G + F W++ FL
Sbjct: 576 CSRCGEHEESFFHCVRDCRFSKIIWHKIGFSSPDFFSSSSVQDWLKDGISCHRPTTFLED 755
Query: 61 VWG 63
G
Sbjct: 756 CGG 764
>BG449067
Length = 578
Score = 38.9 bits (89), Expect = 0.002
Identities = 26/99 (26%), Positives = 47/99 (47%), Gaps = 1/99 (1%)
Frame = -2
Query: 112 RWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSG-GNALIAEATAL 170
+W+ P++ ++KLN D + D G G + R+D+G + R G +A AEA +
Sbjct: 310 KWKKPEKDIIKLNSDANLSSTD-LWGIGVVARNDEGFAMASGTWFRFGFPSATTAEAWGI 134
Query: 171 LLGLELVWDLGYRQVMVEVDCGELLQVMADEESCRFLCL 209
+ + G+ +V E D ++Q++ E L L
Sbjct: 133 YQAMIFAGEYGFSKVQFESDNERVIQMLNGTEEVNRLYL 17
>BG586862
Length = 804
Score = 38.1 bits (87), Expect = 0.004
Identities = 49/203 (24%), Positives = 75/203 (36%), Gaps = 7/203 (3%)
Frame = -1
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNF---GVGDHGSWIRSQARSRNS--- 54
CPRC + E + H +C +Q+ W G+ NF GV WI + +
Sbjct: 540 CPRCESKIETVQHLFLNCEVTQKEWF--GSQLGINFHSSGVLHFHDWITNFILKNDEETI 367
Query: 55 VRFLAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGVAGDHGSWLSSRWQ 114
+ A ++ +W RN VFE+ D +R F +A S L S
Sbjct: 366 IALTALLYSIWHARNQKVFENIDVPGDVVIQRASSSLHSF-----KMAQVSDSVLPSNAI 202
Query: 115 PPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWL-GGFYAHRSGGNALIAEATALLLG 173
P S W G G + R+ +G+ + G + A AEA +
Sbjct: 201 P----------SYSLW------GIGVVARNCEGLAMASGTWLRHGIPCATTAEAWGIYQA 70
Query: 174 LELVWDLGYRQVMVEVDCGELLQ 196
+ D G+ + E D G LL+
Sbjct: 69 MVFAGDCGFSKFEFESDNGNLLR 1
>CB892607 similar to GP|8927657|gb| EST gb|N38213 comes from this gene.
{Arabidopsis thaliana}, partial (1%)
Length = 782
Score = 37.4 bits (85), Expect = 0.006
Identities = 26/74 (35%), Positives = 37/74 (49%), Gaps = 1/74 (1%)
Frame = +1
Query: 115 PPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGG-NALIAEATALLLG 173
PP V L D D GGLI+DDQG ++ YA++ G + L AE + G
Sbjct: 559 PPNSCEVALKCDYVVLNCDLNAACGGLIQDDQGHFV-FHYANKLGSCSVLQAELWGI*HG 735
Query: 174 LELVWDLGYRQVMV 187
L + W+ GY ++ V
Sbjct: 736 LSIDWNRGYSKIRV 777
>TC89958 similar to GP|12483623|dbj|BAB21445. NADH dehydrogenase subunit 4L
{Gonostoma gracile}, partial (17%)
Length = 1040
Score = 36.2 bits (82), Expect = 0.014
Identities = 14/33 (42%), Positives = 22/33 (66%)
Frame = +2
Query: 113 WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDD 145
W+PP G VK NMD + +++ C+GAG + D+
Sbjct: 551 WEPPS*GYVKCNMDAALFKEYNCVGAGFRV*DE 649
>TC88926 similar to GP|7290507|gb|AAF45960.1| peb gene product {Drosophila
melanogaster}, partial (1%)
Length = 1073
Score = 35.4 bits (80), Expect = 0.024
Identities = 26/74 (35%), Positives = 38/74 (51%), Gaps = 4/74 (5%)
Frame = +3
Query: 121 VKLNMDESFWEQDRC--MGAGGLIRDDQGIWLGGFYAHRSGGNALI--AEATALLLGLEL 176
V LN+D S + G GG++ D G WL GF A + N + E A+L GL
Sbjct: 342 VILNVDGSLLREREVPSAGCGGVLSDSSGKWLCGF-AQKLNPNLKVDETEKEAILRGLLW 518
Query: 177 VWDLGYRQVMVEVD 190
V + G R+++V+ D
Sbjct: 519 VKEKGKRKILVKSD 560
>BF006335 similar to GP|14334538|gb| unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 505
Score = 31.2 bits (69), Expect = 0.46
Identities = 18/42 (42%), Positives = 22/42 (51%)
Frame = +2
Query: 137 GAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLLGLELVW 178
G GGL R+ + GFY N L AE LL+GL+L W
Sbjct: 209 GFGGLDRNYDKAFQFGFYGSIGWSNILHAEIQDLLVGLKLWW 334
>BG452711
Length = 672
Score = 30.8 bits (68), Expect = 0.60
Identities = 9/25 (36%), Positives = 16/25 (64%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIW 25
CP C+ +D+LH L C ++++W
Sbjct: 408 CPICNQRSQDMLHALFSCTRAKDVW 482
>AJ496898 similar to GP|23171295|gb CG3731-PA {Drosophila melanogaster},
partial (35%)
Length = 698
Score = 30.8 bits (68), Expect = 0.60
Identities = 18/69 (26%), Positives = 28/69 (40%), Gaps = 22/69 (31%)
Frame = +2
Query: 42 GSWIRSQARSRNSVRFLA----------------------GVWGVWKWRNNMVFEDSPWL 79
G+W RSQ N+ LA G+WGV+ + M E+ W
Sbjct: 92 GAWDRSQGGGANNASGLAAIVAEDNCAHSFQSFNTCYKDTGLWGVYFVSDGMTIENMVWN 271
Query: 80 VDEACRRIC 88
+ EA +++C
Sbjct: 272 IQEAWKKMC 298
>TC93136
Length = 722
Score = 30.8 bits (68), Expect = 0.60
Identities = 22/79 (27%), Positives = 35/79 (43%), Gaps = 4/79 (5%)
Frame = +1
Query: 13 HCLRDCPHSQEIWLR-LGALAWTNFGVGDHGSWIRSQARSRNSVRFL---AGVWGVWKWR 68
H CP +W + LG L ++ F + + RSR +L A +W +WK
Sbjct: 274 HLFLHCPFLSSVWSKILGWLYYSKFVC----VFRLLERRSRTKGVWLIWHATIWVIWKGI 441
Query: 69 NNMVFEDSPWLVDEACRRI 87
NN +F++ +DE I
Sbjct: 442 NNRIFKNISKAIDEIVEEI 498
>TC82520
Length = 833
Score = 30.0 bits (66), Expect = 1.0
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = +2
Query: 55 VRFLAGVWGVWKWRNNMVFEDS 76
+ + A VW +WK RNN VF++S
Sbjct: 497 IMWFASVWVLWKKRNNRVFQNS 562
>BF003590
Length = 531
Score = 29.3 bits (64), Expect = 1.7
Identities = 21/78 (26%), Positives = 33/78 (41%), Gaps = 8/78 (10%)
Frame = +1
Query: 57 FLAGVWGVWKWRNNMVFED--------SPWLVDEACRRICHEHDEFITFSNGVAGDHGSW 108
F W +W+ RN MVF++ S +++ C H T + V D
Sbjct: 313 FCITFWKIWQVRNQMVFQNQQLNPIQISISILEFT*ELNCSNHSAIATVT--VRSD---- 474
Query: 109 LSSRWQPPKQGVVKLNMD 126
+ W PP GV+ LN++
Sbjct: 475 -AEFWCPPPPGVMNLNLN 525
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 29.3 bits (64), Expect = 1.7
Identities = 10/25 (40%), Positives = 14/25 (56%)
Frame = -3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIW 25
C RC E + H L CP+++ IW
Sbjct: 174 CSRCGMESETVNHILFQCPYARLIW 100
>TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (16%)
Length = 704
Score = 27.7 bits (60), Expect = 5.1
Identities = 20/65 (30%), Positives = 27/65 (40%), Gaps = 2/65 (3%)
Frame = +1
Query: 90 EHDEFITFSNGVAGDHGSWLSSRWQPPKQG--VVKLNMDESFWEQDRCMGAGGLIRDDQG 147
EH T S G HG+ ++S WQ + G + N+D W + R G I
Sbjct: 64 EHKLHTTRSWEFLGLHGNDINSAWQKGRFGENTIIANIDTGVWPESRSFSDRG-IGPIPA 240
Query: 148 IWLGG 152
W GG
Sbjct: 241 KWRGG 255
>CA918228 weakly similar to GP|18478606|gb| repetitive proline rich protein
{Oryza sativa}, partial (13%)
Length = 303
Score = 27.7 bits (60), Expect = 5.1
Identities = 15/37 (40%), Positives = 17/37 (45%), Gaps = 2/37 (5%)
Frame = +1
Query: 42 GSWIRSQARSR--NSVRFLAGVWGVWKWRNNMVFEDS 76
GSW+RS R N RF W VW WR + S
Sbjct: 130 GSWLRSWLWMRCWNRFRF---AWWVWIWRRRWALQSS 231
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.333 0.143 0.509
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,062,738
Number of Sequences: 36976
Number of extensions: 201197
Number of successful extensions: 1132
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 1118
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1128
length of query: 278
length of database: 9,014,727
effective HSP length: 95
effective length of query: 183
effective length of database: 5,502,007
effective search space: 1006867281
effective search space used: 1006867281
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0118c.6


