

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
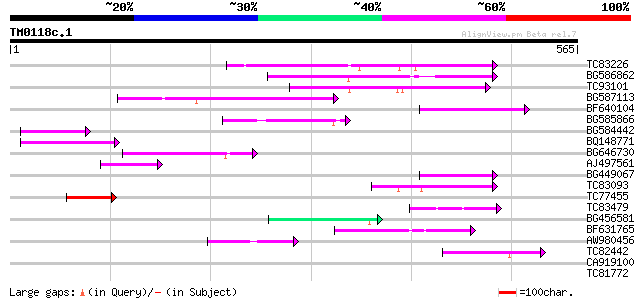
Query= TM0118c.1
(565 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 129 4e-30
BG586862 105 6e-23
TC93101 97 2e-20
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 76 4e-14
BF640104 67 1e-11
BG585866 67 1e-11
BG584442 56 3e-08
BQ148771 55 7e-08
BG646730 52 5e-07
AJ497561 weakly similar to GP|10140689|gb putative non-LTR retro... 48 9e-06
BG449067 48 9e-06
TC83093 48 1e-05
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 47 2e-05
TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protei... 44 2e-04
BG456581 43 3e-04
BF631765 42 5e-04
AW980456 42 6e-04
TC82442 42 6e-04
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 40 0.002
TC81772 40 0.003
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 129 bits (323), Expect = 4e-30
Identities = 85/287 (29%), Positives = 138/287 (47%), Gaps = 17/287 (5%)
Frame = +3
Query: 217 WPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCCETAWRAC 276
W G YS ++GY+ + + + + +++S +W K+W +P+ WR
Sbjct: 6 WMHNPTGIYSVKSGYNTL-RTWQTQQINNTSTSSDETLIWKKIWSLHTIPRHKVLLWRIL 182
Query: 277 LDILPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSV 336
D LPVRS L +RG+ P CPRC ET+ H SCPL + +WF S L + FD +
Sbjct: 183 NDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVWFGSNLCINFDNLPNP 362
Query: 337 HDFFLSFLATA----DTNIAGIFLAVLYSVWQARNELLFKWKDSSIDQLLQRASS----- 387
+ F++ L A D I A++Y++W ARN + + + ++QRAS+
Sbjct: 363 N--FIN*LYEAIL*KDECITI*IAAIIYNLWHARNLSVLEDQTILEMDIIQRASNCISDY 536
Query: 388 ------LRPSRVLLDDGPRAVQ--VLPSSWSRPLSGVCKMNIDASVTSSKEAGAGMVMRN 439
PS PR+ + W RP G+ K+N DA++ + + G G+++R+
Sbjct: 537 KQANTQAPPSMARTGYDPRSQHRPAKNTKWKRPNLGLVKVNTDANLQNHGKWGLGIIIRD 716
Query: 440 NHGEVLAAATTELGPVLSPVLAEALGMRWALRLATKLGFRRIWIETD 486
G V+AA+T E + AEA + +R A GF ++ E D
Sbjct: 717 EVGLVMAASTWETDGNDRALEAEAYALLTGMRFAKDCGFXKVXFEGD 857
>BG586862
Length = 804
Score = 105 bits (261), Expect = 6e-23
Identities = 71/231 (30%), Positives = 106/231 (45%), Gaps = 2/231 (0%)
Frame = -1
Query: 258 KLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLV 317
K+W +P+ WR + LPV+ LH+RG+ + CPRC ETV H +C +
Sbjct: 657 KVWGIKTIPRHKSFLWRLLHNALPVKDELHKRGIRCSLLCPRCESKIETVQHLFLNCEVT 478
Query: 318 RMIWFASPLNLRFDEEGSV--HDFFLSFLATADTNIAGIFLAVLYSVWQARNELLFKWKD 375
+ WF S L + F G + HD+ +F+ D A+LYS+W ARN+ +F+ D
Sbjct: 477 QKEWFGSQLGINFHSSGVLHFHDWITNFILKNDEETIIALTALLYSIWHARNQKVFENID 298
Query: 376 SSIDQLLQRASSLRPSRVLLDDGPRAVQVLPSSWSRPLSGVCKMNIDASVTSSKEAGAGM 435
D ++QRASS S + VLPS+ ++ S G G+
Sbjct: 297 VPGDVVIQRASSSLHSFKMAQVSD---SVLPSN---------------AIPSYSLWGIGV 172
Query: 436 VMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRLATKLGFRRIWIETD 486
V RN G +A+ T + AEA G+ A+ A GF + E+D
Sbjct: 171 VARNCEGLAMASGTWLRHGIPCATTAEAWGIYQAMVFAGDCGFSKFEFESD 19
>TC93101
Length = 675
Score = 96.7 bits (239), Expect = 2e-20
Identities = 64/221 (28%), Positives = 103/221 (45%), Gaps = 21/221 (9%)
Frame = -1
Query: 280 LPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSVH-- 337
LPVR L++RGV P CPRC ET +H SC + +WF S L++RF + +++
Sbjct: 663 LPVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVWFGSQLSIRFPDNSTINFS 484
Query: 338 DFFLSFLATADTNIAGIFLAVLYSVWQARNELLFKWKDSSIDQLLQRA---------SSL 388
D+ ++ I A+ YS+W ARN+ +F+ + S D ++Q A +++
Sbjct: 483 DWLFDAISNQTEEIIIKISAITYSIWHARNKAIFENQFVSEDTIIQ*AQNSILAYEQATI 304
Query: 389 RP----------SRVLLDDGPRAVQVLPSSWSRPLSGVCKMNIDASVTSSKEAGAGMVMR 438
+P S + R + S W +PL+ + K N DA++ G G ++R
Sbjct: 303 KPQNPNIVLSSLSATSNTNTTRRRSNVRSRWQKPLNNILKANCDANLQVQGRWGLGCIIR 124
Query: 439 NNHGEVLAAATTELGPVLSPVLAEALGMRWALRLATKLGFR 479
N GE AT + AE + A+ LA GF+
Sbjct: 123 NADGEAKVTATWCINGFDCAATAETYAILAAMYLAKDYGFK 1
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 75.9 bits (185), Expect = 4e-14
Identities = 55/229 (24%), Positives = 93/229 (40%), Gaps = 9/229 (3%)
Frame = -3
Query: 108 LEAKKGYRPSYVWLSLQKSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAES 167
L A G SY W S+ + + ++G +GNG+N E + N +
Sbjct: 762 LNAPLGSWASYAWRSIHSAQHLIKQGAKVIIGNGENTNIWEREMA--WKLTCVTNHPNKH 589
Query: 168 LGVVHVSDLMVTNGGCW---------DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWP 218
+ + + GC + N + + T R ILSI +D + W
Sbjct: 588 SSRAY*APTLYGYEGCRSDDPMRRERNANLINSIFPEGTRRKILSIHPQGPIGEDSYSWE 409
Query: 219 ETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCCETAWRACLD 278
++ G YS ++GY + + + L+ ++WK + P+ WR +
Sbjct: 408 YSKSGHYSVKSGYYVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHFLWRCISN 229
Query: 279 ILPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPLN 327
LP + + R + + C RC ETV+H LF CP R+IW SP++
Sbjct: 228 SLPTAANMRSRHISKDGSCSRCGMESETVNHILFQCPYARLIWATSPIH 82
>BF640104
Length = 344
Score = 67.4 bits (163), Expect = 1e-11
Identities = 38/111 (34%), Positives = 62/111 (55%), Gaps = 1/111 (0%)
Frame = -1
Query: 409 WSRPLSGVCKMNIDASVTSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRW 468
W +P GV K+N DA++TS G G++ RN++G V+A+ T + P+ AEA G+
Sbjct: 338 WIKPHQGVIKINCDANLTSEDVWGIGVITRNDNGIVMASGTWNRPGFMCPITAEAWGVYQ 159
Query: 469 ALRLATKLGFRRIWIETDCLELYTFWNR-QGGEFSHLATVVFDCRSLLPNF 518
A A GF+ + E D +L + +R + G S+L +++ SL+P F
Sbjct: 158 AALFALDQGFQNVLFENDNEKLISMLSREEEGHRSYLGSIIQSIISLVPRF 6
>BG585866
Length = 828
Score = 67.4 bits (163), Expect = 1e-11
Identities = 38/131 (29%), Positives = 62/131 (47%), Gaps = 4/131 (3%)
Frame = +3
Query: 213 DCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCCETA 272
D + WP +G Y+ ++GYS+I + ++SS W +W+ +
Sbjct: 348 DAYIWPHNSNGVYTAKSGYSWILSQTETVNYNNSS--------WSWIWRLKIPEKYKFFL 503
Query: 273 WRACLDILPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLVRMIW----FASPLNL 328
W AC + +P S+L+ R +V + C RC E EE+ H + C ++IW F+SP
Sbjct: 504 WLACHNAVPTLSLLNHRNMVNSAICSRCGEHEESFFHCVRDCRFSKIIWHKIGFSSP--- 674
Query: 329 RFDEEGSVHDF 339
F SV D+
Sbjct: 675 DFFSSSSVQDW 707
>BG584442
Length = 775
Score = 56.2 bits (134), Expect = 3e-08
Identities = 30/71 (42%), Positives = 42/71 (58%), Gaps = 1/71 (1%)
Frame = +1
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRT-MHWLKWDTLCRPKGCGGIGFRD 69
+I SYVMS FLLL+ +IE++++ F W +R MHW+ + L K GG+GF D
Sbjct: 454 SISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVGENRKGMHWMS*EKLFVHKNYGGMGFTD 633
Query: 70 FKAFNTALLAK 80
F FN +L K
Sbjct: 634 FTTFNIPMLGK 666
>BQ148771
Length = 680
Score = 55.1 bits (131), Expect = 7e-08
Identities = 27/99 (27%), Positives = 46/99 (46%)
Frame = -3
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDF 70
A+P Y M ++ +I+++ +F WG R H + W+T+ +PK G+G R
Sbjct: 504 AVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTEVSRRYHAVGWETMSKPKTIYGLGLRRL 325
Query: 71 KAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLE 109
N A + K W I + S+ ++ + Y ES E
Sbjct: 324 DVMNKACIMKLGWSIYSGSNSLCTEVMRGKYQRSESLEE 208
>BG646730
Length = 799
Score = 52.4 bits (124), Expect = 5e-07
Identities = 38/137 (27%), Positives = 62/137 (44%), Gaps = 2/137 (1%)
Frame = +1
Query: 113 GYRPSYVWLSLQKSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVH 172
G +PSY W S+ + G W + NG+N+R +D W+P+ + + +
Sbjct: 388 GNQPSYAWRSMFNIKDVIDLGSRWSISNGQNVRIWKDDWLPNQTGFKVCSPMVDFEEATC 567
Query: 173 VSDLMVTNGGCWDRNKVEFLCWPPTARAILSIPLPLQQRDD--CFFWPETRDGCYSTRTG 230
+S+L+ TN R+ V + A+ IL+IPL + D +W RD YS R+
Sbjct: 568 ISELIDTNTKSSKRDLVRNVFSAFEAKQILNIPLSWRLWPDRRILYW--ERDRNYSVRSA 741
Query: 231 YSFI*KKFSEGEASSSS 247
+ I + SSS
Sbjct: 742 HHLIKVATKDSPEVSSS 792
>AJ497561 weakly similar to GP|10140689|gb putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 621
Score = 48.1 bits (113), Expect = 9e-06
Identities = 22/62 (35%), Positives = 32/62 (51%)
Frame = +2
Query: 91 SMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWIFQEGGLWRVGNGKNIRA*EDK 150
S+L+KI K YFP+ F A G+ PS+ W SL + + G W +G+G I
Sbjct: 410 SLLSKILKFKYFPQWDFSYANLGHNPSFTWRSLLSTQSLLTLGHRWMIGDGSQINVSSMS 589
Query: 151 WV 152
W+
Sbjct: 590 WI 595
>BG449067
Length = 578
Score = 48.1 bits (113), Expect = 9e-06
Identities = 26/78 (33%), Positives = 43/78 (54%)
Frame = -2
Query: 409 WSRPLSGVCKMNIDASVTSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRW 468
W +P + K+N DA+++S+ G G+V RN+ G +A+ T S AEA G+
Sbjct: 307 WKKPEKDIIKLNSDANLSSTDLWGIGVVARNDEGFAMASGTWFRFGFPSATTAEAWGIYQ 128
Query: 469 ALRLATKLGFRRIWIETD 486
A+ A + GF ++ E+D
Sbjct: 127 AMIFAGEYGFSKVQFESD 74
>TC83093
Length = 477
Score = 47.8 bits (112), Expect = 1e-05
Identities = 34/138 (24%), Positives = 63/138 (45%), Gaps = 12/138 (8%)
Frame = -3
Query: 361 SVWQARNELLFKWKDSSIDQLLQRAS----------SLRPSRVLLDDGPRAVQVLPSSW- 409
++W +RN+L+F SS + +++++ S+ + + Q L +
Sbjct: 475 NIWFSRNKLIFNNNFSSEEDTIKQSNTNIQDYITCDSINKTSTCMTQTGTTTQTLSRNVY 296
Query: 410 -SRPLSGVCKMNIDASVTSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRW 468
P + K+ I+A+++ G + RNN G+VLA A L + EA +
Sbjct: 295 *EPPSNNTIKVKINANLSKENTWGLSAICRNNFGDVLATAIWCLPGTHNSAAVEAFAIYL 116
Query: 469 ALRLATKLGFRRIWIETD 486
A+ LA K GF + +E+D
Sbjct: 115 AMELADKKGFHELELESD 62
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 47.0 bits (110), Expect = 2e-05
Identities = 19/50 (38%), Positives = 31/50 (62%)
Frame = -3
Query: 57 CRPKGCGGIGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKES 106
C P+ GG+G RD + N +LLAK WWR++ S+ ++ + +Y P+ S
Sbjct: 969 CLPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYGPRVS 820
>TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 1222
Score = 43.5 bits (101), Expect = 2e-04
Identities = 27/92 (29%), Positives = 47/92 (50%)
Frame = +2
Query: 399 PRAVQVLPSSWSRPLSGVCKMNIDASVTSSKEAGAGMVMRNNHGEVLAAATTELGPVLSP 458
P + ++ W +P G K+N D SV + + AG G ++R+ GE + A ++ P
Sbjct: 947 PVSRSIIWCEWKKPEIGWTKLNTDGSV-NKETAGFGGLLRDYRGEPICAFVSK-APQGDT 1120
Query: 459 VLAEALGMRWALRLATKLGFRRIWIETDCLEL 490
L E + L L+ LG + IW+E+D + +
Sbjct: 1121 FLVELWAIWRGLVLSLGLGIKSIWVESDSMSV 1216
>BG456581
Length = 683
Score = 43.1 bits (100), Expect = 3e-04
Identities = 32/118 (27%), Positives = 46/118 (38%), Gaps = 5/118 (4%)
Frame = +2
Query: 259 LWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLVR 318
+W + P WR D LP L RG C C E ET H C V
Sbjct: 80 IWNSCIPPSHSFICWRLAHDRLPTDDNLSSRGCALVSMCSFCLEQVETSDHLFLRCKFVV 259
Query: 319 MIWFASPLNLRFDEEGSVHDFFLSFLAT-ADTNIAGIFLA----VLYSVWQARNELLF 371
+W LR + S LS L + + +++A +++S+W ARN + F
Sbjct: 260 TLWSWLCSQLRVGLDFSSFKALLSSLPRHCSSQVRDLYVAAVVHMVHSIWWARNNVRF 433
>BF631765
Length = 498
Score = 42.4 bits (98), Expect = 5e-04
Identities = 37/141 (26%), Positives = 58/141 (40%)
Frame = +1
Query: 324 SPLNLRFDEEGSVHDFFLSFLATADTNIAGIFLAVLYSVWQARNELLFKWKDSSIDQLLQ 383
SPL + + + + LA + + +F L+ +W+ RN+ +F D +
Sbjct: 25 SPLGIHVPVQLDLLSWMEKCLAVPEQRVKQLFGTSLWLIWKHRNQKVFNNVDFDPPLVSA 204
Query: 384 RASSLRPSRVLLDDGPRAVQVLPSSWSRPLSGVCKMNIDASVTSSKEAGAGMVMRNNHGE 443
+ + L G V V+P P G K+N+ A G G + RN G
Sbjct: 205 LVEEFSIANLPLKRG--QVCVVPR*QPPPF-GSIKVNVGAGCFGDGSTGWGFIARNAAGL 375
Query: 444 VLAAATTELGPVLSPVLAEAL 464
L+A T G SP+LAE L
Sbjct: 376 ALSAETKSDGISYSPLLAECL 438
>AW980456
Length = 779
Score = 42.0 bits (97), Expect = 6e-04
Identities = 25/90 (27%), Positives = 37/90 (40%)
Frame = -2
Query: 198 ARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWM 257
A I+S PL D W + + G Y ++ Y F ++ + S+ + P W
Sbjct: 253 ADKIMSTPLISHA*LDRLIWKDEKHGKYYVKSAYRFCVEELFD------SSYLHRPGNWS 92
Query: 258 KLWKADAVPQCCETAWRACLDILPVRSILH 287
+WK P+ WR C LP R LH
Sbjct: 91 GIWKLKVPPKVQNLVWRMCRGCLPTRIRLH 2
>TC82442
Length = 578
Score = 42.0 bits (97), Expect = 6e-04
Identities = 27/106 (25%), Positives = 50/106 (46%), Gaps = 3/106 (2%)
Frame = -3
Query: 432 GAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRLATKLGFRRIWIETDCLELY 491
G G ++ N+ G+ + +AT AEA + A+ LA + + E+DC +
Sbjct: 321 GLGCIIHNSAGQAMTSATWSRNGFNCAATAEAYAILAAMHLALTSDLKHMIFESDCEAVI 142
Query: 492 TFWNR---QGGEFSHLATVVFDCRSLLPNFDVFNFTFVRRTSNGAA 534
NR + + S+L +V+ + + FD+ +F F+ R+ N A
Sbjct: 141 RKLNRGIEKDIDISYLGSVLEEIWKISEKFDMCSFRFIPRSGNRVA 4
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 40.4 bits (93), Expect = 0.002
Identities = 39/156 (25%), Positives = 62/156 (39%), Gaps = 11/156 (7%)
Frame = -2
Query: 244 SSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVV---GNPCCPRC 300
SS S ++ P M +W + AWR D LP +S L RGV+ C C
Sbjct: 626 SSGSLSLILPYAEM-IWHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGC 450
Query: 301 SEVEETVHHALFSCPLVRMIWFASPLNLRF--DEEGSVHDFFLSFLATADTNIAG----- 353
+ E+ H SC +W + F + + D F+ F+ + N A
Sbjct: 449 GAL-ESAQHLFLSCSYFASLWSLVRDWIGFVGVDTNVLSDHFVQFVHSTGGNKASQSFLQ 273
Query: 354 -IFLAVLYSVWQARNELLFKWKDSSIDQLLQRASSL 388
I+L + +W RN + F + + +LL + L
Sbjct: 272 LIWLLCAWVLWTERNNMCFNDSITPLPRLLDKVKYL 165
>TC81772
Length = 982
Score = 39.7 bits (91), Expect = 0.003
Identities = 14/40 (35%), Positives = 24/40 (60%)
Frame = -2
Query: 113 GYRPSYVWLSLQKSAWIFQEGGLWRVGNGKNIRA*EDKWV 152
G+ PSY+W S+ + +EG W++G+G +I D W+
Sbjct: 714 GHNPSYIWCSV*TLRMVLEEGHHWKIGDGSSINIWTDHWL 595
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.139 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,494,774
Number of Sequences: 36976
Number of extensions: 407203
Number of successful extensions: 2336
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 2294
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2323
length of query: 565
length of database: 9,014,727
effective HSP length: 101
effective length of query: 464
effective length of database: 5,280,151
effective search space: 2449990064
effective search space used: 2449990064
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0118c.1


