

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0117b.14
(458 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
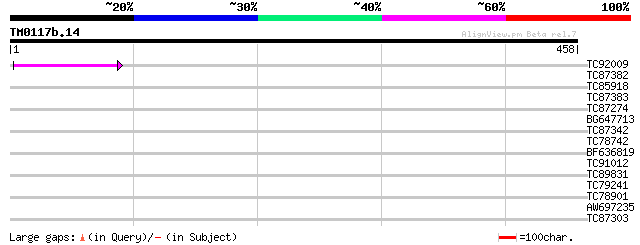
Score E
Sequences producing significant alignments: (bits) Value
TC92009 42 6e-04
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 38 0.007
TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreu... 36 0.035
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 35 0.079
TC87274 similar to PIR|S51232|S51232 gibberellin-responsive ovar... 32 0.51
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 32 0.51
TC87342 PIR|S41431|S41431 GTP-binding protein ras-like - fava b... 30 1.5
TC78742 similar to GP|3819164|emb|CAA09989.1 cytosolic chaperoni... 29 4.3
BF636819 29 4.3
TC91012 similar to GP|13605698|gb|AAK32842.1 AT5g64940/MXK3_17 {... 28 7.4
TC89831 similar to PIR|H96569|H96569 unknown protein 54928-5675... 28 9.7
TC79241 similar to GP|21740501|emb|CAD40825. OSJNBa0006B20.16 {O... 28 9.7
TC78901 similar to GP|21734794|gb|AAM76972.1 reduced vernalizati... 28 9.7
AW697235 28 9.7
TC87303 homologue to GP|4586449|dbj|BAA76393.1 cytochrome c oxid... 28 9.7
>TC92009
Length = 974
Score = 41.6 bits (96), Expect = 6e-04
Identities = 23/90 (25%), Positives = 42/90 (46%), Gaps = 2/90 (2%)
Frame = +2
Query: 4 EFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGE-AFMWWFCWRRRNQKATWWEF 62
+F G + GW+ E + +K A ++ E A +W++ W + N A W F
Sbjct: 695 QFTGTDPAGWITRAEMFFADNEIHSCDKLQWAFMSMEDEKALLWFYSWCQENPDADWKSF 874
Query: 63 VEALLRKFEPELEPYM-PEPVQDSEEEEIP 91
A++R+F +E + VQ+ ++E P
Sbjct: 875 SMAMIREFGARMEHRTDKQMVQNHDQESEP 964
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 38.1 bits (87), Expect = 0.007
Identities = 24/80 (30%), Positives = 37/80 (46%), Gaps = 10/80 (12%)
Frame = +2
Query: 3 PEFDGR----EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQK-- 56
P+F+G + WL +E+ E K V EE+K L A +WW +RR ++
Sbjct: 629 PDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREG 808
Query: 57 ----ATWWEFVEALLRKFEP 72
TW + + L RK+ P
Sbjct: 809 KSKIKTWDKMRQKLTRKYLP 868
>TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreurys tristis},
partial (8%)
Length = 4117
Score = 35.8 bits (81), Expect = 0.035
Identities = 37/114 (32%), Positives = 51/114 (44%), Gaps = 5/114 (4%)
Frame = -1
Query: 71 EPELEPYMPE--PVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPYSPE 128
+PELEP E V E+E K E E + +VE+ EAN A + S E
Sbjct: 2638 KPELEPLATEEPKVTKEPEKESQEKSEEAEQPKAIAIVESTVEANEEKTAAA----ISEE 2471
Query: 129 NS--EMEASRKDSEFAMAAE-QTAEVSRCQPVTVVDYIDCTLTAKGEKIDAEGE 179
E+E + E M E TAE+ + +P VV ++ T T K E + E E
Sbjct: 2470 TKIIEVEPTETVKEEPMVTEVATAEIVKEEP--VVTEVEATETLKEELVATEVE 2315
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 34.7 bits (78), Expect = 0.079
Identities = 23/80 (28%), Positives = 38/80 (46%), Gaps = 10/80 (12%)
Frame = -2
Query: 3 PEFDGR----EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQK-- 56
P+F+G + WL ++E+ + K V EE+K L A +WW +RR ++
Sbjct: 724 PDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREG 545
Query: 57 ----ATWWEFVEALLRKFEP 72
TW + + L RK+ P
Sbjct: 544 KSKIKTWEKMRQKLTRKYLP 485
>TC87274 similar to PIR|S51232|S51232 gibberellin-responsive ovarian protein
G14 - garden pea (fragment), partial (45%)
Length = 734
Score = 32.0 bits (71), Expect = 0.51
Identities = 28/112 (25%), Positives = 52/112 (46%), Gaps = 9/112 (8%)
Frame = +1
Query: 72 PELEPYMPEPVQ--DSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIK-----P 124
P EP E V ++EE E+P +EAE V EAKPE + K E+K
Sbjct: 184 PASEPATTEEVVAVENEETEVP-----VEAESTQVTQEAKPEVENPEPEKVEVKEEVSAE 348
Query: 125 YSPENSEMEAS--RKDSEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKI 174
+ E +E E++ + +++ ++ E+ E + + + ++ T + K+
Sbjct: 349 EAKETTETESAPEKVEAKEEVSTEEAKETTEIESAAIAAPVEEVTTTEA*KV 504
Score = 28.1 bits (61), Expect = 7.4
Identities = 16/44 (36%), Positives = 22/44 (49%)
Frame = +1
Query: 71 EPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANS 114
+PE+E PE V+ EE +E E E VEAK E ++
Sbjct: 286 KPEVENPEPEKVEVKEEVSAEEAKETTETESAPEKVEAKEEVST 417
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 32.0 bits (71), Expect = 0.51
Identities = 22/78 (28%), Positives = 37/78 (47%), Gaps = 10/78 (12%)
Frame = +2
Query: 3 PEFDGR----EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRR------ 52
P+F G+ E WL +E+ + K V+EE+K L A +WW +R
Sbjct: 347 PDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRKRNCEG 526
Query: 53 RNQKATWWEFVEALLRKF 70
+++ TW + + L RK+
Sbjct: 527 KSKIKTWDKMRQKLTRKY 580
>TC87342 PIR|S41431|S41431 GTP-binding protein ras-like - fava bean,
complete
Length = 995
Score = 30.4 bits (67), Expect = 1.5
Identities = 18/62 (29%), Positives = 33/62 (53%), Gaps = 1/62 (1%)
Frame = +3
Query: 125 YSPENSEMEASRKDSEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKIDAEGEIK-IK 183
Y E S +EA+ ++ FA Q + + V + + ++ AKGEKID + ++ +K
Sbjct: 525 YFMETSALEATNVENAFAEVLTQIYRIVSKKAVEGAENGNASVPAKGEKIDLKNDVSALK 704
Query: 184 RL 185
R+
Sbjct: 705 RV 710
>TC78742 similar to GP|3819164|emb|CAA09989.1 cytosolic chaperonin
delta-subunit {Glycine max}, partial (38%)
Length = 755
Score = 28.9 bits (63), Expect = 4.3
Identities = 12/35 (34%), Positives = 21/35 (59%)
Frame = +3
Query: 361 VVFPLSSHIHAFTRSRARLAQLDFSELGQSLTTRP 395
V+ PL +HIH+F +SR + F + S+ ++P
Sbjct: 9 VLLPLITHIHSFPKSRVFIFTFLFVRVSHSILSQP 113
>BF636819
Length = 635
Score = 28.9 bits (63), Expect = 4.3
Identities = 12/23 (52%), Positives = 14/23 (60%)
Frame = -2
Query: 45 MWWFCWRRRNQKATWWEFVEALL 67
+WWF WRR +Q T EF LL
Sbjct: 409 VWWFWWRRCHQPVTR*EFQTKLL 341
>TC91012 similar to GP|13605698|gb|AAK32842.1 AT5g64940/MXK3_17 {Arabidopsis
thaliana}, partial (8%)
Length = 616
Score = 28.1 bits (61), Expect = 7.4
Identities = 13/44 (29%), Positives = 20/44 (44%)
Frame = -3
Query: 22 EAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEFVEA 65
E + + EK + + GE W WRR A W+F+E+
Sbjct: 200 EKREMERVEKEKRLRRVICGERKWSWDSWRREEVVAINWKFLES 69
>TC89831 similar to PIR|H96569|H96569 unknown protein 54928-56750
[imported] - Arabidopsis thaliana, partial (39%)
Length = 897
Score = 27.7 bits (60), Expect = 9.7
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -2
Query: 231 SCLCTAVTANRLPDTFFCWTEVFIQ 255
S LCTA+T+N DTF T F++
Sbjct: 317 SLLCTAITSNSSVDTFVATTARFVK 243
>TC79241 similar to GP|21740501|emb|CAD40825. OSJNBa0006B20.16 {Oryza
sativa}, partial (77%)
Length = 1285
Score = 27.7 bits (60), Expect = 9.7
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = +3
Query: 70 FEPELEPYMPEPVQDSEEEEIPGKQEVLEAE 100
FEP LEP + + + + EE E+ +++V + E
Sbjct: 303 FEPFLEPTIAQEISEDEEVEVEEEEDVDDVE 395
>TC78901 similar to GP|21734794|gb|AAM76972.1 reduced vernalization response
1 {Arabidopsis thaliana}, partial (37%)
Length = 1760
Score = 27.7 bits (60), Expect = 9.7
Identities = 26/82 (31%), Positives = 44/82 (52%), Gaps = 13/82 (15%)
Frame = -2
Query: 354 FRRRWLGVVFPL--SSHIHAFTRSRA------RLAQLDFSEL-----GQSLTTRPKPIML 400
FRRR+L + FPL SS ++ F SRA RL +L++ +L GQ + R I++
Sbjct: 958 FRRRFLDI*FPLLYSSFLNLFISSRAQLQLFIRLFKLNYIKLETYISGQLVVYR---IVI 788
Query: 401 NRRNGPKRNFSFSIIYRPVHSL 422
+ R+ P ++I +R + +
Sbjct: 787 SGRSIPTFTPVYAIKFRSIKQI 722
>AW697235
Length = 846
Score = 27.7 bits (60), Expect = 9.7
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -3
Query: 231 SCLCTAVTANRLPDTFFCWTEVFIQ 255
S LCTA+T+N DTF T F++
Sbjct: 286 SLLCTAITSNSSVDTFVATTARFVK 212
>TC87303 homologue to GP|4586449|dbj|BAA76393.1 cytochrome c oxidase subunit
6b-1 {Oryza sativa (japonica cultivar-group)}, partial
(48%)
Length = 905
Score = 27.7 bits (60), Expect = 9.7
Identities = 19/61 (31%), Positives = 25/61 (40%)
Frame = +1
Query: 63 VEALLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEI 122
VEA++ K E P P ++S E P +E + E VVE K EI
Sbjct: 250 VEAVVEKTVEETIPAAPVVAEESSEVTPPAAEENAQEESSETVVEENSGDQEEAEEKPEI 429
Query: 123 K 123
K
Sbjct: 430 K 432
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,441,964
Number of Sequences: 36976
Number of extensions: 240715
Number of successful extensions: 1522
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1495
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1518
length of query: 458
length of database: 9,014,727
effective HSP length: 99
effective length of query: 359
effective length of database: 5,354,103
effective search space: 1922122977
effective search space used: 1922122977
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0117b.14


