

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
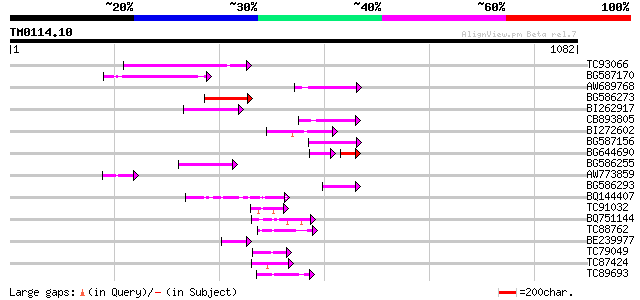
Query= TM0114.10
(1082 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 140 2e-33
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 139 5e-33
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 108 9e-24
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 95 1e-19
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 82 1e-15
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 80 3e-15
BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%) 75 2e-13
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 75 2e-13
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 52 2e-11
BG586255 similar to GP|7682800|gb Hypothetical protein T15F17.l ... 58 2e-08
AW773859 55 1e-07
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 54 2e-07
BQ144407 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 49 8e-06
TC91032 weakly similar to GP|20196995|gb|AAB91977.2 expressed pr... 49 1e-05
BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like ... 48 2e-05
TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Ar... 47 3e-05
BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryz... 46 7e-05
TC79049 weakly similar to PIR|T10798|T10798 pherophorin-S - Volv... 46 9e-05
TC87424 similar to GP|19699309|gb|AAL91265.1 AT4g10840/F25I24_50... 45 1e-04
TC89693 weakly similar to GP|21104633|dbj|BAB93225. hypothetical... 45 2e-04
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 140 bits (354), Expect = 2e-33
Identities = 86/249 (34%), Positives = 125/249 (49%), Gaps = 5/249 (2%)
Frame = +1
Query: 218 FGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRYYVLFLDDFTDFLWTFPLSN 276
FG ++ F ++ + T D +HSDLW S V S G RY + +DDF +W + L
Sbjct: 40 FGNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRY 219
Query: 277 KSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQ 336
K++ F F + T NVK L DN EF ++ F+ +C HG+ + P Q
Sbjct: 220 KNETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQ 399
Query: 337 NGKAERKIRTINNMIRTLLAHASV--PPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLY 394
NG AER IRT+ R +L++A + W A A +L+N P +L P +
Sbjct: 400 NGVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWS 579
Query: 395 RRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYK--CFDLSHRKII 452
Y++LR+FGC Y LV KL PR+ C+FL Y +GY+ C D +K+I
Sbjct: 580 GNLVDYSNLRIFGCPAYALVND---GKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLI 750
Query: 453 ISRHVIFDE 461
+SR V F+E
Sbjct: 751 LSRDVTFNE 777
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 139 bits (351), Expect = 5e-33
Identities = 70/206 (33%), Positives = 118/206 (56%)
Frame = -3
Query: 179 SLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVMPFD 238
+LWH+RLGHP AL + + +E+ N CE+C+ GKH + F ++ V FD
Sbjct: 587 ALWHARLGHPHGRALNLMLPG--VVFENKN----CEACILGKHCKNVFPRTSTVYENCFD 426
Query: 239 ILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHN 298
++++DLWT+P LS H+Y+V F+D+ + + W + +K +V + F + + H+
Sbjct: 425 LIYTDLWTAPSLSRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAK 246
Query: 299 VKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHA 358
+K L+ DNG E+ + +F ++ HG++ + SCP+T QNG A+RK + + + R+L+ A
Sbjct: 245 IKILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQA 66
Query: 359 SVPPSFWHHALQMATYLLNIIPRKSL 384
+V A YL+N IP K L
Sbjct: 65 NV---------STACYLINWIPTKVL 15
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 108 bits (271), Expect = 9e-24
Identities = 58/129 (44%), Positives = 74/129 (56%), Gaps = 1/129 (0%)
Frame = +1
Query: 544 TRSMRGIYKPRKLFNLSVTIDDPTISPLPKN-PKLALSDPNWKSAMQSEFDALIRNKTWD 602
TR+ G KP+ P P++ L LSDP W AM++E+ ALI NKTWD
Sbjct: 4 TRTKTGKLKPKVFLT----------EPCPQDCENLPLSDPRWLQAMKTEYKALIDNKTWD 153
Query: 603 LVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATI 662
LVP P I C W++ K +G ++KARLV G SQ G D ETFSPV+KP TI
Sbjct: 154 LVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFSQTLGCDYTETFSPVIKPVTI 333
Query: 663 RTVLTIALS 671
R +LTIA++
Sbjct: 334 RLILTIAIT 360
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 95.1 bits (235), Expect = 1e-19
Identities = 44/92 (47%), Positives = 62/92 (66%)
Frame = -2
Query: 372 ATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVF 431
A YL+N IP + L + +P ++L +R PS T++RVFGCLCY LVP NKL+ RS +F
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 432 LGYPLHHRGYKCFDLSHRKIIISRHVIFDETR 463
+GY +GYKC+D R++++SR V F E R
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIEER 429
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 81.6 bits (200), Expect = 1e-15
Identities = 42/114 (36%), Positives = 61/114 (52%)
Frame = +1
Query: 332 HTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQ 391
HT QNG AER RT+ R +L A + SFW A++ A Y++N P + +P +
Sbjct: 82 HTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPME 261
Query: 392 LLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFD 445
+ + Y+ L VFGC Y + S KL P+S C+FLGY + +GY +D
Sbjct: 262 MWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 80.5 bits (197), Expect = 3e-15
Identities = 47/118 (39%), Positives = 62/118 (51%)
Frame = +3
Query: 552 KPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVN 611
K + L L++T D T K+ K W+++M +E +A RN TW+L S
Sbjct: 3 KTQNLGMLTMTSDPTTFEEAVKSEK-------WRASMNNEMEATERNNTWELTDLRSGAK 161
Query: 612 IIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIA 669
I WIF K NG E+YKARLV G SQ GVD E F+PV + TIR V+ +A
Sbjct: 162 TIGLKWIFKTKLNENGEIEKYKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALA 335
>BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%)
Length = 488
Score = 74.7 bits (182), Expect = 2e-13
Identities = 49/143 (34%), Positives = 69/143 (47%), Gaps = 8/143 (5%)
Frame = +1
Query: 491 QWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPPQPVPP--------IRTM 542
Q +SP ++ V + S + +++ A+ +SP+PP PP P + +
Sbjct: 13 QTSSPSSPNSNEVAQNIQISSPASPTSNELANITTSPTPPPPPPPTARPHNNHTTNVGGI 192
Query: 543 ATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWD 602
TRS I K K NL V P P N AL D +W+S MQ EFDAL N TWD
Sbjct: 193 ITRSQNNIVKTIKKLNLHVR---PLCPIEPSNITQALRDHDWRSIMQDEFDALQNNNTWD 363
Query: 603 LVPRPSDVNIIRCMWIFHHKTKA 625
LV R N++ C F ++TK+
Sbjct: 364 LVSRSFAQNLVGCKLGFSYQTKS 432
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 74.7 bits (182), Expect = 2e-13
Identities = 40/101 (39%), Positives = 58/101 (56%)
Frame = -1
Query: 571 LPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFE 630
+P++ + A+ D WK ++ +E A+I+N TW P + WIF K KA+G E
Sbjct: 561 IPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWIFTIKYKADGSIE 382
Query: 631 RYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
R K RLV G + G D ETF+PV K TIR VL++A++
Sbjct: 381 RKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVN 259
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 52.0 bits (123), Expect(2) = 2e-11
Identities = 23/51 (45%), Positives = 30/51 (58%)
Frame = -3
Query: 572 PKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHK 622
PKN K AL D +W ++MQ E R+K W LVPRP +I W+F +K
Sbjct: 585 PKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRWVFRNK 433
Score = 35.8 bits (81), Expect(2) = 2e-11
Identities = 17/39 (43%), Positives = 24/39 (60%)
Frame = -2
Query: 631 RYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIA 669
R K++LV G +Q G+D DE FSPV + IR ++ A
Sbjct: 418 RNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFA 302
>BG586255 similar to GP|7682800|gb Hypothetical protein T15F17.l {Arabidopsis
thaliana}, partial (12%)
Length = 436
Score = 58.2 bits (139), Expect = 2e-08
Identities = 36/112 (32%), Positives = 55/112 (48%)
Frame = -1
Query: 323 GLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRK 382
G ++ S H +QNG AE I+ I + R LL + P S W HA+ A L+ I P
Sbjct: 415 G*VWNTSVVHVHTQNGLAESFIKRIQLITRPLLMRSKQPVSAWGHAVLHAAELIRIRP-S 239
Query: 383 SLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGY 434
S SP+QLL P +H++ FG Y + K+ P+ +++G+
Sbjct: 238 SEHKYSPSQLLSGHVPDVSHIKTFGYPVYVPIAPPHRTKMGPQRRMGIYVGF 83
>AW773859
Length = 538
Score = 55.1 bits (131), Expect = 1e-07
Identities = 27/69 (39%), Positives = 41/69 (59%), Gaps = 1/69 (1%)
Frame = -3
Query: 178 QSLWHSRLGHPGSSALQSLRSN-KFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVMP 236
++LWH RLGH + L SL SN FIT ++ +VC+ C + +H +LPF S N
Sbjct: 236 EALWHFRLGHLSNRKLLSLHSNFPFIT---IDQNSVCDICHYSRHKKLPFQLSTNRASKC 66
Query: 237 FDILHSDLW 245
+++ H D+W
Sbjct: 65 YELFHFDIW 39
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 54.3 bits (129), Expect = 2e-07
Identities = 29/73 (39%), Positives = 42/73 (56%)
Frame = +2
Query: 597 RNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPV 656
+ +T LV +P+ V I WI+ K +G +YKARLV G + G+D DE F+PV
Sbjct: 47 QKQTLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPV 226
Query: 657 VKPATIRTVLTIA 669
V+ TI +L +A
Sbjct: 227 VRIETI*LLLALA 265
>BQ144407 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (23%)
Length = 1217
Score = 49.3 bits (116), Expect = 8e-06
Identities = 64/205 (31%), Positives = 87/205 (42%), Gaps = 6/205 (2%)
Frame = -3
Query: 335 SQNGKAERKIRTINNMIRTLLAHA---SVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQ 391
S + A +I + + ++ +LL A + PP H+L ++ L ++P L +
Sbjct: 1032 SHSAYAHCEISSRSPIVSSLLFPADRDATPPLTLRHSLSLS--LAALLP------LCASP 877
Query: 392 LLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVF--LGYPLHHRGYKCFDLSHR 449
L R P +H L P PS L P SSP L PLH F LS
Sbjct: 876 PLRLRSPVVSHTT--SILYAPPPPSILPPILTPLSSPISLRSLLSPLH------FTLSRL 721
Query: 450 KIIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVP-PAVT 508
+S + P P PPPS Y S P PS+ T+P P+S+ P PA
Sbjct: 720 SRCVSH--LHSSPPPPTLIHPLPPPSPYP-ISPP--PSLDISITAPSTRPSSLTPTPASP 556
Query: 509 PSQLPTTSTSSSASPPSSPSPPSPP 533
P +P + SS SP S SPP PP
Sbjct: 555 PPPIPRSPHPSSFSPHSR-SPPRPP 484
Score = 41.6 bits (96), Expect = 0.002
Identities = 27/76 (35%), Positives = 39/76 (50%), Gaps = 2/76 (2%)
Frame = -3
Query: 465 PFATMPTPPPSTYDCFSDPIHPS--IIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSAS 522
P A PTP + + P+H + H + P SP +V+P ++ P Q P SS
Sbjct: 390 PRAPRPTPSFPRHSA-APPLHAPYRLPHLFLHPH-SPPAVIPLSLLPVQGPDLPPVSS-P 220
Query: 523 PPSSPSPPSPPQPVPP 538
PP+ P PP+PP+P P
Sbjct: 219 PPTPPPPPAPPRPAAP 172
Score = 36.6 bits (83), Expect = 0.054
Identities = 28/83 (33%), Positives = 36/83 (42%), Gaps = 7/83 (8%)
Frame = -2
Query: 462 TRFPFATMPTPPP--STYDCFSDPIH-PSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
TR + PTPPP + + P P ++P P P S PP P P +
Sbjct: 424 TR*TALSSPTPPPRPAAHSLLPPPFRCPPASCPLSAPPPLPPSAFPPRGHP---PIAAPG 254
Query: 519 SSA-SPP---SSPSPPSPPQPVP 537
SPP S P PP+PP+P P
Sbjct: 253 PRPRSPPRFFSPPHPPAPPRPPP 185
Score = 36.2 bits (82), Expect = 0.070
Identities = 28/73 (38%), Positives = 34/73 (46%), Gaps = 8/73 (10%)
Frame = -2
Query: 465 PFATMPTP--PPST----YDCFSDPIHPSII-HQWTS-PDPSPTSVVPPAVTPSQLPTTS 516
P A +P P PPS Y C PIHPS++ H + PS PP V S P T
Sbjct: 664 PSAALPLPHIPPSLSRHLYHC---PIHPSVLPHPYPRLTTPSYPPFPPPLVVLSPFPLTP 494
Query: 517 TSSSASPPSSPSP 529
+ + ASP P P
Sbjct: 493 SPTLASPFQHPPP 455
Score = 36.2 bits (82), Expect = 0.070
Identities = 51/201 (25%), Positives = 76/201 (37%), Gaps = 36/201 (17%)
Frame = -1
Query: 374 YLLNIIPRKSLSNLSPT---QLLYRRDPSYTHLRVFGCLCYPLVP-STTINKLQPRSSPC 429
Y L+ I R SL++L+ + LL S+T R YP +P ST+++ P P
Sbjct: 755 YYLHSISR-SLASLAASLTCTLLPHLRLSFTPFRRPPLTPYPPLPLSTSLSLPHPPVRPP 579
Query: 430 VFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSII 489
L P H R + I H + P + T PS+ S P+HP
Sbjct: 578 SPLPPPHHPLLSPVPPTPRRSLPIPAHPL----AHPGFALSTSTPSSPIPPSHPLHPLNC 411
Query: 490 HQWTSPDPSPTSVVPPA----VTPSQLPTTSTSSSAS----------------------- 522
+ P P+P +PP+ + P +P + +S+S
Sbjct: 410 AFFPHPAPAPRGPLPPSPAIPLPPRFMPLIGSPTSSSIRIPPPRSSPYRCSRSKAPISPP 231
Query: 523 -----PPSSPSPPSPPQPVPP 538
PP P PP P +P PP
Sbjct: 230 FLLPPPPPRPPPPPPGRPRPP 168
Score = 30.4 bits (67), Expect = 3.9
Identities = 31/93 (33%), Positives = 37/93 (39%), Gaps = 11/93 (11%)
Frame = -2
Query: 464 FPFATMPT-------PPPSTYDCFSDPIHPSIIHQWTSPDPSPT----SVVPPAVTPSQL 512
FP PT PPP F PI + +SP P P S++PP P +
Sbjct: 508 FPLTPSPTLASPFQHPPPHLQ--FRLPILFTR*TALSSPTPPPRPAAHSLLPP---PFRC 344
Query: 513 PTTSTSSSASPPSSPSPPSPPQPVPPIRTMATR 545
P S SA PP P PP+ PPI R
Sbjct: 343 PPASCPLSAPPPLPP-SAFPPRGHPPIAAPGPR 248
>TC91032 weakly similar to GP|20196995|gb|AAB91977.2 expressed protein
{Arabidopsis thaliana}, partial (37%)
Length = 1366
Score = 48.5 bits (114), Expect = 1e-05
Identities = 33/84 (39%), Positives = 38/84 (44%), Gaps = 10/84 (11%)
Frame = -2
Query: 459 FDETRFPFATMPTPP-----PSTYDCFSDPIHPSIIHQWTSPDPSPTS-----VVPPAVT 508
F T P A + PP PST P PS H T P SPTS +PP +T
Sbjct: 564 FSTTHTPTALVHLPPQFSTPPSTPSTL--PQQPSTSHSSTLPSHSPTSHSTLQSLPPTLT 391
Query: 509 PSQLPTTSTSSSASPPSSPSPPSP 532
P+T S +SPP PSPP P
Sbjct: 390 TLSSPSTPHFSPSSPPQPPSPPHP 319
>BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like protein
(clone pMG14) - common tobacco (fragment), partial (11%)
Length = 632
Score = 47.8 bits (112), Expect = 2e-05
Identities = 41/139 (29%), Positives = 62/139 (44%), Gaps = 17/139 (12%)
Frame = +1
Query: 462 TRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSA 521
TR ++P+ PPST P +++ T+P +P S PPA +PS PT +S +
Sbjct: 163 TRRRLLSLPSSPPST-----SPSPHALLRSPTAPPAAPLS--PPASSPSS-PTRPSSLFS 318
Query: 522 SPPSSPS-----PPSPPQPVPPIRTMATRSMRGIYKPRK---------LFNLSVTIDDPT 567
PP SPS PPS P P+ P+ + + + P + V + P+
Sbjct: 319 PPPPSPSRSRPPPPSTPSPLLPLPLLPLATEPSPFPPSSPPPRPASSPSLSPPVLLFPPS 498
Query: 568 ISP---LPKNPKLALSDPN 583
SP P P AL+ P+
Sbjct: 499 ASPPPWSPSRPPSALAPPS 555
Score = 39.7 bits (91), Expect = 0.006
Identities = 29/83 (34%), Positives = 41/83 (48%), Gaps = 2/83 (2%)
Frame = +1
Query: 465 PFATMPTP-PPSTYDCFSDPIHPSIIHQWTSPDPSPTSVV-PPAVTPSQLPTTSTSSSAS 522
P AT P+P PPS S P P+ +SP SP ++ PP+ +P + S+ +
Sbjct: 397 PLATEPSPFPPS-----SPPPRPA-----SSPSLSPPVLLFPPSASPPPWSPSRPPSALA 546
Query: 523 PPSSPSPPSPPQPVPPIRTMATR 545
PPS PS P+ P P T +R
Sbjct: 547 PPSRPSLPASPPSSPRSSTRPSR 615
Score = 29.6 bits (65), Expect = 6.6
Identities = 35/154 (22%), Positives = 62/154 (39%), Gaps = 7/154 (4%)
Frame = +3
Query: 410 CYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATM 469
C P TTI K++ ++ F G + H G++ + + + + S T A
Sbjct: 72 CSKQPPFTTI-KMKYSAAVLAFAGAVMAHGGHE--EEAAQSTVFSTKYF---TITSCAPE 233
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSS--- 526
T P+ S + P I + P + T+V P TP+ + ++ + +PP+
Sbjct: 234 VTNCPARSTVVSSSVVPVITNSAELPVLTTTAVPVPVTTPAAVYPIPSAPAPAPPAGNGT 413
Query: 527 -PSPPSPPQP---VPPIRTMATRSMRGIYKPRKL 556
P PP P V P+ ++ + KP L
Sbjct: 414 VPVPPVKPTTETGVKPVPVTTGAAVPSVSKPATL 515
>TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 47.4 bits (111), Expect = 3e-05
Identities = 39/118 (33%), Positives = 54/118 (45%), Gaps = 4/118 (3%)
Frame = +3
Query: 474 PSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSS--SASPPSSPSP-- 529
PS Y F++P PS +SP + TS PPA TP+ P ++T+S +++P SSP P
Sbjct: 60 PSPYQ-FNNPTSPSSSPSASSPVSTTTS--PPASTPASSPVSTTTSPPASTPASSPVPTT 230
Query: 530 PSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSA 587
SPP P P ++T S PT SP P A P+ +A
Sbjct: 231 TSPPAPTPASSPVSTNS-------------------PTASPAGSLPAAATPSPSSTTA 347
Score = 42.0 bits (97), Expect = 0.001
Identities = 25/73 (34%), Positives = 37/73 (50%), Gaps = 2/73 (2%)
Frame = +3
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSP--DPSPTSVVPPAVTPSQLPTTSTSSSAS 522
P +T +PP ST S P+ + ++P P PT+ PPA TP+ P ++ S +AS
Sbjct: 120 PVSTTTSPPASTPA--SSPVSTTTSPPASTPASSPVPTTTSPPAPTPASSPVSTNSPTAS 293
Query: 523 PPSSPSPPSPPQP 535
P S + P P
Sbjct: 294 PAGSLPAAATPSP 332
Score = 36.6 bits (83), Expect = 0.054
Identities = 25/72 (34%), Positives = 36/72 (49%), Gaps = 4/72 (5%)
Frame = +3
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVV--PPAVTP--SQLPTTSTSSSASPPS 525
PT P S+ S P+ + ++P SP S PPA TP S +PTT++ + +P S
Sbjct: 84 PTSPSSSPSA-SSPVSTTTSPPASTPASSPVSTTTSPPASTPASSPVPTTTSPPAPTPAS 260
Query: 526 SPSPPSPPQPVP 537
SP + P P
Sbjct: 261 SPVSTNSPTASP 296
Score = 32.7 bits (73), Expect = 0.78
Identities = 29/78 (37%), Positives = 39/78 (49%), Gaps = 4/78 (5%)
Frame = +3
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVT--PSQLPTTSTSSSAS 522
P +T +PP ST S P+ + TSP P+PT P T P+ P S ++A+
Sbjct: 168 PVSTTTSPPASTPA--SSPVPTT-----TSP-PAPTPASSPVSTNSPTASPAGSLPAAAT 323
Query: 523 P-PSSPSPPSP-PQPVPP 538
P PSS + SP P PP
Sbjct: 324 PSPSSTTAGSPSPSSTPP 377
>BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 514
Score = 46.2 bits (108), Expect = 7e-05
Identities = 21/58 (36%), Positives = 31/58 (53%)
Frame = -2
Query: 404 RVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDE 461
R+FGC + + S +K R+ CVF+ Y +GY+C+ RK +SR V F E
Sbjct: 456 RIFGCTSFVHIHSDGRSKFDHRALKCVFIRYSSTQKGYRCYHPPSRKYFVSRDVTFHE 283
>TC79049 weakly similar to PIR|T10798|T10798 pherophorin-S - Volvox carteri,
partial (6%)
Length = 957
Score = 45.8 bits (107), Expect = 9e-05
Identities = 31/83 (37%), Positives = 40/83 (47%), Gaps = 7/83 (8%)
Frame = +1
Query: 463 RFPFAT---MPTPPPSTYDCFSDPIHPSI-IHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
+FP+ + P+ PP +S P PS +HQ+ SP+PSP S P S LP S +
Sbjct: 688 QFPYPSHQLYPSVPPQP*SSYSSPQFPSSQVHQFFSPNPSP-SPHSPLSPSSTLPLPSPT 864
Query: 519 SSASPP---SSPSPPSPPQPVPP 538
PP PS P PP PP
Sbjct: 865 PRPYPPCPSKPPSNPQPPSSSPP 933
>TC87424 similar to GP|19699309|gb|AAL91265.1 AT4g10840/F25I24_50
{Arabidopsis thaliana}, partial (32%)
Length = 1014
Score = 45.4 bits (106), Expect = 1e-04
Identities = 27/85 (31%), Positives = 38/85 (43%), Gaps = 5/85 (5%)
Frame = +2
Query: 462 TRFPFATMPTPPPSTYDCFSDPIHPSIIH----QWTSPDPSPTSVVPPAVTPSQLPTTST 517
T FPF + P + P+H + +H Q+T P PP +T + PT +
Sbjct: 65 TTFPFVLVEIVPQFLSLFYLSPLHTNFLHTILIQFTMPGLVSPGGPPPRITLPETPTQRS 244
Query: 518 SSSASPPSSPSP-PSPPQPVPPIRT 541
SS +P SP P PP P P R+
Sbjct: 245 ESSKTPTQSPLPNKKPPSPSPSSRS 319
>TC89693 weakly similar to GP|21104633|dbj|BAB93225. hypothetical
protein~similar to Arabidopsis thaliana chromosome 3
At3g46550, partial (53%)
Length = 1346
Score = 45.1 bits (105), Expect = 2e-04
Identities = 37/115 (32%), Positives = 50/115 (43%), Gaps = 3/115 (2%)
Frame = +2
Query: 471 TPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTP-SQLPTTSTSSSASPPSSPSP 529
+PP S I P+ +H +SP P+P S PP +T + P S +SSA+ SS S
Sbjct: 155 SPPYFLTHPLSPLIFPTALHSLSSPSPTPIS--PPTLTSITSPPPPSPTSSATTSSSNSS 328
Query: 530 PSPPQ--PVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDP 582
P P +PP+ + S P T P ISP P+ L S P
Sbjct: 329 PGPTSNTSLPPVNSSPLSSKPPAAPP--------TTSAPLISPTPRTLTLLQSIP 469
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.329 0.140 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,927,341
Number of Sequences: 36976
Number of extensions: 844455
Number of successful extensions: 18091
Number of sequences better than 10.0: 644
Number of HSP's better than 10.0 without gapping: 7526
Number of HSP's successfully gapped in prelim test: 386
Number of HSP's that attempted gapping in prelim test: 1601
Number of HSP's gapped (non-prelim): 11929
length of query: 1082
length of database: 9,014,727
effective HSP length: 106
effective length of query: 976
effective length of database: 5,095,271
effective search space: 4972984496
effective search space used: 4972984496
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0114.10


