

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.1
(661 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
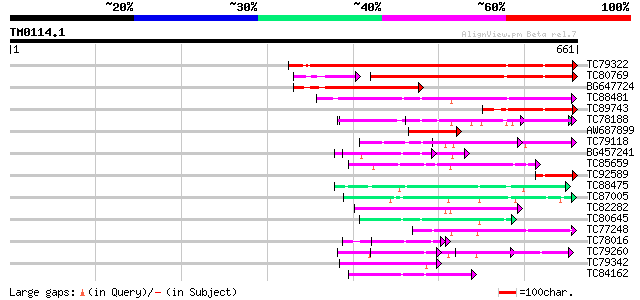
Score E
Sequences producing significant alignments: (bits) Value
TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16 ... 368 e-102
TC80769 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis... 259 9e-82
BG647724 similar to GP|11141605|gb LEUNIG {Arabidopsis thaliana}... 136 2e-32
TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclea... 98 9e-21
TC89743 weakly similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arab... 94 1e-19
TC78188 similar to PIR|C84870|C84870 probable splicing factor [i... 76 5e-14
AW687899 weakly similar to GP|11141605|gb| LEUNIG {Arabidopsis t... 70 2e-12
TC79118 similar to GP|16323188|gb|AAL15328.1 AT5g67320/K8K14_4 {... 61 1e-09
BG457241 homologue to PIR|F84600|F8 coatomer alpha subunit [impo... 59 6e-09
TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein ... 57 3e-08
TC92589 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis... 57 3e-08
TC88475 similar to GP|20260442|gb|AAM13119.1 unknown protein {Ar... 55 7e-08
TC87005 similar to GP|9294061|dbj|BAB02018.1 WD40-repeat protein... 52 1e-06
TC82282 similar to PIR|T08544|T08544 hypothetical protein F27B13... 50 2e-06
TC80645 similar to PIR|T50983|T50983 probable pleiotropic regula... 50 3e-06
TC77248 similar to GP|13489182|gb|AAK27816.1 putative WD-repeat ... 50 3e-06
TC78016 similar to GP|21537191|gb|AAM61532.1 PRL1 protein {Arabi... 50 3e-06
TC79260 weakly similar to GP|9759350|dbj|BAB10005.1 WD-repeat pr... 49 6e-06
TC79342 similar to GP|10177096|dbj|BAB10430. Notchless protein h... 48 1e-05
TC84162 similar to GP|15810485|gb|AAL07130.1 putative WD40-repea... 47 2e-05
>TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16
{Arabidopsis thaliana}, partial (59%)
Length = 1824
Score = 368 bits (945), Expect = e-102
Identities = 180/336 (53%), Positives = 238/336 (70%)
Frame = +2
Query: 326 DAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKV 385
D D+NVESFLS +D D + + F LKR A + + +KGFS E+ + KS KV
Sbjct: 641 DVGSLDDNVESFLSQDD--GDGK-DLFGTLKRNPAEHATDSSKGFSFSEVSSIRKSNGKV 811
Query: 386 LTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFD 445
+ CHFSSDGK++ASAGH+KKV +WNME + ++ + H+ +ITDVRF+P ST ATSSFD
Sbjct: 812 VCCHFSSDGKLLASAGHDKKVVLWNMETLKTQSTPEEHTVIITDVRFRPNSTQLATSSFD 991
Query: 446 RSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHT 505
+VRLWDA++P SL +GH V SLDFHP++ DL CS D N+ IR WN++Q C
Sbjct: 992 TTVRLWDAADPSVSLQAYSGHTSHVASLDFHPKKNDLFCSCDDNNEIRFWNISQYSCTRV 1171
Query: 506 TRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
+GGS QVRFQP+ G LLA A GN +++ DVETD +H L+GH +V +CWD +G++LA
Sbjct: 1172FKGGSTQVRFQPRSGHLLAAASGNVVSLFDVETDRQMHSLQGHSGEVHCVCWDTNGDYLA 1351
Query: 566 SVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
SVS++S ++WS AS G CI EL+S GN F SC+FHP Y +LL+IGGYQSLE W E +K
Sbjct: 1352SVSQESVKVWSLAS-GDCIHELNSSGNMFHSCVFHPSYSNLLVIGGYQSLELWHMAE-NK 1525
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+I AH+G+I+ LA SPV ++ASASHD SVK+WK
Sbjct: 1526CMTIPAHEGVISALAQSPVTGMVASASHDKSVKIWK 1633
>TC80769 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis thaliana},
partial (40%)
Length = 1362
Score = 259 bits (662), Expect(2) = 9e-82
Identities = 127/241 (52%), Positives = 166/241 (68%)
Frame = +3
Query: 421 DTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRM 480
+ HS LITDVRF ATSSFD++VR+WD NP SL TGH VMSLDFHP +
Sbjct: 483 EEHSALITDVRFSASMPRLATSSFDKTVRVWDVDNPGYSLRTFTGHSTSVMSLDFHPNKD 662
Query: 481 DLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDS 540
DL+CS D + IR W++N C+ ++GG+ Q+RFQP+ G+ LA A N ++I+DVET +
Sbjct: 663 DLICSCDGDGEIRYWSINNGSCVRVSKGGTTQMRFQPRLGRYLAAAAENIVSILDVETQA 842
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFH 600
+ LKGH K + S+CWD SG LASVSEDS RIW+ +G+C+ EL G+KF SC+FH
Sbjct: 843 CRYSLKGHTKTIDSVCWDPSGELLASVSEDSVRIWTL--EGECVHELSCNGSKFHSCVFH 1016
Query: 601 PGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
P + SLL+IG YQSLE + E +KT +++AH GLI LA S V L+ASASHD +K+W
Sbjct: 1017PTFPSLLVIGCYQSLELCNMAE-NKTMTLSAHDGLITALAVSTVNGLVASASHDKFIKLW 1193
Query: 661 K 661
K
Sbjct: 1194K 1196
Score = 63.5 bits (153), Expect(2) = 9e-82
Identities = 34/80 (42%), Positives = 47/80 (58%), Gaps = 1/80 (1%)
Frame = +2
Query: 331 DENVESFLSFED-EPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCH 389
D+NVESFLS +D +P D P +S KGF+ ++ + S +K+ CH
Sbjct: 242 DDNVESFLSQDDTDPRD----PVGRCMDVS--------KGFTFSDVNSVRASSSKIACCH 385
Query: 390 FSSDGKVMASAGHEKKVFIW 409
FSSDGK++AS GH+KK IW
Sbjct: 386 FSSDGKLLASGGHDKKAVIW 445
>BG647724 similar to GP|11141605|gb LEUNIG {Arabidopsis thaliana}, partial
(25%)
Length = 757
Score = 136 bits (343), Expect = 2e-32
Identities = 71/152 (46%), Positives = 95/152 (61%)
Frame = +2
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
D+NVESFLS +D +P + R+ + KGF+ E+ + S NKV+ HF
Sbjct: 314 DDNVESFLSHDDT------DPRDPVGRMDVS------KGFTFSELNSVRASTNKVVCSHF 457
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDGK++AS GH+KKV +W ++ + A+ + HS LITDVRF P ATSS+D++VR+
Sbjct: 458 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 637
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
WD NP SL TGH VMSLDFHP + DL
Sbjct: 638 WDVDNPGYSLRTFTGHSAAVMSLDFHPNKDDL 733
>TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclear
ribonucleoprotein [imported] - Arabidopsis thaliana,
partial (60%)
Length = 1836
Score = 98.2 bits (243), Expect = 9e-21
Identities = 80/310 (25%), Positives = 133/310 (42%), Gaps = 7/310 (2%)
Frame = +1
Query: 358 ISATCSRNENKGFSLQEIGCLHKSINKVLT-CHFSSDGKVMASAGHEKKVFIWNMENFQY 416
I T + N EIG ++ LT C FS DGK +A+ +W+M N +
Sbjct: 889 IDWTLKQAANLNLEFSEIGD-----DRPLTGCSFSRDGKGLATCSFTGATKLWSMPNVKK 1053
Query: 417 FASADTHSHLITDVRFQP-GSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDF 475
++ H+ TDV + P AT+S DR+ + W+ + LG GH E++ + F
Sbjct: 1054 VSTLKGHTQRATDVAYSPVHKNHLATASADRTAKYWNDQGAL--LGTFKGHLERLARIAF 1227
Query: 476 HPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNSIT 532
HP L ++ + RLW+V E + G S+ V F + +
Sbjct: 1228 HPSG-KYLGTASYDKTWRLWDVETEEELLLQEGHSRSVYGLDFHHDGSLAASCGLDALAR 1404
Query: 533 IIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVG 591
+ D+ T + L+GH K +L I + +G LA+ ED + RIW K + + +
Sbjct: 1405 VWDLRTGRSVLALEGHVKSILGISFSPNGYHLATGGEDNTCRIWDLRKK-KSLYTIPAHS 1581
Query: 592 NKFQSCIFHPGYRSLLIIGGY-QSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIAS 650
N F P L+ Y + + WS + +++ H+ + L G I +
Sbjct: 1582 NLISQVKFEPQEGYFLVTASYDMTAKVWSSRDFKPVKTLSGHEAKVTSLDVLGDGGYIVT 1761
Query: 651 ASHDNSVKVW 660
SHD ++K+W
Sbjct: 1762 VSHDRTIKLW 1791
>TC89743 weakly similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis
thaliana}, partial (8%)
Length = 603
Score = 94.4 bits (233), Expect = 1e-19
Identities = 48/110 (43%), Positives = 70/110 (63%)
Frame = +3
Query: 552 VLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGG 611
+L ICW L +IW+ S G+C+ EL+S GN+F S +FHP Y +LL++GG
Sbjct: 60 LLEICWHLLVQILV-------KIWNLTS-GECVQELNSTGNQFYSGLFHPSYATLLVVGG 215
Query: 612 YQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
SLE W+ E +K+ +I AH+ +I+ LA+ ++ASAS DNSVK+WK
Sbjct: 216 ISSLELWNMAE-NKSMTIPAHENVISALAECAATGMVASASRDNSVKLWK 362
Score = 32.0 bits (71), Expect = 0.78
Identities = 12/22 (54%), Positives = 17/22 (76%)
Frame = +2
Query: 547 GHDKDVLSICWDRSGNFLASVS 568
GH + V +ICWD +G+ LAS+S
Sbjct: 23 GHPEPVNNICWDTTGDMLASIS 88
Score = 30.8 bits (68), Expect = 1.7
Identities = 18/89 (20%), Positives = 40/89 (44%), Gaps = 4/89 (4%)
Frame = +3
Query: 491 VIRLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKG 547
++++WN+ EC+ Q F P Y LL +S+ + ++ + + +
Sbjct: 96 LVKIWNLTSGECVQELNSTGNQFYSGLFHPSYATLLVVGGISSLELWNMAENKSM-TIPA 272
Query: 548 HDKDVLSICWDRSGNFLASVSED-SARIW 575
H+ + ++ + +AS S D S ++W
Sbjct: 273 HENVISALAECAATGMVASASRDNSVKLW 359
>TC78188 similar to PIR|C84870|C84870 probable splicing factor [imported] -
Arabidopsis thaliana, partial (95%)
Length = 1436
Score = 75.9 bits (185), Expect = 5e-14
Identities = 61/284 (21%), Positives = 112/284 (38%), Gaps = 11/284 (3%)
Frame = +1
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNME-NFQYFASADTHSHLITDVRFQPGSTMFATSS 443
V T F+ G V+AS H+K++F+WN+ + + F H + + D+ + T ++S
Sbjct: 322 VYTMKFNPTGSVVASGSHDKEIFLWNVHGDCKNFMVLKGHKNAVLDLHWTSDGTQIISAS 501
Query: 444 FDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM 503
D+++RLWD + + + H V S R L+ S + +LW++ Q +
Sbjct: 502 PDKTLRLWDTETG-KQIKKMVEHLSYVNSCCPTRRGPPLVVSGSDDGTAKLWDMRQRGSI 678
Query: 504 HT--TRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
T + V F ++ I N + I D+ L+GH + S+ G
Sbjct: 679 QTFPDKYQITAVSFSDASDKIYTGGIDNDVKIWDLRKGEVTMTLQGHQDMITSMQLSPDG 858
Query: 562 NFLASVSED-SARIWSA---ASDGKCI----GELHSVGNKFQSCIFHPGYRSLLIIGGYQ 613
++L + D IW A +C+ G H+ C + P + +
Sbjct: 859 SYLLTNGMDCKLCIWDMRPYAPQNRCVKILEGHQHNFEKNLLKCGWSPDGSKVTAGSSDR 1038
Query: 614 SLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSV 657
+ W T + + H G + P ++ S S D +
Sbjct: 1039MVYIWDTTSRRILYKLPGHNGSVNECVFHPNEPIVGSCSSDKQI 1170
Score = 57.4 bits (137), Expect = 2e-08
Identities = 56/234 (23%), Positives = 102/234 (42%), Gaps = 15/234 (6%)
Frame = +1
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTH-SHLITDVRFQPGSTMFAT 441
N VL H++SDG + SA +K + +W+ E + H S++ + + G + +
Sbjct: 445 NAVLDLHWTSDGTQIISASPDKTLRLWDTETGKQIKKMVEHLSYVNSCCPTRRGPPLVVS 624
Query: 442 SSFDRSVRLWDASNPIRSLG-ILTGHDE-QVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQ 499
S D + +LWD +R G I T D+ Q+ ++ F + NDV ++W++ +
Sbjct: 625 GSDDGTAKLWD----MRQRGSIQTFPDKYQITAVSFSDASDKIYTGGIDNDV-KIWDLRK 789
Query: 500 SECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVE----TDSHLHDLKGH---- 548
E T +G + + P LL + + I D+ + + L+GH
Sbjct: 790 GEVTMTLQGHQDMITSMQLSPDGSYLLTNGMDCKLCIWDMRPYAPQNRCVKILEGHQHNF 969
Query: 549 DKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHP 601
+K++L W G+ + + S D IW S + + +L C+FHP
Sbjct: 970 EKNLLKCGWSPDGSKVTAGSSDRMVYIWDTTS-RRILYKLPGHNGSVNECVFHP 1128
Score = 53.1 bits (126), Expect = 3e-07
Identities = 48/208 (23%), Positives = 94/208 (45%), Gaps = 9/208 (4%)
Frame = +1
Query: 462 ILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH--TTRGGSKQV---RFQ 516
+LTGH V ++ F+P ++ S + I LWNV+ +C + +G V +
Sbjct: 298 LLTGHQSAVYTMKFNPTG-SVVASGSHDKEIFLWNVH-GDCKNFMVLKGHKNAVLDLHWT 471
Query: 517 PQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGN-FLASVSED-SARI 574
Q+++ + ++ + D ET + + H V S C R G + S S+D +A++
Sbjct: 472 SDGTQIISASPDKTLRLWDTETGKQIKKMVEHLSYVNSCCPTRRGPPLVVSGSDDGTAKL 651
Query: 575 WSAASDGKCIGELHSVGNKFQ-SCIFHPGYRSLLIIGGYQS-LEAWSPTEGSKTWSIAAH 632
W D + G + + +K+Q + + + GG + ++ W +G T ++ H
Sbjct: 652 W----DMRQRGSIQTFPDKYQITAVSFSDASDKIYTGGIDNDVKIWDLRKGEVTMTLQGH 819
Query: 633 QGLIAGLADSPVGELIASASHDNSVKVW 660
Q +I + SP G + + D + +W
Sbjct: 820 QDMITSMQLSPDGSYLLTNGMDCKLCIW 903
Score = 35.0 bits (79), Expect = 0.092
Identities = 16/75 (21%), Positives = 33/75 (43%)
Frame = +1
Query: 376 GCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPG 435
G H +L C +S DG + + ++ V+IW+ + + H+ + + F P
Sbjct: 952 GHQHNFEKNLLKCGWSPDGSKVTAGSSDRMVYIWDTTSRRILYKLPGHNGSVNECVFHPN 1131
Query: 436 STMFATSSFDRSVRL 450
+ + S D+ + L
Sbjct: 1132 EPIVGSCSSDKQIYL 1176
>AW687899 weakly similar to GP|11141605|gb| LEUNIG {Arabidopsis thaliana},
partial (6%)
Length = 193
Score = 70.5 bits (171), Expect = 2e-12
Identities = 30/61 (49%), Positives = 40/61 (65%)
Frame = +1
Query: 466 HDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLAT 525
H +MSLDFHP++ D C DS + IR +N+ S C ++GG+ +VRFQP GQLLA
Sbjct: 1 HSSAIMSLDFHPKKTDFFCFCDSENEIRYYNITSSSCTRVSKGGNAKVRFQPGSGQLLAA 180
Query: 526 A 526
A
Sbjct: 181 A 183
>TC79118 similar to GP|16323188|gb|AAL15328.1 AT5g67320/K8K14_4 {Arabidopsis
thaliana}, partial (44%)
Length = 1098
Score = 61.2 bits (147), Expect = 1e-09
Identities = 54/205 (26%), Positives = 95/205 (46%), Gaps = 15/205 (7%)
Frame = +2
Query: 408 IWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDA--SNPIRSLGILTG 465
+W+++ + HS DV ++ +T FA+SS D + + ++P ++ G
Sbjct: 146 VWDVKAEECKQQFGFHSGPTLDVDWRT-NTSFASSSTDNMIYVCKIGENHPTQTFA---G 313
Query: 466 HDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRF---------- 515
H +V + + P LL S + ++W+V Q + +H R SK++
Sbjct: 314 HQGEVNCVKWDPTGT-LLASCSDDITAKIWSVKQDKYLHDFREHSKEIYTIRWSPTGPGT 490
Query: 516 -QPQYGQLLATA-IGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SA 572
P LLA+A +++ + D+E +H L GH V S+ + +G ++AS S D S
Sbjct: 491 NNPNKKLLLASASFDSTVKLWDIELGKLIHSLNGHRHPVYSVAFSPNGEYIASGSLDKSL 670
Query: 573 RIWSAASDGKCIGELHSVGNKFQSC 597
IWS +GK I + G F+ C
Sbjct: 671 HIWS-LKEGKIIRTYNGSGGIFEVC 742
Score = 55.1 bits (131), Expect = 9e-08
Identities = 45/180 (25%), Positives = 78/180 (43%), Gaps = 13/180 (7%)
Frame = +2
Query: 494 LWNVNQSECMHTT---RGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDK 550
+W+V EC G + V ++ +++ N I + + + GH
Sbjct: 146 VWDVKAEECKQQFGFHSGPTLDVDWRTNTS-FASSSTDNMIYVCKIGENHPTQTFAGHQG 322
Query: 551 DVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIF--------HP 601
+V + WD +G LAS S+D +A+IWS D K + + + + + +P
Sbjct: 323 EVNCVKWDPTGTLLASCSDDITAKIWSVKQD-KYLHDFREHSKEIYTIRWSPTGPGTNNP 499
Query: 602 GYRSLLIIGGYQS-LEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ LL + S ++ W G S+ H+ + +A SP GE IAS S D S+ +W
Sbjct: 500 NKKLLLASASFDSTVKLWDIELGKLIHSLNGHRHPVYSVAFSPNGEYIASGSLDKSLHIW 679
Score = 40.0 bits (92), Expect = 0.003
Identities = 17/56 (30%), Positives = 30/56 (53%)
Frame = +2
Query: 396 VMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLW 451
++ASA + V +W++E + S + H H + V F P A+ S D+S+ +W
Sbjct: 512 LLASASFDSTVKLWDIELGKLIHSLNGHRHPVYSVAFSPNGEYIASGSLDKSLHIW 679
>BG457241 homologue to PIR|F84600|F8 coatomer alpha subunit [imported] -
Arabidopsis thaliana, partial (15%)
Length = 654
Score = 58.9 bits (141), Expect = 6e-09
Identities = 30/109 (27%), Positives = 58/109 (52%)
Frame = +1
Query: 389 HFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSV 448
HF + + S G + K+ +WN + + + H I V+F S ++S D+++
Sbjct: 244 HFHNSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQFHHESPWIVSASDDQTI 423
Query: 449 RLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNV 497
R+W+ + + +LTGH+ VM FHP+ DL+ S+ + +R+W++
Sbjct: 424 RIWNWQSR-TCISVLTGHNHYVMCALFHPKD-DLVVSASLDQTVRVWDI 564
Score = 47.8 bits (112), Expect = 1e-05
Identities = 35/172 (20%), Positives = 72/172 (41%), Gaps = 14/172 (8%)
Frame = +1
Query: 379 HKSINKVLTCHFSSDGKVMASAGHEKKVFI-----------WNMENFQYFASADTHSHLI 427
H ++K+LT + +V + H K+ +I W+ D H +
Sbjct: 55 H*PLSKMLTKFETKSNRVKGLSFHPKRPWILASLHSGVIQLWDYRMGTLIDKFDEHDGPV 234
Query: 428 TDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSD 487
V F +F + D +++W+ R L L GH + + ++ FH ++ +SD
Sbjct: 235 RGVHFHNSQPLFVSGGDDYKIKVWNYKLH-RCLFTLLGHLDYIRTVQFHHESPWIVSASD 411
Query: 488 SNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDV 536
+ IR+WN C+ G + V F P+ +++ ++ ++ + D+
Sbjct: 412 -DQTIRIWNWQSRTCISVLTGHNHYVMCALFHPKDDLVVSASLDQTVRVWDI 564
Score = 34.3 bits (77), Expect = 0.16
Identities = 30/151 (19%), Positives = 60/151 (38%), Gaps = 1/151 (0%)
Frame = +1
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
K + F P+ +LA+ I + D + + HD V + + S S +D
Sbjct: 109 KGLSFHPKRPWILASLHSGVIQLWDYRMGTLIDKFDEHDGPVRGVHFHNSQPLFVSGGDD 288
Query: 571 -SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSI 629
++W+ +C+ L + ++ FH ++ Q++ W+ + +
Sbjct: 289 YKIKVWNYKLH-RCLFTLLGHLDYIRTVQFHHESPWIVSASDDQTIRIWNWQSRTCISVL 465
Query: 630 AAHQGLIAGLADSPVGELIASASHDNSVKVW 660
H + P +L+ SAS D +V+VW
Sbjct: 466 TGHNHYVMCALFHPKDDLVVSASLDQTVRVW 558
>TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein beta
subunit-like protein. [Alfalfa] {Medicago sativa},
complete
Length = 1261
Score = 56.6 bits (135), Expect = 3e-08
Identities = 55/236 (23%), Positives = 100/236 (42%), Gaps = 13/236 (5%)
Frame = +2
Query: 396 VMASAGHEKKVFIWNM--ENFQYFASADT---HSHLITDVRFQPGSTMFATSSFDRSVRL 450
++ +A +K + +W++ E+ Y HSH + DV + S+D +RL
Sbjct: 149 MIVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHFVQDVVLSSDGQFALSGSWDGELRL 328
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WD N S GH + V+S+ F ++ S+ + I+LWN EC +T + G
Sbjct: 329 WDL-NAGTSARRFVGHTKDVLSVAFSIDNRQIV-SASRDRTIKLWN-TLGECKYTIQDGD 499
Query: 511 KQ------VRFQPQYGQ--LLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGN 562
VRF P Q +++ + ++ + ++ + L GH V ++ G+
Sbjct: 500 AHSDWVSCVRFSPSTLQPTIVSASWDRTVKVWNLTNCKLRNTLAGHSGYVNTVAVSPDGS 679
Query: 563 FLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAW 618
AS +D + ++GK + L G+ + F P R L S++ W
Sbjct: 680 LCASGGKDGVILLWDLAEGKRLYSL-DAGSIIHALCFSPN-RYWLCAATESSIKIW 841
Score = 33.1 bits (74), Expect = 0.35
Identities = 51/244 (20%), Positives = 83/244 (33%), Gaps = 6/244 (2%)
Frame = +2
Query: 423 HSHLITDVRFQ-PGSTMFATSSFDRSVRLWDASNPIRSLGI----LTGHDEQVMSLDFHP 477
H+ ++T + S M T+S D+S+ LW + ++ G+ LTGH V +
Sbjct: 101 HTDVVTAIATPIDNSDMIVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHFVQDVVLSS 280
Query: 478 RRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVE 537
L S + +RLW++N G+ RF
Sbjct: 281 DGQFALSGSWDGE-LRLWDLN---------AGTSARRFV--------------------- 367
Query: 538 TDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQS 596
GH KDVLS+ + + S S D + ++W+ + K + + + S
Sbjct: 368 ---------GHTKDVLSVAFSIDNRQIVSASRDRTIKLWNTLGECKYTIQDGDAHSDWVS 520
Query: 597 CIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNS 656
C+ +SP+ T I SAS D +
Sbjct: 521 CV------------------RFSPSTLQPT---------------------IVSASWDRT 583
Query: 657 VKVW 660
VKVW
Sbjct: 584 VKVW 595
>TC92589 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis thaliana},
partial (5%)
Length = 415
Score = 56.6 bits (135), Expect = 3e-08
Identities = 29/48 (60%), Positives = 36/48 (74%)
Frame = +1
Query: 614 SLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
SLE W+ TE +KT +++AH GLIA LA S V L+ASASHD VK+WK
Sbjct: 10 SLELWNMTE-NKTMTLSAHDGLIAALAVSTVNGLVASASHDRFVKLWK 150
>TC88475 similar to GP|20260442|gb|AAM13119.1 unknown protein {Arabidopsis
thaliana}, partial (96%)
Length = 1315
Score = 55.5 bits (132), Expect = 7e-08
Identities = 59/286 (20%), Positives = 114/286 (39%), Gaps = 10/286 (3%)
Frame = +3
Query: 379 HKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTM 438
H I +T S + AS E++ N+ + +Y H + V + T
Sbjct: 87 HPRIRTTITVSVS---RKSASKAMEEQFSFKNLHSREYSG----HKKKVHSVAWNCIGTK 245
Query: 439 FATSSFDRSVRLWD----ASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRL 494
A+ S D++ R+W A ++ + L GH + V L + P+ DL+ ++ + +RL
Sbjct: 246 LASGSVDQTARIWHIDPHAHGKVKDIE-LKGHTDSVDQLCWDPKHPDLIATASGDKTVRL 422
Query: 495 WNVNQSECMHTTR--GGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDV 552
W+ +C G + + ++P + + +TI+DV +H K + +V
Sbjct: 423 WDARSGKCSQQAELSGENINITYKPDGTHVAVGNRDDELTILDVRKFKPMHRRK-FNYEV 599
Query: 553 LSICWDRSGN-FLASVSEDSARIWSAASDGKCIGELHSVGNKFQSC---IFHPGYRSLLI 608
I W+ +G F + + + S S + L ++ C P R +
Sbjct: 600 NEIAWNMTGEMFFLTTGNGTVEVLSYPS----LRPLDTLMAHTAGCYCIAIDPTGRHFAV 767
Query: 609 IGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHD 654
+ W +E + + + ++ + G+LIASAS D
Sbjct: 768 GSADSLVSLWVISEMLCVRTFTKLEWPVRTISFNHTGDLIASASED 905
>TC87005 similar to GP|9294061|dbj|BAB02018.1 WD40-repeat protein
{Arabidopsis thaliana}, partial (74%)
Length = 1480
Score = 51.6 bits (122), Expect = 1e-06
Identities = 65/297 (21%), Positives = 120/297 (39%), Gaps = 26/297 (8%)
Frame = +3
Query: 390 FSSDGKVMASAGHEKKVFIWNMEN-FQYFASADTHSHLITDVRFQPGSTMFATS------ 442
FS +G+ +ASA V IW N F S I D+++ P S
Sbjct: 231 FSPNGEWVASADVSGTVRIWGTRNEFVLKKEFRVLSGRIDDLQWSPDGIRIVASGEAKGN 410
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
SF R+ +WD+ ++G GH +V+S + P R + + + ++ +
Sbjct: 411 SFVRAF-MWDSGT---NVGEFDGHSRRVLSCAYKPTRPFRIVTCGEDFLVNFYEGPPFRF 578
Query: 503 MHTTRGGS---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLK---GHDKDVLSIC 556
+ R S VR+ P + + + I D +T + +L GH + ++
Sbjct: 579 KQSHRDHSNFVNCVRYSPDGSKFVTGSSDKKGIIFDGKTGEKIGELSSEGGHTGSIYAVS 758
Query: 557 WDRSGNFLASVSED-SARIWSAASDGKCIGELH---------SVGNKFQSCIFHPGYRSL 606
W G + +VS D SA++W + D IG++ V + C++ Y
Sbjct: 759 WSPDGKQVLTVSADKSAKVWDISEDN--IGKVKKTLICSASGGVEDMLVGCLWLNDYLVT 932
Query: 607 LIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLA---DSPVGELIASASHDNSVKVW 660
+ +GG S+ + S + + S + H ++ L +P ++ S S+D + W
Sbjct: 933 VSLGGTISIFSASDLDKAPR-SFSGHMKNVSSLTIRRSNP--RVLLSCSYDGLIVKW 1094
Score = 35.4 bits (80), Expect = 0.071
Identities = 44/226 (19%), Positives = 91/226 (39%), Gaps = 11/226 (4%)
Frame = +3
Query: 446 RSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM-- 503
RSV + + NP++ + + H F P + + S+D + +R+W +
Sbjct: 147 RSVIMMNLHNPLQ-VSVYGEHAYPATVARFSPNG-EWVASADVSGTVRIWGTRNEFVLKK 320
Query: 504 --HTTRGGSKQVRFQPQYGQLLAT--AIGNS-ITIIDVETDSHLHDLKGHDKDVLSICWD 558
G +++ P +++A+ A GNS + ++ +++ + GH + VLS +
Sbjct: 321 EFRVLSGRIDDLQWSPDGIRIVASGEAKGNSFVRAFMWDSGTNVGEFDGHSRRVLSCAYK 500
Query: 559 RSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEA- 617
+ F + + + H + F +C+ + S + G
Sbjct: 501 PTRPFRIVTCGEDFLVNFYEGPPFRFKQSHRDHSNFVNCVRYSPDGSKFVTGSSDKKGII 680
Query: 618 WSPTEGSKTWSIAA---HQGLIAGLADSPVGELIASASHDNSVKVW 660
+ G K +++ H G I ++ SP G+ + + S D S KVW
Sbjct: 681 FDGKTGEKIGELSSEGGHTGSIYAVSWSPDGKQVLTVSADKSAKVW 818
>TC82282 similar to PIR|T08544|T08544 hypothetical protein F27B13.70 -
Arabidopsis thaliana, partial (84%)
Length = 904
Score = 50.4 bits (119), Expect = 2e-06
Identities = 43/212 (20%), Positives = 87/212 (40%), Gaps = 16/212 (7%)
Frame = +2
Query: 403 EKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGI 462
++ V +W ++ + H + V P ++ A+SS D VR++D + ++
Sbjct: 191 DETVRLWKSDDLVLERTNTGHCLGVASVAAHPLGSIAASSSLDSFVRVFDVDSNA-TIAT 367
Query: 463 LTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTT------------RGGS 510
L +V + F P+ L + + + LW+ + E + T + GS
Sbjct: 368 LEAPPSEVWQMRFDPKGAILAVAGGGSASVNLWDTSTWELVVTLSIPRVEGPKPSDKSGS 547
Query: 511 KQ----VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
K+ V + P +L ++ +I++ DV+ LH L+GH V S+ + L
Sbjct: 548 KKFVLSVAWSPDGKRLACGSMDGTISVFDVQRAKFLHHLEGHFMPVRSLVYSPYDPRLLF 727
Query: 567 VSEDSARIWSAASDGKCIGELHSVGNKFQSCI 598
+ D + ++GK + S + C+
Sbjct: 728 SASDDGNVHMYDAEGKALXGTMSXHASWVLCV 823
>TC80645 similar to PIR|T50983|T50983 probable pleiotropic regulator 1
(PLRG1) [imported] - Neurospora crassa, partial (41%)
Length = 1141
Score = 50.1 bits (118), Expect = 3e-06
Identities = 44/194 (22%), Positives = 78/194 (39%), Gaps = 11/194 (5%)
Frame = +2
Query: 408 IWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHD 467
+W+M H+ ++D+ Q T S D +VRLWD + +S+G+LT H
Sbjct: 32 VWDMRTRSNVHVLSGHTGTVSDLTCQEADPQVITGSLDATVRLWDLAAG-KSMGVLTHHK 208
Query: 468 EQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQP--QYGQLLAT 525
+ V +L HP + +S S+ I+ W + M G + + + L +
Sbjct: 209 KGVRALATHPE--EFTFASGSSGSIKQWKCPEGAFMQNFEGHNAIINTLSVNRDNVLFSG 382
Query: 526 AIGNSITIIDVETDSHLHDLK--------GHDKDVLSICWDRSG-NFLASVSEDSARIWS 576
S+ D +T L+ + ++S +D SG + ++ + +IW
Sbjct: 383 GDNGSMNFWDWKTGHRFQSLETTAQPGSLDAEAGIMSSTFDNSGLRLICGEADKTIKIWK 562
Query: 577 AASDGKCIGELHSV 590
D K E H V
Sbjct: 563 --QDEKATPETHPV 598
>TC77248 similar to GP|13489182|gb|AAK27816.1 putative WD-repeat containing
protein {Oryza sativa}, partial (92%)
Length = 2005
Score = 50.1 bits (118), Expect = 3e-06
Identities = 46/199 (23%), Positives = 88/199 (44%), Gaps = 8/199 (4%)
Frame = +1
Query: 470 VMSLDF-HPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLAT 525
+++LD H + + + D+N VI ++ + + T G SK+V +F Q ++
Sbjct: 769 IIALDILHSKDLIATGNIDTNAVI--FDRPSGQVLATLTGHSKKVTSVKFVGQGESIITG 942
Query: 526 AIGNSITIIDVETDSHLHD---LKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGK 582
+ ++ + D H + LK H +V ++ + N+ + S D + + S G
Sbjct: 943 SADKTVRLWQGSDDGHYNCKQILKDHSAEVEAVTVHATNNYFVTASLDGSWCFYELSSGT 1122
Query: 583 CIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSL-EAWSPTEGSKTWSIAAHQGLIAGLAD 641
C+ ++ + S FHP +L G SL + W + H G + ++
Sbjct: 1123CLTQVSDSSEGYTSAAFHPD-GLILGTGTTDSLVKIWDVKSQANVAKFDGHVGHVTAISF 1299
Query: 642 SPVGELIASASHDNSVKVW 660
S G +A+A+HD VK+W
Sbjct: 1300SENGYYLATAAHD-GVKLW 1353
Score = 29.6 bits (65), Expect = 3.9
Identities = 16/72 (22%), Positives = 34/72 (47%)
Frame = +1
Query: 428 TDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSD 487
T F P + T + D V++WD + ++ GH V ++ F L ++
Sbjct: 1159 TSAAFHPDGLILGTGTTDSLVKIWDVKSQ-ANVAKFDGHVGHVTAISFSENGYYL--ATA 1329
Query: 488 SNDVIRLWNVNQ 499
++D ++LW++ +
Sbjct: 1330 AHDGVKLWDLRK 1365
>TC78016 similar to GP|21537191|gb|AAM61532.1 PRL1 protein {Arabidopsis
thaliana}, partial (81%)
Length = 1759
Score = 50.1 bits (118), Expect = 3e-06
Identities = 26/86 (30%), Positives = 45/86 (52%)
Frame = +1
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H + V P +T FAT S DR++++WD ++ + L LTGH EQV L + +
Sbjct: 583 HLGWVRSVAVDPSNTWFATGSADRTIKIWDLASGVLKL-TLTGHIEQVRGLAISHKHTYM 759
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRG 508
+ D V + W++ Q++ + + G
Sbjct: 760 FSAGDDKQV-KCWDLEQNKVIRSYHG 834
Score = 43.1 bits (100), Expect = 3e-04
Identities = 28/125 (22%), Positives = 55/125 (43%)
Frame = +3
Query: 389 HFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSV 448
HF+ V+ SAG +W++ + + H + + V +P T S D ++
Sbjct: 879 HFTDWEDVILSAG------VWDIRSKMQIHALSGHENTVCSVFTRPTDPQVVTGSHDSTI 1040
Query: 449 RLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRG 508
++WD +++ LT H + V ++ HP+ +S S D I+ + + + E H
Sbjct: 1041KMWDLRYG-KTMSTLTNHKKSVRAMAQHPKEQAF--ASASADNIKKFTLPKGEFCHNMLS 1211
Query: 509 GSKQV 513
K +
Sbjct: 1212QQKTI 1226
Score = 28.9 bits (63), Expect = 6.6
Identities = 9/47 (19%), Positives = 22/47 (46%)
Frame = +3
Query: 614 SLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
S W + +++ H+ + + P + + SHD+++K+W
Sbjct: 909 SAGVWDIRSKMQIHALSGHENTVCSVFTRPTDPQVVTGSHDSTIKMW 1049
>TC79260 weakly similar to GP|9759350|dbj|BAB10005.1 WD-repeat protein-like
{Arabidopsis thaliana}, partial (50%)
Length = 1227
Score = 48.9 bits (115), Expect = 6e-06
Identities = 44/144 (30%), Positives = 69/144 (47%), Gaps = 6/144 (4%)
Frame = +2
Query: 520 GQLLATAIGNSITII-DVETDSHL---HDLKGHDKDVLSICWDRSGNFLASVS-EDSARI 574
G+ LATA + II +V+T+ L H L GH K V S+ W +G L + E++ R
Sbjct: 581 GKYLATASNDRTAIIWEVDTNDGLSMKHKLSGHQKSVSSVSWSPNGQELLTCGVEEAVRR 760
Query: 575 WSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQG 634
W S GKC+ G+ SC + P + +L +S+ W +G + S +
Sbjct: 761 WD-VSTGKCLQVYEKNGSGLISCAWFPCGKYILSGLSDKSICMWE-LDGKEVESWKGQKT 934
Query: 635 L-IAGLADSPVGELIASASHDNSV 657
L I+ L + GE I S +N++
Sbjct: 935 LKISDLEITGDGEHILSICKENAI 1006
Score = 43.5 bits (101), Expect = 3e-04
Identities = 43/215 (20%), Positives = 88/215 (40%), Gaps = 8/215 (3%)
Frame = +2
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFA---SADTHSHLITDVRFQPGSTMF 439
++V FS DGK +A+A +++ IW ++ + H ++ V + P
Sbjct: 548 DEVWFVQFSHDGKYLATASNDRTAIIWEVDTNDGLSMKHKLSGHQKSVSSVSWSPNGQEL 727
Query: 440 ATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQ 499
T + +VR WD S + L + + ++S + P +L S S+ I +W ++
Sbjct: 728 LTCGVEEAVRRWDVSTG-KCLQVYEKNGSGLISCAWFPCGKYIL-SGLSDKSICMWELDG 901
Query: 500 SECMHTTRGGS----KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSI 555
E + + +G + +L+ N+I + + ET + D+ + S
Sbjct: 902 KE-VESWKGQKTLKISDLEITGDGEHILSICKENAILLFNKET--KVERFIEEDQTITSF 1072
Query: 556 CWDRSGNF-LASVSEDSARIWSAASDGKCIGELHS 589
+ F L ++ +W+ D K +G+ S
Sbjct: 1073SLSKDSRFLLVNLLNQEIHLWNIEGDLKLVGKYKS 1177
Score = 42.0 bits (97), Expect = 8e-04
Identities = 23/85 (27%), Positives = 43/85 (50%), Gaps = 2/85 (2%)
Frame = +2
Query: 421 DTHSHLITDVRFQPGSTMFATSSFDRSVRLW--DASNPIRSLGILTGHDEQVMSLDFHPR 478
D H + V+F AT+S DR+ +W D ++ + L+GH + V S+ + P
Sbjct: 536 DAHDDEVWFVQFSHDGKYLATASNDRTAIIWEVDTNDGLSMKHKLSGHQKSVSSVSWSPN 715
Query: 479 RMDLLCSSDSNDVIRLWNVNQSECM 503
+LL + + +R W+V+ +C+
Sbjct: 716 GQELL-TCGVEEAVRRWDVSTGKCL 787
>TC79342 similar to GP|10177096|dbj|BAB10430. Notchless protein homolog
{Arabidopsis thaliana}, partial (47%)
Length = 873
Score = 47.8 bits (112), Expect = 1e-05
Identities = 26/123 (21%), Positives = 53/123 (42%), Gaps = 4/123 (3%)
Frame = +3
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
VL+ FS DG+ +AS + V W++ + H + + + + P + S
Sbjct: 504 VLSVAFSPDGRQLASGSGDTTVRFWDLGTQTPMYTCTGHKNWVLCIAWSPDGKYLVSGSM 683
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLC----SSDSNDVIRLWNVNQS 500
+ WD + LTGH + + + + P ++ C S+ + R+W+V+
Sbjct: 684 SGELICWDPQTGKQLGNALTGHKKWITGISWEPVHLNAPCRRFVSASKDGDARIWDVSLK 863
Query: 501 ECM 503
+C+
Sbjct: 864 KCI 872
Score = 40.4 bits (93), Expect = 0.002
Identities = 43/173 (24%), Positives = 64/173 (36%), Gaps = 6/173 (3%)
Frame = +3
Query: 418 ASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHP 477
A+ H + V F P A+ S D +VR WD + TGH V+ + + P
Sbjct: 477 ATISGHGEAVLSVAFSPDGRQLASGSGDTTVRFWDLGTQ-TPMYTCTGHKNWVLCIAWSP 653
Query: 478 RRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVE 537
L+ S S ++I W+ PQ G+ L GN++T
Sbjct: 654 DGKYLVSGSMSGELI-CWD--------------------PQTGKQL----GNALT----- 743
Query: 538 TDSHLHDLKGHDKDVLSICWD------RSGNFLASVSEDSARIWSAASDGKCI 584
GH K + I W+ F+++ + ARIW S KCI
Sbjct: 744 ---------GHKKWITGISWEPVHLNAPCRRFVSASKDGDARIWD-VSLKKCI 872
Score = 36.2 bits (82), Expect = 0.041
Identities = 29/111 (26%), Positives = 47/111 (42%), Gaps = 3/111 (2%)
Frame = +3
Query: 478 RRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSK---QVRFQPQYGQLLATAIGNSITII 534
+ + ++C + V R+ VN+ C T G + V F P QL + + ++
Sbjct: 411 KALPIVCQPQA--VFRIRPVNR--CSATISGHGEAVLSVAFSPDGRQLASGSGDTTVRFW 578
Query: 535 DVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIG 585
D+ T + ++ GH VL I W G +L S S I GK +G
Sbjct: 579 DLGTQTPMYTCTGHKNWVLCIAWSPDGKYLVSGSMSGELICWDPQTGKQLG 731
>TC84162 similar to GP|15810485|gb|AAL07130.1 putative WD40-repeat protein
{Arabidopsis thaliana}, partial (29%)
Length = 628
Score = 47.0 bits (110), Expect = 2e-05
Identities = 33/149 (22%), Positives = 62/149 (41%)
Frame = +1
Query: 396 VMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASN 455
++ S ++ +W + + H I V F P T+S D+++R+W S+
Sbjct: 28 LVCSGSQDRTACVWRLPDLVSVVVFKGHKRGIWSVEFSPVDQCVVTASGDKTIRIWAISD 207
Query: 456 PIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRF 515
L GH V+ F R ++ S ++ +++LW V +EC+ T +V
Sbjct: 208 G-SCLKTFEGHTSSVLRALFVTRGTQIV-SCGADGLVKLWTVKSNECVATYDNHEDKVWA 381
Query: 516 QPQYGQLLATAIGNSITIIDVETDSHLHD 544
+ A G S ++++ DS D
Sbjct: 382 LAVGRKTETLATGGSDAVVNLWLDSTAAD 468
Score = 32.7 bits (73), Expect = 0.46
Identities = 25/116 (21%), Positives = 45/116 (38%), Gaps = 1/116 (0%)
Frame = +1
Query: 546 KGHDKDVLSICWDRSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYR 604
KGH + + S+ + + + S D RIW A SDG C+ + +F
Sbjct: 103 KGHKRGIWSVEFSPVDQCVVTASGDKTIRIW-AISDGSCLKTFEGHTSSVLRALFVTRGT 279
Query: 605 SLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ G ++ W+ + H+ + LA E +A+ D V +W
Sbjct: 280 QIVSCGADGLVKLWTVKSNECVATYDNHEDKVWALAVGRKTETLATGGSDAVVNLW 447
Score = 29.6 bits (65), Expect = 3.9
Identities = 17/69 (24%), Positives = 30/69 (42%)
Frame = +1
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
+ VL F + G + S G + V +W +++ + A+ D H + + + AT
Sbjct: 241 SSVLRALFVTRGTQIVSCGADGLVKLWTVKSNECVATYDNHEDKVWALAVGRKTETLATG 420
Query: 443 SFDRSVRLW 451
D V LW
Sbjct: 421 GSDAVVNLW 447
Score = 29.3 bits (64), Expect = 5.1
Identities = 27/108 (25%), Positives = 46/108 (42%), Gaps = 5/108 (4%)
Frame = +1
Query: 482 LLCSSDSNDVIRLWNVNQSECMHTTRG---GSKQVRFQPQYGQLLATAIGN-SITIIDVE 537
L+CS + +W + + +G G V F P Q + TA G+ +I I +
Sbjct: 28 LVCSGSQDRTACVWRLPDLVSVVVFKGHKRGIWSVEFSP-VDQCVVTASGDKTIRIWAIS 204
Query: 538 TDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCI 584
S L +GH VL + G + S D ++W+ S+ +C+
Sbjct: 205 DGSCLKTFEGHTSSVLRALFVTRGTQIVSCGADGLVKLWTVKSN-ECV 345
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.130 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,435,747
Number of Sequences: 36976
Number of extensions: 321862
Number of successful extensions: 1928
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 1730
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1864
length of query: 661
length of database: 9,014,727
effective HSP length: 102
effective length of query: 559
effective length of database: 5,243,175
effective search space: 2930934825
effective search space used: 2930934825
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0114.1


