

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.7
(147 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
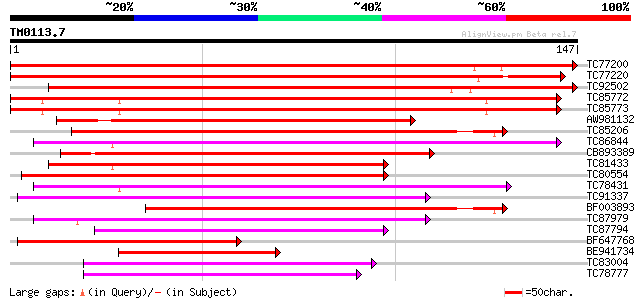
Score E
Sequences producing significant alignments: (bits) Value
TC77200 similar to GP|22597162|gb|AAN03468.1 bZIP transcription ... 209 3e-55
TC77220 similar to GP|22597162|gb|AAN03468.1 bZIP transcription ... 192 5e-50
TC92502 weakly similar to GP|22597162|gb|AAN03468.1 bZIP transcr... 143 2e-35
TC85772 weakly similar to GP|9650828|emb|CAC00658.1 common plant... 133 2e-32
TC85773 weakly similar to GP|9650828|emb|CAC00658.1 common plant... 133 2e-32
AW981132 similar to GP|4457221|gb|A putative bZIP DNA-binding pr... 105 9e-24
TC85206 similar to GP|16580134|gb|AAK92215.1 bZIP transcription ... 89 8e-19
TC86844 similar to GP|16797791|gb|AAL27150.1 bZIP transcription ... 83 4e-17
CB893389 similar to PIR|T48034|T480 bZIP transcription factor-li... 79 5e-16
TC81433 similar to GP|13430400|gb|AAK25822.1 bZip transcription ... 79 9e-16
TC80554 similar to GP|13430400|gb|AAK25822.1 bZip transcription ... 77 3e-15
TC78431 similar to GP|16797791|gb|AAL27150.1 bZIP transcription ... 77 3e-15
TC91337 weakly similar to GP|13365772|dbj|BAB39174. RISBZ4 {Oryz... 70 4e-13
BF003893 weakly similar to GP|14289165|dbj glip19 {Oryza sativa}... 67 3e-12
TC87979 weakly similar to GP|10954097|gb|AAG25728.1 bZIP protein... 65 8e-12
TC87794 similar to GP|13775107|gb|AAK39130.1 bZIP transcription ... 56 5e-09
BF647768 weakly similar to PIR|T03990|T039 probable transcriptio... 52 1e-07
BE941734 similar to GP|7406677|emb| putative ripening-related bZ... 51 2e-07
TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - ki... 50 3e-07
TC78777 similar to PIR|T10985|T10985 regulator protein ROM2 - ki... 49 7e-07
>TC77200 similar to GP|22597162|gb|AAN03468.1 bZIP transcription factor ATB2
{Glycine max}, partial (84%)
Length = 1481
Score = 209 bits (532), Expect = 3e-55
Identities = 109/156 (69%), Positives = 136/156 (86%), Gaps = 9/156 (5%)
Frame = +1
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
MASSSGTSSGSS++QNSGSEE+LQALMDQRKRKRMISNRESARRSRMRKQKHLD+L +QV
Sbjct: 532 MASSSGTSSGSSLLQNSGSEEDLQALMDQRKRKRMISNRESARRSRMRKQKHLDDLVSQV 711
Query: 61 AQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVF- 119
++LR ENQ++LTS+N+TTQ+++ ++AENSVL AQMGELS+RLESLNEI+ +N++NGVF
Sbjct: 712 SKLRKENQEILTSVNITTQKYLSVEAENSVLRAQMGELSNRLESLNEIVGALNSSNGVFG 891
Query: 120 AANSFVD--------PIPSAYYLNHPIMASADILQY 147
A+N+FV+ + + Y+N PIMASADILQY
Sbjct: 892 ASNAFVEQNNGFFFNSLNNMSYMNQPIMASADILQY 999
>TC77220 similar to GP|22597162|gb|AAN03468.1 bZIP transcription factor ATB2
{Glycine max}, partial (65%)
Length = 1341
Score = 192 bits (487), Expect = 5e-50
Identities = 103/151 (68%), Positives = 123/151 (81%), Gaps = 7/151 (4%)
Frame = +3
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
MASSSGTSSG S + NSGSEE+L LMDQRKRKRMISNRESARRSRMRKQKHLD+LA Q+
Sbjct: 561 MASSSGTSSGYSTLPNSGSEEDLMLLMDQRKRKRMISNRESARRSRMRKQKHLDDLAVQL 740
Query: 61 AQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVFA 120
+QLR+ENQQ+LTS+NLTTQRF+ +++ENSVL AQ+ EL+ R ESLNEII+F+N NGVF
Sbjct: 741 SQLRNENQQILTSVNLTTQRFLAVESENSVLRAQLNELNSRFESLNEIINFMNVANGVFE 920
Query: 121 -------ANSFVDPIPSAYYLNHPIMASADI 144
N F +P+ + YLN PIMASAD+
Sbjct: 921 PVDNNINENYFNNPL-NMGYLNQPIMASADM 1010
>TC92502 weakly similar to GP|22597162|gb|AAN03468.1 bZIP transcription
factor ATB2 {Glycine max}, partial (59%)
Length = 597
Score = 143 bits (361), Expect = 2e-35
Identities = 79/150 (52%), Positives = 106/150 (70%), Gaps = 13/150 (8%)
Frame = +3
Query: 11 SSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQM 70
S+ Q+ GSEE+LQ LMDQRKRKR S RESARRSRMRKQKH+D+L A+V +LR+EN ++
Sbjct: 15 STESQSYGSEEDLQRLMDQRKRKRKQSGRESARRSRMRKQKHMDDLIAEVERLRNENSEI 194
Query: 71 LTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVN---ATNGV--------- 118
LT +N+TTQ ++ I+AEN VL AQM EL+ RL+SLN+II+ +N TNGV
Sbjct: 195 LTRMNMTTQHYLKIEAENCVLRAQMCELNQRLQSLNDIINLINITTTTNGVNYQNNDCFL 374
Query: 119 -FAANSFVDPIPSAYYLNHPIMASADILQY 147
+ N F++ + +Y PIMASAD+ +
Sbjct: 375 TISDNCFMNQMNMSYLNQQPIMASADMFMW 464
>TC85772 weakly similar to GP|9650828|emb|CAC00658.1 common plant regulatory
factor 7 {Petroselinum crispum}, partial (64%)
Length = 1224
Score = 133 bits (335), Expect = 2e-32
Identities = 74/155 (47%), Positives = 107/155 (68%), Gaps = 12/155 (7%)
Frame = +1
Query: 1 MASSSGT-SSGSSMIQNSGSEEELQALM----DQRKRKRMISNRESARRSRMRKQKHLDE 55
MASS GT SSG+S +QNSGSE + + DQ+KRKRM SNRESARRSRM+KQ+H+++
Sbjct: 655 MASSGGTYSSGTSSLQNSGSEGDHNHVQVNITDQKKRKRMQSNRESARRSRMKKQQHMED 834
Query: 56 LAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNAT 115
L+ Q+ QL+ EN Q+ T++ +TTQ ++ +++EN++L QM ELSHRL+SLN+II ++ ++
Sbjct: 835 LSNQIEQLKKENIQISTNVGVTTQMYLNVESENAILRVQMAELSHRLQSLNDIIHYIESS 1014
Query: 116 NGVFAAN-------SFVDPIPSAYYLNHPIMASAD 143
N +F F D + PIMAS+D
Sbjct: 1015NSLFQETDQLFNDCGFSDTWNTFPVNQQPIMASSD 1119
>TC85773 weakly similar to GP|9650828|emb|CAC00658.1 common plant regulatory
factor 7 {Petroselinum crispum}, partial (62%)
Length = 945
Score = 133 bits (335), Expect = 2e-32
Identities = 74/155 (47%), Positives = 107/155 (68%), Gaps = 12/155 (7%)
Frame = +1
Query: 1 MASSSGT-SSGSSMIQNSGSEEELQALM----DQRKRKRMISNRESARRSRMRKQKHLDE 55
MASS GT SSG+S +QNSGSE + + DQ+KRKRM SNRESARRSRM+KQ+H+++
Sbjct: 166 MASSGGTYSSGTSSLQNSGSEGDHNHVQVNITDQKKRKRMQSNRESARRSRMKKQQHMED 345
Query: 56 LAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNAT 115
L+ Q+ QL+ EN Q+ T++ +TTQ ++ +++EN++L QM ELSHRL+SLN+II ++ ++
Sbjct: 346 LSNQIEQLKKENIQISTNVGVTTQMYLNVESENAILRVQMAELSHRLQSLNDIIHYIESS 525
Query: 116 NGVFAAN-------SFVDPIPSAYYLNHPIMASAD 143
N +F F D + PIMAS+D
Sbjct: 526 NSLFQETDQLFNDCGFSDTWNTFPVNQQPIMASSD 630
>AW981132 similar to GP|4457221|gb|A putative bZIP DNA-binding protein
{Capsicum chinense}, partial (47%)
Length = 587
Score = 105 bits (261), Expect = 9e-24
Identities = 50/93 (53%), Positives = 74/93 (78%)
Frame = +2
Query: 13 MIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLT 72
++++SG E+EL MDQRK KR SNRESA+R RMRK KH+D++ +Q++QL +N ++L
Sbjct: 227 ILESSGPEKEL---MDQRKNKRKQSNRESAKRCRMRKHKHVDDMMSQMSQLTKDNSEILN 397
Query: 73 SLNLTTQRFMIIDAENSVLSAQMGELSHRLESL 105
S+N+TTQ ++ ++AENS+L AQ+GELS R +SL
Sbjct: 398 SINITTQHYLNVEAENSILRAQIGELSQRFQSL 496
>TC85206 similar to GP|16580134|gb|AAK92215.1 bZIP transcription factor
BZI-4 {Nicotiana tabacum}, partial (31%)
Length = 1139
Score = 88.6 bits (218), Expect = 8e-19
Identities = 47/114 (41%), Positives = 72/114 (62%), Gaps = 1/114 (0%)
Frame = +2
Query: 17 SGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNL 76
SG + + +D+RKRKRM+SNRESARRSR+RKQ+ +++L + +L+ EN ++ S+
Sbjct: 503 SGGSDGMDLQIDERKRKRMLSNRESARRSRLRKQQQVEDLTGEAGKLKIENDRLARSIKA 682
Query: 77 TTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVFAANSF-VDPIP 129
T + ++ ++A N V+ AQ EL + LN +ID A ANSF VD +P
Sbjct: 683 TEEAYLKMEAANDVIRAQTRELEAQFRFLNSVIDAAAAEE----ANSFSVDDVP 832
>TC86844 similar to GP|16797791|gb|AAL27150.1 bZIP transcription factor
{Nicotiana tabacum}, partial (47%)
Length = 1609
Score = 83.2 bits (204), Expect = 4e-17
Identities = 52/139 (37%), Positives = 75/139 (53%), Gaps = 2/139 (1%)
Frame = +1
Query: 7 TSSGSSMIQNSGSEEELQA--LMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLR 64
T+SGSS + E + D ++ +RM+SNRESARRSR RKQ HL EL QV++LR
Sbjct: 598 TTSGSSDDEEGDGEINMNGDNPTDAKRVRRMLSNRESARRSRRRKQAHLTELETQVSELR 777
Query: 65 SENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVFAANSF 124
EN +L L TQ+F +N +L A + L +++ E + +N VF A S
Sbjct: 778 GENSSLLKRLTDVTQKFNNSAVDNRILKADVETLRAKVKMAEETVKRFTGSNPVFNAMSE 957
Query: 125 VDPIPSAYYLNHPIMASAD 143
V + + + P +SAD
Sbjct: 958 VSSMGMSLFDGSPSESSAD 1014
>CB893389 similar to PIR|T48034|T480 bZIP transcription factor-like protein -
Arabidopsis thaliana, partial (35%)
Length = 724
Score = 79.3 bits (194), Expect = 5e-16
Identities = 44/97 (45%), Positives = 66/97 (67%)
Frame = +1
Query: 14 IQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTS 73
+ +SGS+ + MD+RKRKRMISNRESARRSR RKQK L++ + +LR+EN+++ +
Sbjct: 163 VTSSGSDG-VNGAMDERKRKRMISNRESARRSRERKQKLLEDYQDEANRLRNENRRLSEN 339
Query: 74 LNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIID 110
+ + + F +A N VL AQ EL+ +L+ L II+
Sbjct: 340 IRVREEGFNANEAANGVLRAQTQELTDQLKFLKSIIE 450
>TC81433 similar to GP|13430400|gb|AAK25822.1 bZip transcription factor
{Phaseolus vulgaris}, partial (68%)
Length = 774
Score = 78.6 bits (192), Expect = 9e-16
Identities = 44/89 (49%), Positives = 60/89 (66%), Gaps = 1/89 (1%)
Frame = +2
Query: 11 SSMIQNSGSEEELQA-LMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQ 69
SS S +ELQ ++D+RK +RMISNRESARRSRMRKQKHLDEL +QV +LR+EN
Sbjct: 215 SSNSTTSDEADELQFNIIDERKHRRMISNRESARRSRMRKQKHLDELWSQVMKLRTENHN 394
Query: 70 MLTSLNLTTQRFMIIDAENSVLSAQMGEL 98
++ LN ++ + EN+ L + +L
Sbjct: 395 LVDKLNHVSESHDTVVQENARLKEETFDL 481
>TC80554 similar to GP|13430400|gb|AAK25822.1 bZip transcription factor
{Phaseolus vulgaris}, partial (52%)
Length = 839
Score = 77.0 bits (188), Expect = 3e-15
Identities = 45/95 (47%), Positives = 60/95 (62%)
Frame = +1
Query: 4 SSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQL 63
S +S SS + ++E+ L+++RK +RM+SNRESARRSRMRKQK LDEL +QV L
Sbjct: 229 SPPSSCISSNSTSDEADEQNLGLINERKHRRMVSNRESARRSRMRKQKQLDELWSQVVWL 408
Query: 64 RSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGEL 98
R+EN Q+L LN + + EN L Q EL
Sbjct: 409 RNENHQLLDKLNNFCETHDKVVQENVQLKEQASEL 513
>TC78431 similar to GP|16797791|gb|AAL27150.1 bZIP transcription factor
{Nicotiana tabacum}, partial (61%)
Length = 1546
Score = 76.6 bits (187), Expect = 3e-15
Identities = 47/131 (35%), Positives = 70/131 (52%), Gaps = 7/131 (5%)
Frame = +1
Query: 7 TSSGSSMIQNSGSEEELQALM-------DQRKRKRMISNRESARRSRMRKQKHLDELAAQ 59
++SGSS Q+ E E + M D ++ +RM+SNRESARRSR RKQ HL EL Q
Sbjct: 661 STSGSSREQSDDDEAEGETYMIDSTDPTDVKRVRRMLSNRESARRSRRRKQAHLTELETQ 840
Query: 60 VAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVF 119
V+QLR EN ++ L Q++ +N VL A + L +++ E + V N +F
Sbjct: 841 VSQLRGENSTLVKRLTDVNQKYTDSAVDNRVLKADVETLRAKVKMAEETVKRVTGLNPLF 1020
Query: 120 AANSFVDPIPS 130
+ +P+
Sbjct: 1021HVMPDMSSMPA 1053
>TC91337 weakly similar to GP|13365772|dbj|BAB39174. RISBZ4 {Oryza sativa},
partial (38%)
Length = 1112
Score = 69.7 bits (169), Expect = 4e-13
Identities = 39/107 (36%), Positives = 62/107 (57%)
Frame = +1
Query: 3 SSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQ 62
++SG+S S SG E++ +D ++++R SN ESARRSR RKQ HL EL AQV +
Sbjct: 337 TASGSSDPSDEDNESGPCEQITNPVDMKRQRRKDSNCESARRSRWRKQAHLSELEAQVEK 516
Query: 63 LRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEII 109
L+ EN + T+Q+F D N VL + + L +++ +++
Sbjct: 517 LKLENATLYKQFTDTSQQFHEADTNNRVLKSDVEALRAKVKLAEDMV 657
>BF003893 weakly similar to GP|14289165|dbj glip19 {Oryza sativa}, partial
(20%)
Length = 533
Score = 66.6 bits (161), Expect = 3e-12
Identities = 36/95 (37%), Positives = 58/95 (60%), Gaps = 1/95 (1%)
Frame = -2
Query: 36 ISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQM 95
++ RESARRSR+RKQ+ +++L + +L+ EN ++ S+ T + ++ ++A N V+ AQ
Sbjct: 490 LNQRESARRSRLRKQQQVEDLTGEAGKLKIENDRLARSIKATEEAYLKMEAANDVIRAQT 311
Query: 96 GELSHRLESLNEIIDFVNATNGVFAANSF-VDPIP 129
EL + LN +ID A ANSF VD +P
Sbjct: 310 RELEAQFRFLNSVIDAAAAEE----ANSFSVDDVP 218
>TC87979 weakly similar to GP|10954097|gb|AAG25728.1 bZIP protein BZO2H2
{Arabidopsis thaliana}, partial (49%)
Length = 1346
Score = 65.5 bits (158), Expect = 8e-12
Identities = 39/108 (36%), Positives = 61/108 (56%), Gaps = 5/108 (4%)
Frame = +2
Query: 7 TSSGSSMIQN-----SGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVA 61
T+SGSS + +G E+ +D ++ +R +SNRESARRSR RKQ HL +L QV
Sbjct: 482 TTSGSSRDPSDEDDEAGPCEQSTNPVDMKRLRRKVSNRESARRSRRRKQAHLADLEVQVE 661
Query: 62 QLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEII 109
QLR EN + L +Q+F + N VL + + L +++ +++
Sbjct: 662 QLRLENASLFKQLTDASQQFRDANTNNRVLKSDVEALRAKVKLAEDMV 805
>TC87794 similar to GP|13775107|gb|AAK39130.1 bZIP transcription factor 2
{Phaseolus vulgaris}, partial (50%)
Length = 1264
Score = 56.2 bits (134), Expect = 5e-09
Identities = 30/76 (39%), Positives = 45/76 (58%)
Frame = +2
Query: 23 LQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFM 82
LQ + ++++R SNRESARRSR+RKQ DELA + L EN + L+ +
Sbjct: 368 LQDERELKRQRRKQSNRESARRSRLRKQAECDELAQRAEVLNQENASLRAELSPIKSEYE 547
Query: 83 IIDAENSVLSAQMGEL 98
I +EN+ L ++GE+
Sbjct: 548 KIPSENASLKERLGEI 595
>BF647768 weakly similar to PIR|T03990|T039 probable transcription
activator - rice, partial (29%)
Length = 660
Score = 51.6 bits (122), Expect = 1e-07
Identities = 28/58 (48%), Positives = 38/58 (65%)
Frame = +1
Query: 3 SSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
++SG+S S SG E++ +D ++++R SN ESARRSR RKQ HL EL AQV
Sbjct: 40 TASGSSDPSDEDNESGPCEQITNPVDMKRQRRKDSNCESARRSRWRKQAHLSELEAQV 213
>BE941734 similar to GP|7406677|emb| putative ripening-related bZIP protein
{Vitis vinifera}, partial (18%)
Length = 607
Score = 50.8 bits (120), Expect = 2e-07
Identities = 23/42 (54%), Positives = 34/42 (80%)
Frame = +2
Query: 29 QRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQM 70
+R+++RMI NRESA RSR RKQ + EL A+VA+L+ EN+++
Sbjct: 8 ERRQRRMIKNRESAARSRARKQAYTMELEAEVAKLKEENEEL 133
>TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - kidney bean,
partial (78%)
Length = 1250
Score = 50.1 bits (118), Expect = 3e-07
Identities = 27/76 (35%), Positives = 44/76 (57%)
Frame = +1
Query: 20 EEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQ 79
E +Q + ++ +R SNRESARRSR+RKQ +ELA +V L +EN + + +N +
Sbjct: 748 EAWMQNERELKRERRKQSNRESARRSRLRKQAEAEELARRVDALTAENLALKSEMNELAE 927
Query: 80 RFMIIDAENSVLSAQM 95
+ EN+ L ++
Sbjct: 928 NSAKLKIENATLKEKL 975
>TC78777 similar to PIR|T10985|T10985 regulator protein ROM2 - kidney bean,
partial (84%)
Length = 1822
Score = 48.9 bits (115), Expect = 7e-07
Identities = 26/72 (36%), Positives = 42/72 (58%)
Frame = +1
Query: 20 EEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQ 79
E +Q + ++ +R SNRESARRSR+RKQ +ELA +V L +E+ + + +N +
Sbjct: 967 EAWIQNERELKRERRKQSNRESARRSRLRKQAEAEELARKVESLNAESASLRSEINRLAE 1146
Query: 80 RFMIIDAENSVL 91
+ EN+ L
Sbjct: 1147 NSERLRMENAAL 1182
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.124 0.319
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,709,739
Number of Sequences: 36976
Number of extensions: 22998
Number of successful extensions: 193
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 181
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 188
length of query: 147
length of database: 9,014,727
effective HSP length: 87
effective length of query: 60
effective length of database: 5,797,815
effective search space: 347868900
effective search space used: 347868900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0113.7


