

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.13
(359 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
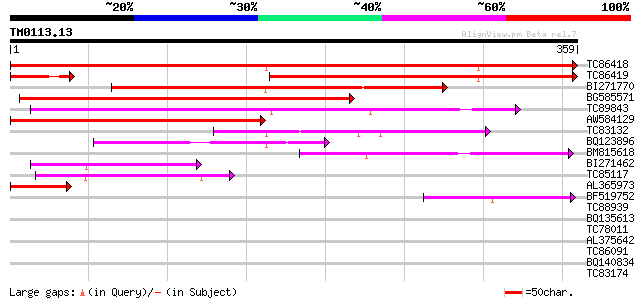
Score E
Sequences producing significant alignments: (bits) Value
TC86418 weakly similar to GP|18461100|dbj|BAB84353. S-adenosyl-L... 577 e-165
TC86419 weakly similar to GP|18461100|dbj|BAB84353. S-adenosyl-L... 308 2e-84
BI271770 weakly similar to GP|13235641|em S-adenosyl-L-methionin... 240 5e-64
BG585571 weakly similar to GP|13235641|em S-adenosyl-L-methionin... 223 9e-59
TC89843 similar to PIR|E85430|E85430 hypothetical protein AT4g36... 211 5e-55
AW584129 weakly similar to GP|13235641|em S-adenosyl-L-methionin... 180 7e-46
TC83132 similar to GP|21593254|gb|AAM65203.1 S-adenosyl-L-methio... 113 1e-25
BQ123896 weakly similar to PIR|A86285|A86 protein F9L1.6 [import... 110 6e-25
BM815618 weakly similar to GP|6651395|gb|A floral nectary-specif... 101 4e-22
BI271462 similar to GP|6651395|gb|A floral nectary-specific prot... 69 2e-12
TC85117 similar to GP|21593254|gb|AAM65203.1 S-adenosyl-L-methio... 68 5e-12
AL365973 similar to GP|13235641|emb S-adenosyl-L-methionine:sali... 47 1e-05
BF519752 weakly similar to GP|13235641|emb S-adenosyl-L-methioni... 44 1e-04
TC88939 similar to PIR|C84538|C84538 probable LRR receptor prote... 31 0.65
BQ135613 similar to GP|14017929|db KIAA1856 protein {Homo sapien... 28 4.2
TC78011 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 28 4.2
AL375642 28 7.1
TC86091 similar to PIR|T09896|T09896 hypothetical protein T22A6.... 28 7.1
BQ140834 27 9.3
TC83174 27 9.3
>TC86418 weakly similar to GP|18461100|dbj|BAB84353.
S-adenosyl-L-methionine:salicylic acid calboxyl
methyltransferase-like protein {Cucumis sativus},
partial (39%)
Length = 1306
Score = 577 bits (1487), Expect = e-165
Identities = 284/369 (76%), Positives = 321/369 (86%), Gaps = 10/369 (2%)
Frame = +2
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCS 60
MA+EQVLHMNGG+GD SYANNS+FQR VMLT KHILE+SI+RLYC TFP+CLKVADLGCS
Sbjct: 89 MAVEQVLHMNGGEGDTSYANNSTFQRMVMLTAKHILEESIMRLYCDTFPNCLKVADLGCS 268
Query: 61 SGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
SGPNAL+VASNIINT+DAVSQ L+ E P+FQFFLNDLFGNDFNTTFK LPDF KRLQ+EK
Sbjct: 269 SGPNALLVASNIINTIDAVSQKLSHESPMFQFFLNDLFGNDFNTTFKLLPDFIKRLQEEK 448
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNKGAIYL 174
G KF PCFFSGTPGSFYGRLFP++S+HFFHSSYSLHWLSK +PLNKG IYL
Sbjct: 449 GQKFSPCFFSGTPGSFYGRLFPDNSIHFFHSSYSLHWLSKTPDALQDAAIEPLNKGNIYL 628
Query: 175 TKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWVVIGMV 234
T+ SPPAV K YF QF++DF LFL SRS ELLP GAMV+TLIGRDE NEL+NAWVVIGM
Sbjct: 629 TRASPPAVQKTYFEQFQQDFSLFLRSRSSELLPSGAMVLTLIGRDEQNELMNAWVVIGMA 808
Query: 235 LNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSD 294
LNDMA+ L+E++KLD+ NIPSY PT++EIR+VIEEEGSFD+QRLETIRTDW+KN++V D
Sbjct: 809 LNDMAAVKLVEQSKLDSLNIPSYRPTSDEIRKVIEEEGSFDVQRLETIRTDWVKNVDVID 988
Query: 295 D----IDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELAN 350
D +DE TRAE VAK+IRAVAE ILKSEF E IMDELF RFKNKI+KL+GVEKLE+AN
Sbjct: 989 DEYTVVDEQTRAEGVAKFIRAVAEPILKSEFAEEIMDELFIRFKNKIIKLYGVEKLEVAN 1168
Query: 351 LVMHITKST 359
LVMHITK T
Sbjct: 1169LVMHITKRT 1195
>TC86419 weakly similar to GP|18461100|dbj|BAB84353.
S-adenosyl-L-methionine:salicylic acid calboxyl
methyltransferase-like protein {Cucumis sativus},
partial (32%)
Length = 879
Score = 308 bits (790), Expect = 2e-84
Identities = 152/199 (76%), Positives = 176/199 (88%), Gaps = 4/199 (2%)
Frame = +2
Query: 165 KPLNKGAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNEL 224
+PLNKG IYLT+ SPPAV K YF QF++DF LFL SRS ELLP GAMV+TLIGRDE NEL
Sbjct: 134 EPLNKGNIYLTRASPPAVQKTYFEQFQQDFSLFLRSRSSELLPSGAMVLTLIGRDEQNEL 313
Query: 225 INAWVVIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRT 284
+NAWVVIGM LNDMA+ L+E++KLD+FNIPSYCPT++EIR+VIEEEGSFD+QRLETIRT
Sbjct: 314 MNAWVVIGMALNDMAAVKLVEQSKLDSFNIPSYCPTSDEIRKVIEEEGSFDVQRLETIRT 493
Query: 285 DWIKNINVSDD----IDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKL 340
DW+KN++V DD +DE+TRAE VAK+IRAVAE ILKSEFGE IMDELF RFKNKI+KL
Sbjct: 494 DWVKNVDVIDDEYTVVDEETRAEGVAKFIRAVAEPILKSEFGEEIMDELFIRFKNKIIKL 673
Query: 341 HGVEKLELANLVMHITKST 359
+GVEKLE+ANLVMHITK T
Sbjct: 674 YGVEKLEVANLVMHITKRT 730
Score = 50.4 bits (119), Expect = 1e-06
Identities = 25/41 (60%), Positives = 30/41 (72%)
Frame = +2
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSII 41
MA+EQVLHMNGG+GD SYANNS+FQ T L+D+ I
Sbjct: 29 MAVEQVLHMNGGEGDTSYANNSTFQ------TPDALQDAAI 133
>BI271770 weakly similar to GP|13235641|em S-adenosyl-L-methionine:salicylic
acid carboxyl methyltransferase {Stephanotis
floribunda}, partial (31%)
Length = 676
Score = 240 bits (613), Expect = 5e-64
Identities = 118/218 (54%), Positives = 155/218 (70%), Gaps = 5/218 (2%)
Frame = +1
Query: 65 ALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKF 124
AL+V SNI+ +D +S +LN E P FQ +LNDL+ NDFNT K LPDF++ +QQE+G
Sbjct: 1 ALLVVSNIMKVIDKISLSLNHELPAFQIYLNDLYENDFNTILKLLPDFHQSIQQERGENH 180
Query: 125 GPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSK-----VNYTKPLNKGAIYLTKTSP 179
GPCF + TPGSFYGRLFPN+ + FFHSSY +HWLS+ +PL KG I +T+ SP
Sbjct: 181 GPCFINATPGSFYGRLFPNNYIDFFHSSYCVHWLSQAPKYSTKKAEPLIKGNICITRMSP 360
Query: 180 PAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWVVIGMVLNDMA 239
P+V++ Y QF DFK FL SRS EL G MV+TLIGR++N E I ++ + MVL++M
Sbjct: 361 PSVYEVYVEQFGRDFKNFLRSRSDELAMHGVMVLTLIGREKNGE-ITSYEALXMVLDEMV 537
Query: 240 SENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQ 277
L+E+AKLD FN+P Y PT EE++Q+IE EGSF +Q
Sbjct: 538 QXGLVEEAKLDMFNLPLYHPTIEEVKQMIEAEGSFTLQ 651
>BG585571 weakly similar to GP|13235641|em S-adenosyl-L-methionine:salicylic
acid carboxyl methyltransferase {Stephanotis
floribunda}, partial (54%)
Length = 723
Score = 223 bits (568), Expect = 9e-59
Identities = 111/212 (52%), Positives = 138/212 (64%)
Frame = +2
Query: 7 LHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSGPNAL 66
+HMNGGDG SYANNS FQ + TKHI E++I LY +T P L +ADLGCS G N L
Sbjct: 2 VHMNGGDGKTSYANNSFFQGKGISLTKHIREEAITSLYSSTLPRSLAIADLGCSCGQNTL 181
Query: 67 MVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGP 126
V S II V+ + Q L P ++ F NDL GNDFN FKSL F +L E + P
Sbjct: 182 SVVSEIIMVVEKLCQQLKYASPEYKIFFNDLSGNDFNNIFKSLDSFKHKLLDEIKTEMSP 361
Query: 127 CFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAVHKAY 186
C+F G PGSFY R+FP+ S+HF H SYSLHWLSKV NKG IY++ TSP V KAY
Sbjct: 362 CYFFGVPGSFYDRVFPDRSLHFVHCSYSLHWLSKVPEGIDNNKGNIYISDTSPSNVVKAY 541
Query: 187 FAQFREDFKLFLGSRSCELLPGGAMVITLIGR 218
+ QF+ D +FL R+ EL+ GG +V+T++GR
Sbjct: 542 YEQFQRDLSIFLKCRAKELVEGGRIVLTMVGR 637
>TC89843 similar to PIR|E85430|E85430 hypothetical protein AT4g36470
[imported] - Arabidopsis thaliana, partial (54%)
Length = 940
Score = 211 bits (536), Expect = 5e-55
Identities = 117/320 (36%), Positives = 170/320 (52%), Gaps = 10/320 (3%)
Frame = +1
Query: 14 GDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSGPNALMVASNII 73
G SYA NSS Q+ KHI+ +++ LY T P + +ADLGCSSGPN L + +I
Sbjct: 1 GKTSYAKNSSLQKKASDKVKHIIIETVEELYIETTPKSIGIADLGCSSGPNTLSIIKDIF 180
Query: 74 NTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGPCFFSGTP 133
T+ S + F+ + NDL NDFN+ FK+LP+F K L Q++ + F F G P
Sbjct: 181 QTIQVTSHKIMHHSTEFRVYFNDLPTNDFNSIFKALPEFQKLLNQDRKNGFPSIFMGGYP 360
Query: 134 GSFYGRLFPNDSVHFFHSSYSLHWLSKVNYT------KPLNKGAIYLTKTSPPAVHKAYF 187
GSFYGRLFPN +HF HSS+ LHWLS+V T + LNKG +Y+ SP V +AY+
Sbjct: 361 GSFYGRLFPNSYLHFVHSSHCLHWLSRVPPTIYDEQKRSLNKGCVYICDKSPEVVSQAYY 540
Query: 188 AQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINA----WVVIGMVLNDMASENL 243
QF+EDF L + ++ N L W ++ + S+
Sbjct: 541 KQFQEDFPCSFDQGLKN*LLAEKWCLHF*EEEDLNMLDRGNSFFWEILTRSFTILVSQGE 720
Query: 244 IEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAE 303
IE+ KLD++++ Y P+ EEI + + GS ++RLE D + ++
Sbjct: 721 IEQEKLDSYDVHFYAPSREEIEDEVMKAGSLKLERLEMFDID-------KKEQGRESYGT 879
Query: 304 AVAKYIRAVAESILKSEFGE 323
VAK +RA+ ES++ + FGE
Sbjct: 880 DVAKAVRAIQESMVSNHFGE 939
>AW584129 weakly similar to GP|13235641|em S-adenosyl-L-methionine:salicylic
acid carboxyl methyltransferase {Stephanotis
floribunda}, partial (31%)
Length = 568
Score = 180 bits (457), Expect = 7e-46
Identities = 88/162 (54%), Positives = 109/162 (66%)
Frame = +2
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCS 60
M + QV+HMNGGDG+ SYA NS Q N + TK I +++I LY T L +ADLGCS
Sbjct: 62 MDLGQVVHMNGGDGETSYAKNSFHQGNAVSLTKPIRDEAITSLYSKTLFKSLAIADLGCS 241
Query: 61 SGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
SGPN L V S+II V+ + Q LN P ++ F ND+ GNDFN FKSL +F ++LQ E
Sbjct: 242 SGPNTLFVVSDIIMVVEKLCQQLNHSSPEYKIFFNDVSGNDFNNIFKSLDNFKEKLQDEI 421
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVN 162
K C+F G PGSFY R+FPN S+HF HSS+SL WLSKVN
Sbjct: 422 KTKMSSCYFFGVPGSFYSRVFPNRSLHFIHSSHSLQWLSKVN 547
>TC83132 similar to GP|21593254|gb|AAM65203.1
S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase-like protein, partial (63%)
Length = 1075
Score = 113 bits (283), Expect = 1e-25
Identities = 70/187 (37%), Positives = 100/187 (53%), Gaps = 12/187 (6%)
Frame = +1
Query: 130 SGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNKGAIYLTKTSPPAVH 183
+G PGSFY RLFP SV FHS++SLHWLSK+ + NKG +++ + +
Sbjct: 1 AGVPGSFYRRLFPARSVDVFHSAFSLHWLSKIPESVLDKKSIAYNKGKVFIHGANESTAN 180
Query: 184 KAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRD--ENNELINAWVVIGM----VLND 237
AY QF+ D FL +RS E+ G+M + +GR + E A V+ G +D
Sbjct: 181 -AYKRQFKTDLASFLSARSVEMKREGSMFLVCLGRTSVDPTEQGGAGVLFGTHFQDAWDD 357
Query: 238 MASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDID 297
+ E LI K D FNIP Y P+ ++ ++V+E GSF I +LE + +N DD +
Sbjct: 358 LVQEGLISSTKRDNFNIPVYAPSMQDFKEVVEANGSFVINKLEVFKGGXPLVLNKPDDAN 537
Query: 298 EDTRAEA 304
E RA A
Sbjct: 538 EVGRALA 558
>BQ123896 weakly similar to PIR|A86285|A86 protein F9L1.6 [imported] -
Arabidopsis thaliana, partial (31%)
Length = 537
Score = 110 bits (276), Expect = 6e-25
Identities = 63/155 (40%), Positives = 86/155 (54%), Gaps = 6/155 (3%)
Frame = +1
Query: 54 VADLGCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFY 113
+ADLGCS+GPN + II ++ ++ L P FQ F ND NDFNT FK LP
Sbjct: 109 IADLGCSTGPNTFIAIQCIIEAIELQYKSQGLAIPEFQVFFNDQISNDFNTLFKKLPSNR 288
Query: 114 KRLQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPL 167
F +G PGSFYGRLFP +S++ HSS SL+W+SKV +
Sbjct: 289 N------------YFAAGVPGSFYGRLFPKESLNVVHSSASLNWISKVPKEITDRSSAAC 432
Query: 168 NKGAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRS 202
NKG I+ T +P V AY Q+++D ++FL +R+
Sbjct: 433 NKGRIHYT-NAPKEVVDAYANQYQKDMEIFLHARA 534
>BM815618 weakly similar to GP|6651395|gb|A floral nectary-specific protein
{Brassica rapa subsp. pekinensis}, partial (24%)
Length = 664
Score = 101 bits (252), Expect = 4e-22
Identities = 59/179 (32%), Positives = 96/179 (52%), Gaps = 5/179 (2%)
Frame = +1
Query: 184 KAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNEL-----INAWVVIGMVLNDM 238
KAY+ QF+ DF +FL R+ EL+ GG MV+T+ GR + W ++ VLN M
Sbjct: 4 KAYYDQFKRDFSVFLKCRAEELVEGGCMVLTMPGRRNEDPCDIKYCCYYWELLAAVLNGM 183
Query: 239 ASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDE 298
E +I++ +++ FN+P Y P+ E+ + EGSF I RLE D +
Sbjct: 184 VLEGIIKEDQVNTFNVPQYYPSPYEVELEVLNEGSFAINRLELFEA-------YVDGSNH 342
Query: 299 DTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHITK 357
A+ +RA+AE ++ S FGE I++E+F R + I+ EKL+ +++ +T+
Sbjct: 343 HEYVYNAARLMRAMAEPLVVSHFGEDIIEEIFSRHQKIIIDKLPKEKLKAVKVIISLTR 519
>BI271462 similar to GP|6651395|gb|A floral nectary-specific protein
{Brassica rapa subsp. pekinensis}, partial (6%)
Length = 492
Score = 69.3 bits (168), Expect = 2e-12
Identities = 41/112 (36%), Positives = 61/112 (53%), Gaps = 4/112 (3%)
Frame = +2
Query: 14 GDASYANNSSFQRNVMLTTKHILEDSIIRLYCAT--FPSC-LKVADLGCSSGPNALMVAS 70
G+ SYA NS QR ++ T + +I+ + C+T +P + +ADLGCSSGPNAL V S
Sbjct: 116 GETSYAMNSFVQRKIISLTNQATKKAIVEILCSTKRWPIMKMGIADLGCSSGPNALRVIS 295
Query: 71 NIINTVDAVSQNLNLEQP-VFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKG 121
I+ ++A S LN P ++NDLF F F ++++ KG
Sbjct: 296 EIVEAINATSSMLNRPAPKELMLYMNDLFHQ*LQQHFCFFAIFPQKIKPTKG 451
Score = 37.4 bits (85), Expect = 0.009
Identities = 15/25 (60%), Positives = 19/25 (76%)
Frame = +3
Query: 98 FGNDFNTTFKSLPDFYKRLQQEKGH 122
F NDFN F SLP F+K+L Q+KG+
Sbjct: 381 FTNDFNNIFASLPSFHKKLSQQKGN 455
>TC85117 similar to GP|21593254|gb|AAM65203.1
S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase-like protein, partial (29%)
Length = 557
Score = 68.2 bits (165), Expect = 5e-12
Identities = 45/134 (33%), Positives = 62/134 (45%), Gaps = 8/134 (5%)
Frame = +1
Query: 17 SYANNSSFQRNVMLTTKHILEDSIIRLYCA-----TFPSCLKVADLGCSSGPNALMVASN 71
+YANNS Q + H L +++ ++ VADLGCS G N + V +
Sbjct: 154 AYANNSQAQAIHAKSMIHFLRETLDKVKLGGGGGGDGDKAFVVADLGCSCGSNTINVVNV 333
Query: 72 IINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK---GHKFGPCF 128
IIN + + L P F + +DL NDFNT F+ LP + E+ F
Sbjct: 334 IINHIIKRYEALGCNPPEFSAYFSDLPSNDFNTLFQLLPPLANGISMEECLXADNQRSYF 513
Query: 129 FSGTPGSFYGRLFP 142
+G PGSFY LFP
Sbjct: 514 VAGVPGSFYRXLFP 555
>AL365973 similar to GP|13235641|emb S-adenosyl-L-methionine:salicylic acid
carboxyl methyltransferase {Stephanotis floribunda},
partial (9%)
Length = 437
Score = 47.0 bits (110), Expect = 1e-05
Identities = 20/39 (51%), Positives = 29/39 (74%)
Frame = +3
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDS 39
M +EQVLHMNGG G+ SY+NNS Q+ V+ TK + +++
Sbjct: 51 MKVEQVLHMNGGIGETSYSNNSLLQKKVISLTKEMRDEA 167
>BF519752 weakly similar to GP|13235641|emb S-adenosyl-L-methionine:salicylic
acid carboxyl methyltransferase {Stephanotis
floribunda}, partial (14%)
Length = 543
Score = 43.5 bits (101), Expect = 1e-04
Identities = 30/103 (29%), Positives = 51/103 (49%), Gaps = 7/103 (6%)
Frame = -1
Query: 263 EIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEA-------VAKYIRAVAES 315
E++ + +EGSF I RL +W + E E+ V + +R V E
Sbjct: 543 EVKLEVLKEGSFTIDRLGVSEVNWNALDQWNALACESQMFESLGDGAYNVMQCMRPVFEP 364
Query: 316 ILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHITKS 358
+L FGE+I+DELF R++ +V EK + N+ + +T++
Sbjct: 363 LLVRHFGESIIDELFDRYQEILVDRMSKEKTKFVNVPVVLTRN 235
>TC88939 similar to PIR|C84538|C84538 probable LRR receptor protein kinase
[imported] - Arabidopsis thaliana, partial (20%)
Length = 1565
Score = 31.2 bits (69), Expect = 0.65
Identities = 20/65 (30%), Positives = 32/65 (48%), Gaps = 7/65 (10%)
Frame = +2
Query: 145 SVHFFHSS----YSLHWLSKVNYTKPLNKGAIYLTKTSPPAVHKAYFAQ---FREDFKLF 197
S+H H S S+ W++++ + +G YL SPP VH+ A + F++
Sbjct: 455 SLHKVHDSDGKLQSMDWITRLKIATGVAEGLAYLHDCSPPLVHRDIRASSILLDDKFEVR 634
Query: 198 LGSRS 202
LGS S
Sbjct: 635 LGSLS 649
>BQ135613 similar to GP|14017929|db KIAA1856 protein {Homo sapiens}, partial
(3%)
Length = 1048
Score = 28.5 bits (62), Expect = 4.2
Identities = 13/34 (38%), Positives = 19/34 (55%)
Frame = +2
Query: 10 NGGDGDASYANNSSFQRNVMLTTKHILEDSIIRL 43
NGG+GD S F +NV++T K + IR+
Sbjct: 560 NGGEGDGETMGGSMFVQNVLVTAKFEGSEYAIRI 661
>TC78011 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (5%)
Length = 637
Score = 28.5 bits (62), Expect = 4.2
Identities = 13/30 (43%), Positives = 19/30 (63%)
Frame = +3
Query: 135 SFYGRLFPNDSVHFFHSSYSLHWLSKVNYT 164
SF F + SVH F SS S++ +S +N+T
Sbjct: 39 SFSFHFFYHPSVHHFPSSLSIYHISSINHT 128
>AL375642
Length = 271
Score = 27.7 bits (60), Expect = 7.1
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +1
Query: 155 LHWLSKVNYTKPLNKGAIYLTKTSPPAVHKAY 186
LHW+ +NY KP+ Y + +S PAV Y
Sbjct: 13 LHWVVGLNYDKPVCDFYCYASASSLPAVSCFY 108
>TC86091 similar to PIR|T09896|T09896 hypothetical protein T22A6.160 -
Arabidopsis thaliana, partial (69%)
Length = 1865
Score = 27.7 bits (60), Expect = 7.1
Identities = 18/67 (26%), Positives = 33/67 (48%)
Frame = +1
Query: 275 DIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFK 334
D+QR TI T VSDD+ + + VA + + ++++ FG D+ F +F
Sbjct: 916 DLQRFATIMTPPTSRKWVSDDLAVISESREVASDL--ITDALIDQVFG----DKSFEKFG 1077
Query: 335 NKIVKLH 341
++ +H
Sbjct: 1078KGLISVH 1098
>BQ140834
Length = 560
Score = 27.3 bits (59), Expect = 9.3
Identities = 24/69 (34%), Positives = 32/69 (45%)
Frame = +3
Query: 55 ADLGCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYK 114
A L P +LMV + ++ V L E F+F N+L D FK LP Y
Sbjct: 294 AKLHFELSPTSLMVRTRRLDHVSRTLHLLECEPEKFEFGRNNLVKLD--PIFK-LPLRYG 464
Query: 115 RLQQEKGHK 123
+L +EK HK
Sbjct: 465 KLFKEKMHK 491
>TC83174
Length = 1323
Score = 27.3 bits (59), Expect = 9.3
Identities = 24/69 (34%), Positives = 32/69 (45%)
Frame = +2
Query: 55 ADLGCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYK 114
A L P +LMV + ++ V L E F+F N+L D FK LP Y
Sbjct: 314 AKLHFELSPTSLMVRTRRLDHVSRTLHLLECEPEKFEFGRNNLVKLD--PIFK-LPLRYG 484
Query: 115 RLQQEKGHK 123
+L +EK HK
Sbjct: 485 KLFKEKMHK 511
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.137 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,564,964
Number of Sequences: 36976
Number of extensions: 148318
Number of successful extensions: 823
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 803
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 808
length of query: 359
length of database: 9,014,727
effective HSP length: 97
effective length of query: 262
effective length of database: 5,428,055
effective search space: 1422150410
effective search space used: 1422150410
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0113.13


