

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0112.2
(150 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
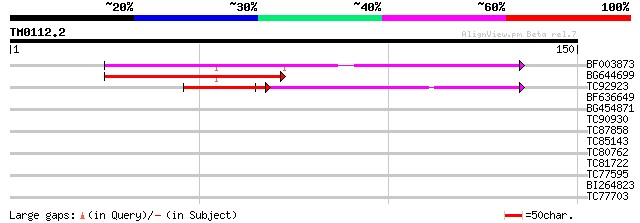
Score E
Sequences producing significant alignments: (bits) Value
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 53 5e-08
BG644699 similar to PIR|T07863|T078 probable polyprotein - pinea... 50 5e-07
TC92923 similar to GP|6466937|gb|AAF13073.1| putative retroeleme... 34 9e-05
BF636649 similar to GP|21628724|emb OSJNBa0033H08.7 {Oryza sativ... 39 0.001
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 32 0.075
TC90930 similar to GP|1742955|emb|CAA96058.1 CLC-b chloride chan... 27 2.4
TC87858 similar to GP|9757903|dbj|BAB08350.1 gene_id:MMN10.5~unk... 27 2.4
TC85143 similar to PIR|T11852|T11852 lipoxygenase (EC 1.13.11.12... 27 4.1
TC80762 27 4.1
TC81722 weakly similar to GP|21593895|gb|AAM65862.1 unknown {Ara... 26 5.3
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 26 7.0
BI264823 similar to GP|12848193|dbj data source:MGD source key:... 26 7.0
TC77703 similar to GP|14334884|gb|AAK59620.1 unknown protein {Ar... 25 9.1
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 52.8 bits (125), Expect = 5e-08
Identities = 35/117 (29%), Positives = 58/117 (48%), Gaps = 6/117 (5%)
Frame = +2
Query: 26 QLTAPYYGPYPIIQRIGAVAYRLQLPEG-GRVHPVFHASLLKEAVGN-----NSVELQLL 79
+LT + GPY I +R+G VAYR+ LP +H VFH S L++ V + S ++Q+
Sbjct: 29 KLTVRFIGPYQISERVGTVAYRVGLPPHLLNLHDVFHVSQLRKYVPDPSHVIQSDDVQVR 208
Query: 80 DHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQFP 136
D+LT E + P + + T + +P V + W + TW+ + +P
Sbjct: 209 DNLTVETL----PVRIDDRKVKTLRGKEIPLVRVVWDRANGESLTWELESKMVESYP 367
>BG644699 similar to PIR|T07863|T078 probable polyprotein - pineapple
retrotransposon dea1 (fragment), partial (5%)
Length = 231
Score = 49.7 bits (117), Expect = 5e-07
Identities = 24/49 (48%), Positives = 32/49 (64%), Gaps = 1/49 (2%)
Frame = +2
Query: 26 QLTAPYYGPYPIIQRIGAVAYRLQLPEG-GRVHPVFHASLLKEAVGNNS 73
+L+ Y GP+ +I+RIG VAY L LP G VHPVFH S+ K G+ +
Sbjct: 65 KLSLRYIGPFEVIKRIGEVAYELALPPGLSGVHPVFHVSMFKRYHGDGN 211
>TC92923 similar to GP|6466937|gb|AAF13073.1| putative retroelement pol
polyprotein {Arabidopsis thaliana}, partial (1%)
Length = 625
Score = 33.9 bits (76), Expect(2) = 9e-05
Identities = 23/71 (32%), Positives = 35/71 (48%)
Frame = +2
Query: 66 KEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTW 125
K +VG+ VELQL + + E P ++ +R + T I+W+ K + +
Sbjct: 266 KSSVGSYPVELQLPEGMEVEINDEADPQFMLATRQIREGDITT**A-IKWKAKTLEHASL 442
Query: 126 KDTLNIRSQFP 136
KD IRSQFP
Sbjct: 443 KDDFTIRSQFP 475
Score = 27.3 bits (59), Expect(2) = 9e-05
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = +3
Query: 47 RLQLPEGGRVHPVFHASLLKEAV 69
+LQL + +H +FH S LK AV
Sbjct: 207 KLQLTQTSTIHSIFHVSQLKRAV 275
>BF636649 similar to GP|21628724|emb OSJNBa0033H08.7 {Oryza sativa}, partial
(4%)
Length = 653
Score = 38.5 bits (88), Expect = 0.001
Identities = 16/33 (48%), Positives = 25/33 (75%)
Frame = +1
Query: 25 PQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVH 57
P+ A YGPY +I++IG+VA++L LPE ++H
Sbjct: 382 PKYVA*CYGPYQVIKQIGSVAFKL*LPEQHQIH 480
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 32.3 bits (72), Expect = 0.075
Identities = 14/29 (48%), Positives = 18/29 (61%)
Frame = +2
Query: 43 AVAYRLQLPEGGRVHPVFHASLLKEAVGN 71
+ AY+L LP +V P+FH S LK GN
Sbjct: 551 SAAYKLSLPSTAKVPPIFHVSQLKPFHGN 637
>TC90930 similar to GP|1742955|emb|CAA96058.1 CLC-b chloride channel protein
{Arabidopsis thaliana}, partial (35%)
Length = 857
Score = 27.3 bits (59), Expect = 2.4
Identities = 14/38 (36%), Positives = 18/38 (46%)
Frame = +3
Query: 88 ASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTW 125
+ +HP ITSR T LP W W+ + PTW
Sbjct: 276 SKMHPL*SITSRVRT-----LPHEWKVWKFQTIQLPTW 374
>TC87858 similar to GP|9757903|dbj|BAB08350.1 gene_id:MMN10.5~unknown
protein {Arabidopsis thaliana}, partial (34%)
Length = 2037
Score = 27.3 bits (59), Expect = 2.4
Identities = 11/40 (27%), Positives = 23/40 (57%)
Frame = -2
Query: 33 GPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLKEAVGNN 72
GP P++++I ++ E GR+H ++H ++ G+N
Sbjct: 107 GP*PMVKKIDSII------EAGRLHDLYHRRMIGTLTGDN 6
>TC85143 similar to PIR|T11852|T11852 lipoxygenase (EC 1.13.11.12) - kidney
bean, partial (93%)
Length = 3009
Score = 26.6 bits (57), Expect = 4.1
Identities = 19/66 (28%), Positives = 29/66 (43%)
Frame = +1
Query: 34 PYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPF 93
P+P ++ GAV+ L PEG V V N++ QL+ H + PF
Sbjct: 1456 PHPDGEKYGAVSKVLLPPEGHGVQRTIWQLAKAYVVVNDACFHQLMSHWLNTHCV-IEPF 1632
Query: 94 SVITSR 99
+ T+R
Sbjct: 1633 IIATNR 1650
>TC80762
Length = 1121
Score = 26.6 bits (57), Expect = 4.1
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = -3
Query: 100 FTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQFPVFNLE 141
FT+ + PQ I +Q + D+PT L+ R FPVF ++
Sbjct: 264 FTSATTTIQPQT-IMFQARYKDDPTTLYPLS*RQLFPVFTIK 142
>TC81722 weakly similar to GP|21593895|gb|AAM65862.1 unknown {Arabidopsis
thaliana}, partial (69%)
Length = 552
Score = 26.2 bits (56), Expect = 5.3
Identities = 10/34 (29%), Positives = 17/34 (49%)
Frame = -3
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFS 94
HA+ + N +LQ++ E + +HPFS
Sbjct: 430 HATKCYDCFSNKEPQLQVISQDVNEHLVMIHPFS 329
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 25.8 bits (55), Expect = 7.0
Identities = 15/49 (30%), Positives = 22/49 (44%)
Frame = +2
Query: 48 LQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVI 96
++L G V+P FH LL+ A D L G+EV P ++
Sbjct: 1151 VELNVPGTVYPKFHVDLLRRAAS---------DPLPGQEVVDPQPPPIV 1270
>BI264823 similar to GP|12848193|dbj data source:MGD source key:MGI:1913153
evidence:ISS~mitochondrial ribosomal protein
S31~putative, partial (6%)
Length = 667
Score = 25.8 bits (55), Expect = 7.0
Identities = 17/67 (25%), Positives = 33/67 (48%), Gaps = 6/67 (8%)
Frame = +1
Query: 28 TAPYYGPYPIIQRIGA------VAYRLQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDH 81
T + P+ IIQR G + ++ + GR++ V E G +++++LLD
Sbjct: 193 TNSFTTPWSIIQRRGNKIVGSDIRVGKKIGKQGRIYEVLKVDHSHEGRGKATLKVELLDI 372
Query: 82 LTGEEVA 88
+ G +V+
Sbjct: 373 IQGTKVS 393
>TC77703 similar to GP|14334884|gb|AAK59620.1 unknown protein {Arabidopsis
thaliana}, partial (77%)
Length = 2410
Score = 25.4 bits (54), Expect = 9.1
Identities = 13/25 (52%), Positives = 18/25 (72%), Gaps = 1/25 (4%)
Frame = +3
Query: 69 VGNNSVEL-QLLDHLTGEEVASVHP 92
+G N VEL Q + +T EEVA++HP
Sbjct: 1245 MGANIVELFQQMTVMTDEEVAAIHP 1319
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,848,661
Number of Sequences: 36976
Number of extensions: 60941
Number of successful extensions: 276
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 275
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 275
length of query: 150
length of database: 9,014,727
effective HSP length: 87
effective length of query: 63
effective length of database: 5,797,815
effective search space: 365262345
effective search space used: 365262345
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0112.2


