

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.9
(341 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
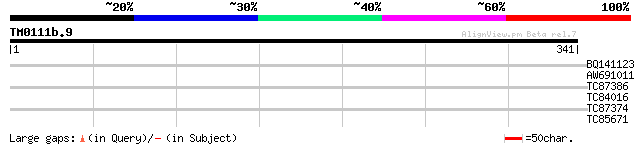
BQ141123 homologue to GP|14488272|db Na+/H+ exchanger protein {I... 28 5.1
AW691011 similar to PIR|T01804|T018 Na+/H+-exchanging protein 3 ... 28 5.1
TC87386 similar to GP|3717239|emb|CAA03732.1 unnamed protein pro... 28 6.7
TC84016 similar to GP|11994521|dbj|BAB02585. gene_id:MUO10.12~un... 28 6.7
TC87374 similar to PIR|T52310|T52310 Lil3 protein [imported] - A... 27 8.7
TC85671 similar to SP|Q9XF97|RL4_PRUAR 60S ribosomal protein L4 ... 27 8.7
>BQ141123 homologue to GP|14488272|db Na+/H+ exchanger protein {Ipomoea nil},
partial (18%)
Length = 655
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +1
Query: 252 STWLVKVIRGFLRAQCQPLPLKLAL 276
++WL+K++ F QC PLP+ L L
Sbjct: 307 TSWLLKLLLLFQNYQCYPLPIMLLL 381
>AW691011 similar to PIR|T01804|T018 Na+/H+-exchanging protein 3 homolog
A_TM021B04.4 - Arabidopsis thaliana, partial (21%)
Length = 654
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +1
Query: 252 STWLVKVIRGFLRAQCQPLPLKLAL 276
++WL+K++ F QC PLP+ L L
Sbjct: 301 TSWLLKLLLLFQNYQCYPLPIMLLL 375
>TC87386 similar to GP|3717239|emb|CAA03732.1 unnamed protein product
{Cyamopsis tetragonoloba}, partial (89%)
Length = 1868
Score = 27.7 bits (60), Expect = 6.7
Identities = 10/25 (40%), Positives = 14/25 (56%)
Frame = +1
Query: 72 SWISWLASSQVGHVGSFEAAWWSWY 96
SWI + +G +GSF WW+ Y
Sbjct: 1474 SWIWFNGKEYLGSLGSFRDLWWAEY 1548
>TC84016 similar to GP|11994521|dbj|BAB02585. gene_id:MUO10.12~unknown
protein {Arabidopsis thaliana}, partial (14%)
Length = 626
Score = 27.7 bits (60), Expect = 6.7
Identities = 11/30 (36%), Positives = 15/30 (49%)
Frame = +2
Query: 74 ISWLASSQVGHVGSFEAAWWSWYPRCDTRE 103
ISW S++ A+WW RCDT +
Sbjct: 284 ISWDGSTRSSSFEQCYASWWGCISRCDTSD 373
>TC87374 similar to PIR|T52310|T52310 Lil3 protein [imported] - Arabidopsis
thaliana, partial (44%)
Length = 1119
Score = 27.3 bits (59), Expect = 8.7
Identities = 18/47 (38%), Positives = 24/47 (50%)
Frame = -2
Query: 6 ISVSLWIKFLVV*PHGRVGF*TLQVEFSFLSLSLPPFQLILLRSFGY 52
+S+S+W F V+ F L++ F LSLSL F LIL Y
Sbjct: 155 VSISVWFVFSVLSFQFLFMFMFLELSFYPLSLSLSHFYLILWNGHKY 15
>TC85671 similar to SP|Q9XF97|RL4_PRUAR 60S ribosomal protein L4 (L1).
[Apricot] {Prunus armeniaca}, complete
Length = 1508
Score = 27.3 bits (59), Expect = 8.7
Identities = 13/27 (48%), Positives = 15/27 (55%), Gaps = 1/27 (3%)
Frame = +2
Query: 75 SWLASSQVGHVGSFEA-AWWSWYPRCD 100
SW Q G V SFEA WW++ CD
Sbjct: 743 SWC*DHQCGKVESFEACTWWAFGEICD 823
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.162 0.621
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,808,456
Number of Sequences: 36976
Number of extensions: 242597
Number of successful extensions: 2782
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1967
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 751
Number of HSP's gapped (non-prelim): 2123
length of query: 341
length of database: 9,014,727
effective HSP length: 97
effective length of query: 244
effective length of database: 5,428,055
effective search space: 1324445420
effective search space used: 1324445420
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0111b.9


