

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.7
(402 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
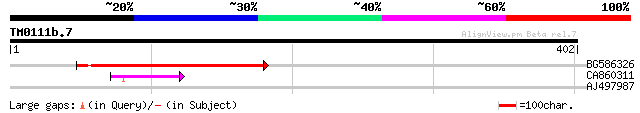
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 162 2e-40
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 41 7e-04
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 35 0.067
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 162 bits (410), Expect = 2e-40
Identities = 79/136 (58%), Positives = 100/136 (73%)
Frame = +2
Query: 48 TAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNYLTHDLELAV 107
T+AP+L LP E Y VY +AS GLGCVL QH + +AYAS QL+ HE NY THDLE+A
Sbjct: 8 TSAPILVLP-ELITYVVYTDASITGLGCVLTQHEKVIAYASRQLRKHEGNYPTHDLEMAA 184
Query: 108 VVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKA 167
VVFALKIWR YLYG+ + ++HK LKY+F +LN+ QR WM F+ DYD + Y+PGKA
Sbjct: 185 VVFALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELNLRQRRWMEFVADYDLDITYYPGKA 364
Query: 168 NVVADALSHQAIHVSS 183
N+VADALS + + VS+
Sbjct: 365 NLVADALSRRRVDVSA 412
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 41.2 bits (95), Expect = 7e-04
Identities = 22/58 (37%), Positives = 31/58 (52%), Gaps = 5/58 (8%)
Frame = +1
Query: 72 GLGCVLMQ-----HREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTF 124
G+G L Q H +AYAS L ERNY + E ++A++ +RHYL+G F
Sbjct: 13 GIGAGLSQKDEENHEHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYLHGPKF 186
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 34.7 bits (78), Expect = 0.067
Identities = 20/73 (27%), Positives = 34/73 (46%), Gaps = 1/73 (1%)
Frame = -2
Query: 108 VVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFTLLYHPG-K 166
+ +A K RHY+ T + S +KY+F+ L W + +YD K
Sbjct: 620 LAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRSQKAIK 441
Query: 167 ANVVADALSHQAI 179
+++AD L+HQ +
Sbjct: 440 GSILADHLAHQPL 402
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.353 0.155 0.544
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,095,238
Number of Sequences: 36976
Number of extensions: 181435
Number of successful extensions: 1921
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1404
Number of HSP's successfully gapped in prelim test: 59
Number of HSP's that attempted gapping in prelim test: 502
Number of HSP's gapped (non-prelim): 1488
length of query: 402
length of database: 9,014,727
effective HSP length: 98
effective length of query: 304
effective length of database: 5,391,079
effective search space: 1638888016
effective search space used: 1638888016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0111b.7


