

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
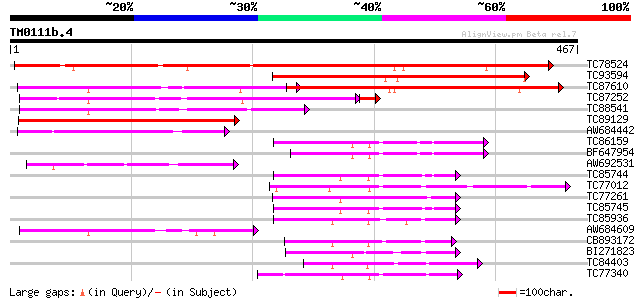
Query= TM0111b.4
(467 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78524 similar to PIR|D84561|D84561 probable AAA-type ATPase [i... 342 1e-94
TC93594 weakly similar to GP|20466452|gb|AAM20543.1 unknown prot... 224 4e-59
TC87610 similar to GP|9294102|dbj|BAB01954.1 contains similarity... 221 6e-58
TC87252 similar to GP|11559424|dbj|BAB18781. mitochondrial prote... 194 9e-52
TC88541 weakly similar to GP|10176993|dbj|BAB10225. contains sim... 138 4e-33
TC89129 weakly similar to GP|17385707|dbj|BAB78658. hypothetical... 136 2e-32
AW684442 similar to GP|9758309|dbj contains similarity to AAA-ty... 87 2e-17
TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AA... 69 4e-12
BF647954 homologue to GP|11094192|db 26S proteasome regulatory p... 67 1e-11
AW692531 weakly similar to GP|11559424|dbj mitochondrial protein... 66 2e-11
TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15... 65 7e-11
TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulator... 64 9e-11
TC77261 homologue to GP|8777330|dbj|BAA96920.1 26S proteasome AA... 64 1e-10
TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome re... 64 2e-10
TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidops... 62 4e-10
AW684609 similar to GP|11559424|dbj mitochondrial protein-like p... 61 1e-09
CB893172 homologue to PIR|T45642|T45 FtsH metalloproteinase-like... 60 2e-09
BI271823 similar to GP|21592745|gb cell division protein FtsH-li... 57 1e-08
TC84403 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotea... 57 2e-08
TC77340 SP|Q9BAE0|FTSH_MEDSA Cell division protein ftsH homolog ... 57 2e-08
>TC78524 similar to PIR|D84561|D84561 probable AAA-type ATPase [imported] -
Arabidopsis thaliana, partial (15%)
Length = 1697
Score = 342 bits (878), Expect = 1e-94
Identities = 194/471 (41%), Positives = 284/471 (60%), Gaps = 27/471 (5%)
Frame = +3
Query: 5 NTKSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFS-----TQFTVVIE 59
+ S AS + ML+R+ ND IP +L +FI K L+R F+ Q ++ I+
Sbjct: 78 SASSWFEVYASFSTFMMLLRTAINDLIPLKLRNFIISK---LTRFFTDYQPNNQVSLQID 248
Query: 60 EFQGMTKNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVT 119
+F + N +Y AA+ Y+ TK + + + +K GK + D + V D+F+ +++
Sbjct: 249 QFWDGSTNHLYYAAKEYIPTKISNTYKSLKVGKISKHNNMVLAFDGKQVVEDEFDDIKLK 428
Query: 120 WKLTCIRVDSSSSRTRYTNMSSSLMS--------EVRSYELSFHKKHKEKMFNSYLPYVL 171
W+L V++S++ + N + + LSF +KH++K+ Y+P+VL
Sbjct: 429 WRL----VENSNNGDGFDNPKKEYKEYKHRSKDYDENGFVLSFDEKHRDKVMEKYIPHVL 596
Query: 172 ERAKAIKQENMAVKIYTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGK 231
+AIK N +KI++++ W ++ HP +F +LA+D LK ++ DLDRFL+ K
Sbjct: 597 STYEAIKAGNKTLKIHSMQSGPWKQSD--LTHPASFDSLAMDPDLKNSIIDDLDRFLRRK 770
Query: 232 EFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAM 291
+ Y + GK WKRGYLLYGPPGTGKSSLIAAMA YL +D+YDLDL+++ N +L +
Sbjct: 771 KLYKKVGKPWKRGYLLYGPPGTGKSSLIAAMAKYLKFDVYDLDLSSVFSNSELMRAMRET 950
Query: 292 SNRSILVIEDIDCSIKLPNREEDE--------EAAKNGDN----KMTLSGLLNVIDGLWS 339
SNRSI+V EDIDC+ ++ +R + + + K G N K TLSGLLN +DGLWS
Sbjct: 951 SNRSIIVFEDIDCNSEVLDRAKPDKFPDMDFLDGIKMGKNMPPRKFTLSGLLNYMDGLWS 1130
Query: 340 CCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGI--SQHKLF 397
CGEERI+IFTTNH++++DPALLRPGRMDMH++LS+ AF LA YL I + H LF
Sbjct: 1131SCGEERILIFTTNHKDKVDPALLRPGRMDMHIHLSFLKAKAFRILAANYLDIEGNHHSLF 1310
Query: 398 EQIGGLLRHVQVTPAEVAGQLTKIADVEECLHGLVKFLQDKTQNDQNINQD 448
EQI LL V V+PA VA L + D + L LVKFLQD+ ++ +Q+
Sbjct: 1311EQIEELLEKVDVSPAVVAEYLLRSEDPDVALGALVKFLQDQEIVNEETSQE 1463
>TC93594 weakly similar to GP|20466452|gb|AAM20543.1 unknown protein
{Arabidopsis thaliana}, partial (40%)
Length = 754
Score = 224 bits (572), Expect = 4e-59
Identities = 123/243 (50%), Positives = 159/243 (64%), Gaps = 31/243 (12%)
Frame = +1
Query: 217 KKELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLT 276
KKE+V DL F + KE+Y + GK WKRGYLLYGPPGTGKSSL+AA+AN+L YDIYD++LT
Sbjct: 19 KKEIVDDLVTFSKAKEYYAKIGKPWKRGYLLYGPPGTGKSSLVAAIANFLKYDIYDIELT 198
Query: 277 ALNDNKDLKNLLLAMSNRSILVIEDIDCSIKL----------------PNREEDEEAA-- 318
+ +N +L+ LL+ ++++S++VIEDIDCS+ L N E ++ AA
Sbjct: 199 NVKNNAELRKLLIGITSKSVVVIEDIDCSLDLTGQRKTDSENDKEKEEKNEEVNQVAAAS 378
Query: 319 ----------KNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMD 368
KN +++TLSGLLN IDG+WS ER+IIFTTN+ E+LD AL+R GRMD
Sbjct: 379 LQGLQAADKEKNKASQVTLSGLLNFIDGIWSASTGERLIIFTTNYVEKLDQALIRRGRMD 558
Query: 369 MHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQL-TKIA--DVE 425
MH+ LSY F F+ LA YL I H LFE I LL +TPA+VA L K+A DVE
Sbjct: 559 MHIELSYWGFDGFKMLAMNYLSIESHPLFETIQRLLEETNMTPADVAENLMPKVAEEDVE 738
Query: 426 ECL 428
L
Sbjct: 739 ASL 747
>TC87610 similar to GP|9294102|dbj|BAB01954.1 contains similarity to AAA-type
ATPase~gene_id:T20D4.3 {Arabidopsis thaliana}, partial
(53%)
Length = 1765
Score = 221 bits (562), Expect = 6e-58
Identities = 117/250 (46%), Positives = 161/250 (63%), Gaps = 22/250 (8%)
Frame = +3
Query: 229 QGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLL 288
+GK + KAWKRGYLL+GPPGTGKS++I+A+AN+++YD+YDL+LT + DN +LK LL
Sbjct: 756 KGKSIMLKLEKAWKRGYLLFGPPGTGKSTMISAIANFMNYDVYDLELTIVKDNNELKRLL 935
Query: 289 LAMSNRSILVIEDIDCSIKLPNR----------EEDE--------EAAKNGDNKMTLSGL 330
+ S++SI+VIEDIDCS+ L + E DE E + ++K+TLSGL
Sbjct: 936 IETSSKSIIVIEDIDCSLDLTGQRKKKKEKDDVENDEKKDPIKKAEKEEKNESKVTLSGL 1115
Query: 331 LNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLG 390
LN IDG+WS CG ERIIIFTTN ++LDPAL+R GRMD H+ +SYC++ AF+ LA YL
Sbjct: 1116 LNFIDGIWSACGSERIIIFTTNFVDKLDPALIRRGRMDKHIEMSYCSYQAFKVLARNYLD 1295
Query: 391 ISQH-KLFEQIGGLLRHVQVTPAEVAGQL---TKIADVEECLHGLVKFLQDKTQNDQNIN 446
+ H LF I LL +TPA+VA L + D E CL L++ L+ + D+
Sbjct: 1296 VETHDDLFPIIEKLLGETNMTPADVAENLMPKSITEDFESCLKNLIQSLEIAKKKDEEEA 1475
Query: 447 QDNASQKEKK 456
+ +E K
Sbjct: 1476 KKKIEDQEAK 1505
Score = 121 bits (303), Expect = 7e-28
Identities = 75/244 (30%), Positives = 130/244 (52%), Gaps = 10/244 (4%)
Frame = +2
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQG--M 64
+ + S S+ A+ M + ++ F P L ++ + + + E G +
Sbjct: 77 REIWSNLGSIMASIMFVYAMYEKFFPPALRRYLRKYTHKFTNFMYPYIKITFYEKSGDNL 256
Query: 65 TKNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKT-LSFGADRDEEVTDDFEGVRVTWKLT 123
N+ Y + YLS ++ A+R+KA +D + L D ++E+TD+F GV+V W
Sbjct: 257 KHNKTYTTIQTYLSANSSQRARRLKAEVIKDSQNPLVLSMDDNQEITDEFNGVKVWWSAN 436
Query: 124 CIRVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMA 183
I +SRT+ ++ S E R L+FHK+H+E + SY+ +VLE+ KAI +N
Sbjct: 437 HI-----TSRTQSFSIYPS-SDEKRFLTLTFHKRHRELITTSYIQHVLEQGKAITMKNRQ 598
Query: 184 VKIYT-------VEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNR 236
+KIYT Y + + T F+HP +F+TLA++ K+E+++DL +F +GKE+Y +
Sbjct: 599 LKIYTNNPSNDWFRYRSTKWSHTTFEHPASFETLALEPKKKEEILNDLVKFKKGKEYYAK 778
Query: 237 TGKA 240
GK+
Sbjct: 779 VGKS 790
>TC87252 similar to GP|11559424|dbj|BAB18781. mitochondrial protein-like
protein {Cucumis sativus}, partial (72%)
Length = 968
Score = 194 bits (492), Expect(2) = 9e-52
Identities = 113/292 (38%), Positives = 172/292 (58%), Gaps = 11/292 (3%)
Frame = +2
Query: 9 LLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFT-VVIEEFQG--MT 65
L S S+ A+ M + ++ + F P L + LK+ N + + + E G +
Sbjct: 65 LWSQLGSIMASIMFVYAMFDKFFPPNLRVYF-LKYTNKFTNYMYPYIHIKFHELSGERLK 241
Query: 66 KNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKT-LSFGADRDEEVTDDFEGVRVTWKLTC 124
++E Y+ + YLS ++ A+R+KA +D + L D +EE+ D+F GV+V W T
Sbjct: 242 QSETYKIIQTYLSDNSSQRARRLKAEVVKDSQNPLVLSMDDNEEIIDEFNGVKVWW--TA 415
Query: 125 IRVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAV 184
S S Y S E R L+FHKKH+E + SY+ +VL+ K+I +N +
Sbjct: 416 NYTTSKSQSFSYYPTSD----EKRFLTLTFHKKHREVITTSYIQHVLDEGKSIMSKNRQL 583
Query: 185 KIYTVE-------YEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRT 237
K+YT Y + N T F+HP F TLA++ K+E+++DL +F +GKE+Y +
Sbjct: 584 KLYTNNPSSNWWGYRSKKWNHTTFEHPARFGTLAMEPEKKQEILNDLLKFKKGKEYYAKV 763
Query: 238 GKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLL 289
GKAWKRGYLLYGPPGTGKS++I+A+ANY++YD+YDL+LT + DN +LK LL+
Sbjct: 764 GKAWKRGYLLYGPPGTGKSTMISAIANYMNYDVYDLELTTVKDNNELKRLLI 919
Score = 28.1 bits (61), Expect(2) = 9e-52
Identities = 12/17 (70%), Positives = 15/17 (87%)
Frame = +3
Query: 289 LAMSNRSILVIEDIDCS 305
L S++SI+VIEDIDCS
Sbjct: 918 LKRSSKSIIVIEDIDCS 968
>TC88541 weakly similar to GP|10176993|dbj|BAB10225. contains similarity to
AAA-type ATPase~gene_id:MYH19.21 {Arabidopsis thaliana},
partial (26%)
Length = 854
Score = 138 bits (348), Expect = 4e-33
Identities = 79/241 (32%), Positives = 135/241 (55%), Gaps = 2/241 (0%)
Frame = +1
Query: 9 LLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQG--MTK 66
+ + S+ A+ M I +I + P++L + I+ L + EF G + +
Sbjct: 154 MFAQIGSIIASLMFIWAIFQQYFPYQLRNLIDKYSQRLVTFIYPYIQITFHEFTGERLMR 333
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
+E Y + E YLS+KA+ A+R+K ++++++L D EE+ D+F G+++ W + +
Sbjct: 334 SEAYSSIENYLSSKASTQAKRLKGDIAKNNQSLILSMDDKEEICDEFNGMKLWWA-SGKK 510
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
+S+S + + N+ E R Y+L+FHK +++ + YL +VL+ KAI+ +N K+
Sbjct: 511 ASNSNSISLHQNID-----EKRYYKLTFHKHNRDVILGKYLSHVLKEGKAIQVKNRQRKL 675
Query: 187 YTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRGYL 246
YT W F+HP TF+TLA+D K+ ++ DL F + EFY R G+AWKRGYL
Sbjct: 676 YTNSGSHWSH--VVFEHPSTFETLAMDLEKKEMIIDDLITFSKAGEFYARIGRAWKRGYL 849
Query: 247 L 247
L
Sbjct: 850 L 852
>TC89129 weakly similar to GP|17385707|dbj|BAB78658. hypothetical
protein~similar to Arabidopsis thaliana F18B3.210,
partial (7%)
Length = 652
Score = 136 bits (342), Expect = 2e-32
Identities = 68/183 (37%), Positives = 117/183 (63%), Gaps = 1/183 (0%)
Frame = +1
Query: 8 SLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTKN 67
++ SA AS+ A+ ML+RS+ + IP + ++ F L + S T++IEE G+T+N
Sbjct: 103 TIFSAYASMTASIMLLRSMAQELIPQPIRGYLYNTFRYLIKPRSPTLTLIIEESTGITRN 282
Query: 68 EVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRV 127
+VY+AAE+YLSTK T +R+K K +K L+ ++ E++TD + G + W+ C
Sbjct: 283 QVYDAAESYLSTKVTPENERLKISKVPKEKKLTIRLEKGEKLTDIYNGFPLKWRFICAET 462
Query: 128 DSSSSRTRYTNMSS-SLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
+ +S+ + N +S S+ SE + +ELSFHKK+KE +SYLP++L++A+ +K E +K+
Sbjct: 463 EKNSANDMHNNNNSVSVRSEKKYFELSFHKKYKEVGLDSYLPFILDKAQEMKDEXRVLKM 642
Query: 187 YTV 189
+T+
Sbjct: 643 HTL 651
>AW684442 similar to GP|9758309|dbj contains similarity to AAA-type
ATPase~gene_id:MUA2.5 {Arabidopsis thaliana}, partial
(33%)
Length = 600
Score = 86.7 bits (213), Expect = 2e-17
Identities = 51/176 (28%), Positives = 87/176 (48%), Gaps = 1/176 (0%)
Frame = +2
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYN-LSRCFSTQFTVVIEEFQGMT 65
K ++ AS+ ++I P EL F + K +N L CFS+ I E G+
Sbjct: 80 KEYWTSLASILGVFAFFQTILQTVFPPEL-RFASAKLFNKLFNCFSSYCYFEITEIDGVN 256
Query: 66 KNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCI 125
NE+Y A + YLS+ +++ R+ ++ + +FG ++ + D F GV V W+
Sbjct: 257 TNELYNAVQLYLSSSVSITGNRLSLTRAVNSSAFTFGLANNDSIIDTFNGVNVVWEHVVT 436
Query: 126 RVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQEN 181
+ +S + R L E R + L KK K+ + NSYL Y++E+A I+++N
Sbjct: 437 QRNSQTFSWR------PLPDEKRGFTLRIKKKDKQLLXNSYLDYIMEKASDIRRKN 586
>TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AAA-ATPase
subunit RPT4a-like protein {Arabidopsis thaliana},
complete
Length = 1640
Score = 68.9 bits (167), Expect = 4e-12
Identities = 57/186 (30%), Positives = 94/186 (49%), Gaps = 9/186 (4%)
Frame = +2
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTA 277
+EL ++ L E + R G +G LLYGPPGTGK+ L A+A+ + + + +A
Sbjct: 515 RELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIASNIDANFLKVVSSA 694
Query: 278 LNDN--KDLKNLLLAMSNRS------ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
+ D + L+ M + I+ +++ID + R E + + + + TL
Sbjct: 695 IIDKYIGESARLIREMFGYARDHQPCIIFMDEIDA---IGGRRFSEGTSADREIQRTLME 865
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCT-FSAFEQLAFTY 388
LLN +DG + G+ ++I+ TN + LDPALLRPGR+D + + S E L
Sbjct: 866 LLNQLDG-FDQLGKVKMIM-ATNRPDVLDPALLRPGRLDRKIEIPLPNEQSRMEILKIHA 1039
Query: 389 LGISQH 394
GI++H
Sbjct: 1040AGIAKH 1057
>BF647954 homologue to GP|11094192|db 26S proteasome regulatory particle
triple-A ATPase subunit4 {Oryza sativa (japonica
cultivar-group)}, partial (53%)
Length = 649
Score = 67.0 bits (162), Expect = 1e-11
Identities = 55/172 (31%), Positives = 88/172 (50%), Gaps = 9/172 (5%)
Frame = +2
Query: 232 EFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDN--KDLKNLLL 289
E + R G +G LLYGPPGTGK+ L A+A+ + + + +A+ D + L+
Sbjct: 11 ELFLRVGIKPPKGVLLYGPPGTGKTLLARAIASNIDANFLKVVSSAIIDKYIGESSRLIR 190
Query: 290 AMSNRS------ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGE 343
M + I+ +++ID + R E + + + + TL LLN +DG + G+
Sbjct: 191 EMFGYARDHQPCIIFMDEIDA---IGGRRFSEGTSADREIQRTLMELLNQLDG-FDQLGK 358
Query: 344 ERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCT-FSAFEQLAFTYLGISQH 394
+II+ TN + LDPALLRPGR+D + + S E L GI++H
Sbjct: 359 VKIIM-ATNRPDVLDPALLRPGRLDRKIEIPLPNEQSRMEILKIHAAGIAKH 511
>AW692531 weakly similar to GP|11559424|dbj mitochondrial protein-like
protein {Cucumis sativus}, partial (12%)
Length = 662
Score = 66.2 bits (160), Expect = 2e-11
Identities = 53/178 (29%), Positives = 89/178 (49%), Gaps = 4/178 (2%)
Frame = +1
Query: 15 SLAATAMLIRSITNDFIPHE---LIDFINLKFYNLSRCF-STQFTVVIEEFQGMTKNEVY 70
S A+ M I ++ + PH+ LI+ + K N F +F I ++ M NE +
Sbjct: 46 SKVASLMFIWAMIQSYCPHQVHALIEKFSEKLANFCYPFVEVRFFENIGDY--MRTNEAF 219
Query: 71 EAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRVDSS 130
E YL++K+T A++++ K+L D ++ D+FEGV+ W L I +++
Sbjct: 220 LYIENYLTSKSTNQAKKLQGEYVR--KSLVLKMDERQKFYDEFEGVKFVWSLIKIVPNTN 393
Query: 131 SSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKIYT 188
S +S S+ R Y L+FH+ H++ + SYL +VLE+ K I K+YT
Sbjct: 394 S-------VSFYPASDKRFYLLTFHRSHRDFVEKSYLNHVLEQGKEIGLSKRQRKLYT 546
>TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15.70 -
Arabidopsis thaliana, complete
Length = 1629
Score = 64.7 bits (156), Expect = 7e-11
Identities = 53/162 (32%), Positives = 81/162 (49%), Gaps = 8/162 (4%)
Frame = +2
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY-----D 272
+E+ ++ L E Y G +G +LYG PGTGK+ L A+AN S +
Sbjct: 683 QEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANSTSATFLRVVGSE 862
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L L D L L +++ SI+ I++ID + + D + + + T+
Sbjct: 863 LIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDA---VGTKRYDAHSGGEREIQRTMLE 1033
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG S G+ ++I+ TN E LDPALLRPGR+D +
Sbjct: 1034LLNQLDGFDSR-GDVKVIL-ATNRIESLDPALLRPGRIDRKI 1153
>TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulatory subunit
6A homolog (TAT-binding protein homolog 1) (TBP-1)
(Mg(2+), complete
Length = 1793
Score = 64.3 bits (155), Expect = 9e-11
Identities = 70/261 (26%), Positives = 114/261 (42%), Gaps = 13/261 (4%)
Frame = +1
Query: 215 GLKK---ELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAM-----ANYL 266
GL+K ELV + + KE + + G +G LLYGPPGTGK+ + A A +L
Sbjct: 652 GLEKQIQELVEAIVLPMTHKERFQKLGIRPPKGVLLYGPPGTGKTLMARACAAQTNATFL 831
Query: 267 SYDIYDLDLTALNDNKDLKNLLLAMSNRS---ILVIEDIDCSIKLPNREEDEEAAKNGDN 323
L + D L ++ I+ I++ID + + D E + + +
Sbjct: 832 KLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDA---IGTKRFDSEVSGDREV 1002
Query: 324 KMTLSGLLNVIDGLWSCCGEERI-IIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFE 382
+ T+ LLN +DG S ++RI +I TN + LDPAL+R GR+D + + T E
Sbjct: 1003QRTMLELLNQLDGFSS---DDRIKVIAATNRADILDPALMRSGRLDRKIEFPHPT----E 1161
Query: 383 QLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQLTKIADVEECLHGLVKFLQDKTQ-N 441
+ L I K+ + + + G K VE G++ +D T+ N
Sbjct: 1162EARARILQIHSRKMNVHPDVNFEELARSTDDFNGAQLKAVCVEA---GMLALRRDATEVN 1332
Query: 442 DQNINQDNASQKEKKNQDHEY 462
++ N+ + KK Y
Sbjct: 1333HEDFNEGIIQVQAKKKASLNY 1395
>TC77261 homologue to GP|8777330|dbj|BAA96920.1 26S proteasome AAA-ATPase
subunit RPT3 {Arabidopsis thaliana}, partial (94%)
Length = 1575
Score = 63.9 bits (154), Expect = 1e-10
Identities = 47/160 (29%), Positives = 74/160 (45%), Gaps = 5/160 (3%)
Frame = +2
Query: 217 KKELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY----- 271
K+E+ ++ L E Y + G RG LLYGPPGTGK+ L A+AN+ +
Sbjct: 602 KQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGTGKTMLAKAVANHTTAAFIRVVGS 781
Query: 272 DLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLL 331
+ L + + + ++ + I ID + D + + + + L LL
Sbjct: 782 EFVQKYLGEGPRMVRDVFRLAKENAPAIIFIDEVDAIATARFDAQTGADREVQRILMELL 961
Query: 332 NVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
N +DG + +I TN + LDPALLRPGR+D +
Sbjct: 962 NQMDGFDQTVNVK--VIMATNRADTLDPALLRPGRLDRKI 1075
>TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome regulatory
particle triple-A ATPase subunit2b {Oryza sativa
(japonica cultivar-group), partial (98%)
Length = 1696
Score = 63.5 bits (153), Expect = 2e-10
Identities = 52/162 (32%), Positives = 81/162 (49%), Gaps = 8/162 (4%)
Frame = +3
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY-----D 272
+E+ ++ L E Y G +G +LYG PGTGK+ L A+AN S +
Sbjct: 681 QEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANSTSATFLRVVGSE 860
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L L D L L +++ +I+ I++ID + + D + + + T+
Sbjct: 861 LIQKYLGDGPKLVRELFRVADDLSPAIVFIDEIDA---VGTKRYDAHSGGEREIQRTMLE 1031
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG S G+ ++I+ TN E LDPALLRPGR+D +
Sbjct: 1032LLNQLDGFDSR-GDVKVIL-ATNRIESLDPALLRPGRIDRKI 1151
>TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidopsis
thaliana}, partial (98%)
Length = 1633
Score = 62.4 bits (150), Expect = 4e-10
Identities = 48/164 (29%), Positives = 82/164 (49%), Gaps = 10/164 (6%)
Frame = +1
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMAN-----YLSYDIYD 272
KE+ ++ ++ E + G A +G LLYGPPGTGK+ L A+A+ ++ +
Sbjct: 652 KEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLARAVAHHTDCTFIRVSGSE 831
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKM--TL 327
L + + + L M+ SI+ +++ID E + NGD+++ T+
Sbjct: 832 LVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSI----GSARMESGSGNGDSEVQRTM 999
Query: 328 SGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG + + ++ TN + LD ALLRPGR+D +
Sbjct: 1000LELLNQLDGFEA--SNKIKVLMATNRIDILDQALLRPGRIDRKI 1125
>AW684609 similar to GP|11559424|dbj mitochondrial protein-like protein
{Cucumis sativus}, partial (8%)
Length = 653
Score = 60.8 bits (146), Expect = 1e-09
Identities = 54/208 (25%), Positives = 91/208 (42%), Gaps = 11/208 (5%)
Frame = +3
Query: 9 LLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQG--MTK 66
+LS S+AA+ M + ++ F P +L F+ + + S + E G + +
Sbjct: 42 ILSQLGSIAASLMFVYAMYEQFCPSDLRKFVENYKHKFTDLMSPYIQITFNESSGERLKQ 221
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKT-LSFGADRDEEVTDDFEGVRVTWKLTCI 125
+E Y + YL ++ A+R++A ED ++ L D +EE+ D+F GV+V W
Sbjct: 222 SETYTIIQTYLGANSSKRAKRLEAEVVEDSQSPLVLSMDDNEEIEDEFNGVKVWW----- 386
Query: 126 RVDSSSSRTRYTNMSSSLMSEVRSYEL--SFHKKHKEKMFNSYL-----PYVLERAKAIK 178
S++S+ SS RS+++ H +K S+L + KAI
Sbjct: 387 ---SANSKAPRRKASSX-----RSFDVVKMLHPNFPQKASRSHLLPLTYNMCWNKGKAII 542
Query: 179 QENMAVKIYTVEYEAW-CRNGTRFDHPM 205
+ A K YT W CR G P+
Sbjct: 543 FKX*ATKAYTNMGGCWGCRGGXHNFAPL 626
>CB893172 homologue to PIR|T45642|T45 FtsH metalloproteinase-like protein -
Arabidopsis thaliana, partial (25%)
Length = 661
Score = 59.7 bits (143), Expect = 2e-09
Identities = 49/150 (32%), Positives = 71/150 (46%), Gaps = 8/150 (5%)
Frame = +3
Query: 227 FLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMAN-----YLSYDIYDLDLTALNDN 281
FL+ + Y R G RG LL G PGTGK+ L A+A ++S + +
Sbjct: 39 FLRNPDRYARLGARPPRGVLLVGLPGTGKTLLAKAVAGEADVPFISCSASEFVELYVGMG 218
Query: 282 KDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLW 338
L A + + SI+ I++ID K +R+ N + + TL+ LL +DG
Sbjct: 219 ASRVRDLFARAKKEAPSIIFIDEIDAVAK--SRDGKFRIVGNDEREQTLNQLLTEMDGFD 392
Query: 339 SCCGEERIIIFTTNHRERLDPALLRPGRMD 368
S I++ TN + LDPAL RPGR D
Sbjct: 393 S--NSPVIVLAATNRADVLDPALRRPGRFD 476
>BI271823 similar to GP|21592745|gb cell division protein FtsH-like protein
{Arabidopsis thaliana}, partial (33%)
Length = 714
Score = 57.4 bits (137), Expect = 1e-08
Identities = 50/154 (32%), Positives = 70/154 (44%), Gaps = 10/154 (6%)
Frame = +3
Query: 228 LQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDN------ 281
LQG Y + G RG LL GPPGTGK+ L A+A + + + +
Sbjct: 156 LQGDINYQKLGAKLPRGVLLVGPPGTGKTLLARAVAGEAGVPFFTVSASEFVEMFVGRGA 335
Query: 282 ---KDLKNLLLAMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLW 338
+DL + + SI+ I+++D R +EE TL+ LL +DG
Sbjct: 336 ARIRDLFSRARKFA-PSIIFIDELDAVGGKRGRGFNEE------RDQTLNQLLTEMDGFE 494
Query: 339 SCCGEERIIIF-TTNHRERLDPALLRPGRMDMHV 371
S E R+++ TN E LDPAL RPGR V
Sbjct: 495 S---EIRVVVIAATNRPEALDPALCRPGRFSXKV 587
>TC84403 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH
{Pisum sativum}, partial (36%)
Length = 897
Score = 57.0 bits (136), Expect = 2e-08
Identities = 49/155 (31%), Positives = 74/155 (47%), Gaps = 8/155 (5%)
Frame = +3
Query: 243 RGYLLYGPPGTGKSSLIAAMAN-----YLSYDIYD-LDLTALNDNKDLKNLLLAMSN--R 294
+G LL GPPGTGK+ L A A +LS D L+L + ++NL
Sbjct: 18 KGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFLELFVGVGSSRVRNLFKEARKCAP 197
Query: 295 SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHR 354
SI+ I++ID + + A N + + TL+ LL +DG + G +++ TN
Sbjct: 198 SIVFIDEIDAIGRARGSRGGDRA--NDERESTLNQLLVEMDGFGTTAGV--VVLAGTNRP 365
Query: 355 ERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYL 389
+ LD ALLRPGR D + + +Q+ YL
Sbjct: 366 DVLDKALLRPGRFDRQITIDKPDIKGRDQIFQIYL 470
>TC77340 SP|Q9BAE0|FTSH_MEDSA Cell division protein ftsH homolog chloroplast
precursor (EC 3.4.24.-). [Alfalfa] {Medicago sativa},
complete
Length = 2473
Score = 56.6 bits (135), Expect = 2e-08
Identities = 51/177 (28%), Positives = 82/177 (45%), Gaps = 8/177 (4%)
Frame = +3
Query: 205 MTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMAN 264
+TF +A K EL +D FL+ + Y G +G LL GPPGTGK+ L A+A
Sbjct: 999 VTFADVAGADQAKLELQEVVD-FLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAG 1175
Query: 265 YLSYDIYD------LDLTALNDNKDLKNLLLAMSNRS--ILVIEDIDCSIKLPNREEDEE 316
+ ++L +++L +++ I+ I++ID + +
Sbjct: 1176 EAGTPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDA---VGRQRGAGL 1346
Query: 317 AAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNL 373
N + + T++ LL +DG G I++ TN + LD ALLRPGR D V +
Sbjct: 1347 GGGNDEREQTINQLLTEMDGFSGNSGV--IVLAATNRPDVLDSALLRPGRFDRQVTV 1511
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,665,143
Number of Sequences: 36976
Number of extensions: 159290
Number of successful extensions: 985
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 951
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 958
length of query: 467
length of database: 9,014,727
effective HSP length: 100
effective length of query: 367
effective length of database: 5,317,127
effective search space: 1951385609
effective search space used: 1951385609
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0111b.4


