

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111a.1
(863 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
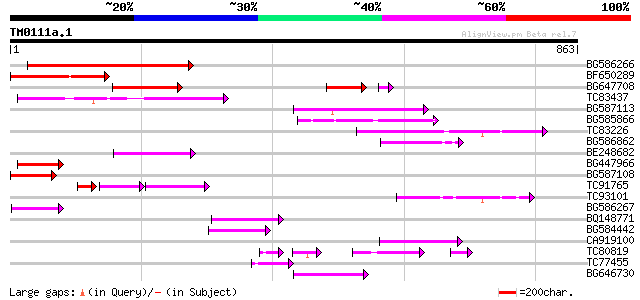
Score E
Sequences producing significant alignments: (bits) Value
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 225 5e-59
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 120 3e-27
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 104 1e-22
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 104 2e-22
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 99 5e-21
BG585866 97 2e-20
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 90 4e-18
BG586862 78 2e-14
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 65 1e-10
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 64 2e-10
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 63 6e-10
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 51 1e-09
TC93101 59 1e-08
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 58 1e-08
BQ148771 58 1e-08
BG584442 57 3e-08
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 53 6e-07
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 46 9e-07
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 50 3e-06
BG646730 50 5e-06
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 225 bits (574), Expect = 5e-59
Identities = 109/254 (42%), Positives = 161/254 (62%)
Frame = -3
Query: 27 ENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQAQSAFISGRQIQDNILIAHEAFH 86
+ + + R I+ CN YK+I KIL +++P + + + +QSAF+ GR I DN+LI H+ H
Sbjct: 784 KRVSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILH 605
Query: 87 FLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFGFSEKWVDRIMQCVTGASFRFK 146
+L+ + K MA+K +M KAYDR+ W+FL+ LT GF W+ IM+CV+ S+ F
Sbjct: 604 YLRQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFL 425
Query: 147 VNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLRMAPECPPITH 206
+NG RV+P RG+RQG PLSPYLF+L E +S L +A +G + G+++A CPPI H
Sbjct: 424 INGGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINH 245
Query: 207 LLFADDSLFFAKANREEAYQLISILNDYTLASGQMISIEKSGIIFSRNTLSSTKHQISSI 266
LLFADD++FF K+N L+SI++ Y ASG+ I+ KS I FS T + ++
Sbjct: 244 LLFADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAITFSSKTSQAIIDRVKGE 65
Query: 267 LRMSAWDSPGKYLG 280
L+++ GKYLG
Sbjct: 64 LKIAKEGGTGKYLG 23
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 120 bits (300), Expect = 3e-27
Identities = 57/151 (37%), Positives = 95/151 (62%)
Frame = +3
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF+ G++P IN T V L+PK ++ RPI+CC+ +YK+I+KIL ++++ +N +
Sbjct: 162 FFKTGFMPKIINCTYVTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVV 341
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
++ QSAF+ GR I DNI+++HE KG + +K+++ KAYD EW F+K +
Sbjct: 342 SENQSAFVKGRVIFDNIILSHELVKSYSR--KGISPRCMVKIDLXKAYDSXEWPFIKHLM 515
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVS 152
GF K+V+ +M +T AS+ F NG+++
Sbjct: 516 LELGFPYKFVNWVMAXLTTASYTFNXNGDLT 608
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 104 bits (260), Expect = 1e-22
Identities = 50/107 (46%), Positives = 73/107 (67%)
Frame = +1
Query: 157 PQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFF 216
P++G+RQG PLSPYLF+L A +S LL + + + G+++A P ITHLLFADDSL F
Sbjct: 4 PEKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLF 183
Query: 217 AKANREEAYQLISILNDYTLASGQMISIEKSGIIFSRNTLSSTKHQI 263
A+AN EA ++ +L+ Y ASGQ+++ EKS + +S+N + K I
Sbjct: 184 ARANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMI 324
Score = 51.6 bits (122), Expect(2) = 7e-08
Identities = 24/61 (39%), Positives = 37/61 (60%)
Frame = +3
Query: 483 LTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYTVKTG 542
L+V+ LI D KQWN + +F A ++ +IPLS+ Q DK+ W KDG Y+V++
Sbjct: 396 LSVDELIDYDTKQWNRDLIFHSFNNYAAHQIIKIPLSMRQPEDKIIWHWEKDGIYSVRSA 575
Query: 543 Y 543
+
Sbjct: 576 H 578
Score = 23.9 bits (50), Expect(2) = 7e-08
Identities = 8/23 (34%), Positives = 11/23 (47%)
Frame = +2
Query: 562 NDPIWNQIWGIQVP*KIKMFLWK 584
N IW IW ++ FLW+
Sbjct: 632 NPVIWKAIWNTPASRSVQNFLWR 700
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 104 bits (259), Expect = 2e-22
Identities = 87/333 (26%), Positives = 142/333 (42%), Gaps = 11/333 (3%)
Frame = +2
Query: 12 INETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQAQSAFISG 71
IN T +AL+PKVD P+ + RPIS LYK++ K+L N+L+ + + + AQSAF+
Sbjct: 50 INSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVVIGSVISDAQSAFVKN 229
Query: 72 RQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFG------ 125
RQI + + + +M ++ K + L W + S G
Sbjct: 230 RQILEMVFL*QMRL------------WMRLR-N*RKIFCCLRWILKRLITLSIGLIWILF 370
Query: 126 -----FSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMS 180
F W I +CV+ A+ VNG S V + + Q + Y F + +
Sbjct: 371 *VGMSFLVLWRKWIKECVSTATTSVLVNG--SPTNVLMKSLVQTQLFTRYSFGVVNPVV- 541
Query: 181 HLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQ 240
++HL FA+D+L N L + L + SG
Sbjct: 542 -----------------------VSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGL 652
Query: 241 MISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRV 300
++ KSG++ N S + +S+L P YLG+P E R + I R+
Sbjct: 653 KVNFHKSGLV-CVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPIVNRI 829
Query: 301 LDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTM 333
++ GW + L+ G+ VL+K+VL ++S Y +
Sbjct: 830 KARLTGWNSRFLSFGGRLVLLKSVLTSLSVYAL 928
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 99.4 bits (246), Expect = 5e-21
Identities = 58/218 (26%), Positives = 98/218 (44%), Gaps = 13/218 (5%)
Frame = -3
Query: 433 SWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDK--WVSDQVLQAPEAGDAELTVNSLIL 490
S+ W+SI + +K+ IIG+G +W+ + W V P + +
Sbjct: 735 SYAWRSIHSAQHLIKQGAKVIIGNGENTNIWEREMAWKLTCVTNHPNKHSSRAY*APTLY 556
Query: 491 ---------PDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYTVKT 541
P ++ N + S FP K+ I D W ++K G Y+VK+
Sbjct: 555 GYEGCRSDDPMRRERNANLINSIFPEGTRRKILSIHPQGPIGEDSYSWEYSKSGHYSVKS 376
Query: 542 GYQVAYSLKEQA--RGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRR 599
GY V ++ A RG D ++ ++W K++ FLW+ + +LP A++ R
Sbjct: 375 GYYVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHFLWRCISNSLPTAANMRSR 196
Query: 600 HIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLH 637
HI+ D C C + E+V H+L CP+ R +W +S +H
Sbjct: 195 HISKDGSCSRCGMESETVNHILFQCPYARLIWATSPIH 82
>BG585866
Length = 828
Score = 97.4 bits (241), Expect = 2e-20
Identities = 64/219 (29%), Positives = 98/219 (44%), Gaps = 5/219 (2%)
Frame = +3
Query: 439 ILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVL--QAP--EAGDAELTVNSLILPDVK 494
I++ K LK W G G+ W W S +L QAP + D LTV + +
Sbjct: 84 IIRDKNVLKSGYTWRAGSGNS-SFWYTNWSSLGLLGTQAPFVDIHDLHLTVKDVFTTGGQ 260
Query: 495 QWNLPKLLSTFPRQIAFKVFQIPLSLT-QVPDKLCWPHTKDGEYTVKTGYQVAYSLKEQA 553
+ L + P IA + L+ + D WPH +G YT K+GY S E
Sbjct: 261 --HTQSLYTILPTDIAEVINNTHLNFNASIGDAYIWPHNSNGVYTAKSGYSWILSQTETV 434
Query: 554 RGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHIAPDPVCFICNQD 613
N+ W+ IW +++P K K FLW AC+ A+P + L R++ +C C +
Sbjct: 435 N------YNNSSWSWIWRLKIPEKYKFFLWLACHNAVPTLSLLNHRNMVNSAICSRCGEH 596
Query: 614 EESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWL 652
EES +H + C +++ +W S +V+DWL
Sbjct: 597 EESFFHCVRDCRFSKIIWHKIGFSSPDFFSSS-SVQDWL 710
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 89.7 bits (221), Expect = 4e-18
Identities = 70/301 (23%), Positives = 132/301 (43%), Gaps = 11/301 (3%)
Frame = +3
Query: 529 WPHTKDGEYTVKTGYQVAYSLK-EQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACN 587
W H G Y+VK+GY + + +Q IW +IW + + K+ LW+ N
Sbjct: 6 WMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSDETLIWKKIWSLHTIPRHKVLLWRILN 185
Query: 588 EALPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLT 647
++LPV++SL +R I P+C C+ E++ HL + CP ++ VW S L + + +
Sbjct: 186 DSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVWFGSNLCINFDNLPNPN 365
Query: 648 VKDWLTQLVIRLQENSEDLSRGLTI-VAYYLWGIWKERNECHFQQRPPDPLGTTIRIRAS 706
+ L + ++ E +TI +A ++ +W RN + + + R
Sbjct: 366 FIN*LYEAIL*KDE-------CITI*IAAIIYNLWHARNLSVLEDQTILEMDIIQRASNC 524
Query: 707 MLNGMEDNTPSP--------IPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIA 758
+ + + NT +P P +P +W P +K+N DA + K +
Sbjct: 525 ISDYKQANTQAPPSMARTGYDPRSQHRPAKNTKWKRPNLGLVKVNTDAN-LQNHGKWGLG 701
Query: 759 VVVFDEKGTNVLSHS-KTIWAASPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLI 817
+++ DE G + + + +T L AEA A+ + A + K+ D L+ ++
Sbjct: 702 IIIRDEVGLVMAASTWETDGNDRALEAEAYALLTGMRFAKDCGFXKVXFEGDNEKLMKMV 881
Query: 818 R 818
+
Sbjct: 882 K 884
>BG586862
Length = 804
Score = 77.8 bits (190), Expect = 2e-14
Identities = 44/126 (34%), Positives = 64/126 (49%)
Frame = -1
Query: 565 IWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGC 624
I ++WGI+ + K FLW+ + ALPVK L +R I +C C E+V HL L C
Sbjct: 666 IREKVWGIKTIPRHKSFLWRLLHNALPVKDELHKRGIRCSLLCPRCESKIETVQHLFLNC 487
Query: 625 PWTRAVWLSSQLHVQHLHQGSLTVKDWLTQLVIRLQENSEDLSRGLTIVAYYLWGIWKER 684
T+ W SQL + G L DW+T +++ N E+ LT + L+ IW R
Sbjct: 486 EVTQKEWFGSQLGINFHSSGVLHFHDWITNFILK---NDEETIIALTAL---LYSIWHAR 325
Query: 685 NECHFQ 690
N+ F+
Sbjct: 324 NQKVFE 307
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 65.1 bits (157), Expect = 1e-10
Identities = 40/125 (32%), Positives = 61/125 (48%)
Frame = +3
Query: 158 QRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFA 217
QRG++QG PL+P+LF+L AE +S L+ A QG + ++HL +ADD+L
Sbjct: 21 QRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADDTLCIG 200
Query: 218 KANREEAYQLISILNDYTLASGQMISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGK 277
+ + L ++L + +ASG ++ KS +I N L P
Sbjct: 201 MPTVDNLWTLKALLQGFEMASGLKVNFHKSSLI-GINVPRDFMEAACRFLNCREESIPFI 377
Query: 278 YLGLP 282
YLGLP
Sbjct: 378 YLGLP 392
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 63.9 bits (154), Expect = 2e-10
Identities = 30/70 (42%), Positives = 43/70 (60%)
Frame = +2
Query: 12 INETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQAQSAFISG 71
+N+T + L+PK P RPIS CN + K+ITK++ N++K + + QSAF+ G
Sbjct: 425 LNKTFIVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIANRVKQTLPDVIDVEQSAFVQG 604
Query: 72 RQIQDNILIA 81
R I DN LIA
Sbjct: 605 RLITDNALIA 634
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 62.8 bits (151), Expect = 6e-10
Identities = 30/71 (42%), Positives = 44/71 (61%)
Frame = +3
Query: 1 SFFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRL 60
SF G L +N T + L+PK +P + +LRPIS CN YK+I+K+L +LK + L
Sbjct: 465 SFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGYKIISKVLCQRLKVCLPSL 644
Query: 61 ATQAQSAFISG 71
++ QSAF+ G
Sbjct: 645 ISETQSAFVHG 677
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 51.2 bits (121), Expect(2) = 1e-09
Identities = 23/69 (33%), Positives = 38/69 (54%)
Frame = +2
Query: 137 CVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLR 196
CV + VN + ++P RG++QG LSPY+F++ E +S L+ A+E G G
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKERGDTHGTS 283
Query: 197 MAPECPPIT 205
+ PP++
Sbjct: 284 I*RGAPPVS 310
Score = 44.3 bits (103), Expect = 2e-04
Identities = 35/97 (36%), Positives = 45/97 (46%)
Frame = +3
Query: 207 LLFADDSLFFAKANREEAYQLISILNDYTLASGQMISIEKSGIIFSRNTLSSTKHQISSI 266
LLF L + A + +IL Y SG+ IS+ KS I SRN K I+ I
Sbjct: 279 LLFEGAPLRSLRVEEHHAQIMKNILILYEEDSGKAISLRKS*IYCSRNVPDILKTSITYI 458
Query: 267 LRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVLDK 303
L + KYLGLP+ G+ R T + IK V K
Sbjct: 459 LGVQFMLGTCKYLGLPSMIGRDRTTTFSSIKGGVWQK 569
Score = 30.4 bits (67), Expect(2) = 1e-09
Identities = 10/29 (34%), Positives = 20/29 (68%)
Frame = +1
Query: 103 LEMNKAYDRLEWDFLKACLTSFGFSEKWV 131
L ++K Y+R++ D+LK + GF+ +W+
Sbjct: 1 LYISKVYNRVD*DYLKEIMIKMGFNNRWI 87
>TC93101
Length = 675
Score = 58.5 bits (140), Expect = 1e-08
Identities = 52/225 (23%), Positives = 91/225 (40%), Gaps = 15/225 (6%)
Frame = -1
Query: 590 LPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVK 649
LPV+ L +R + P+C C + E+ H+ + C T+ VW SQL ++ ++
Sbjct: 663 LPVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVWFGSQLSIRFPDNSTINFS 484
Query: 650 DWLTQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRASMLN 709
DWL + +E++ ++ + Y IW RN+ F+ + T I+ + +
Sbjct: 483 DWLFDAI---SNQTEEIIIKISAITY---SIWHARNKAIFENQFVSE-DTIIQ*AQNSIL 325
Query: 710 GMEDNTPSP---------------IPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCK 754
E T P T R+ N ++RW P LK N DA + +
Sbjct: 324 AYEQATIKPQNPNIVLSSLSATSNTNTTRRRSNVRSRWQKPLNNILKANCDAN-LQVQGR 148
Query: 755 GSIAVVVFDEKGTNVLSHSKTIWAASPLAAEALAVREALLLAMNL 799
+ ++ + G ++ W + A A A+L AM L
Sbjct: 147 WGLGCIIRNADGEAKVT---ATWCINGFDCAATAETYAILAAMYL 22
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 58.2 bits (139), Expect = 1e-08
Identities = 30/79 (37%), Positives = 47/79 (58%)
Frame = +1
Query: 4 QEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQ 63
++G L + LVPK + + + + RPIS CN YK+++K+L +LK + + T+
Sbjct: 556 EQGNLKRESTKQTSGLVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSVLPWIITE 735
Query: 64 AQSAFISGRQIQDNILIAH 82
Q+AF + I DNILIAH
Sbjct: 736 TQAAFGRRQLISDNILIAH 792
>BQ148771
Length = 680
Score = 58.2 bits (139), Expect = 1e-08
Identities = 33/111 (29%), Positives = 50/111 (44%)
Frame = -3
Query: 307 WKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHW 366
WK L+ A + L K+V++A+ Y M P+ + I +F W R H
Sbjct: 564 WKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTEVSRRYHA 385
Query: 367 KDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLY 417
W +SK K GLG + MN A + K W IY+ ++L ++G Y
Sbjct: 384 VGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRGKY 232
>BG584442
Length = 775
Score = 57.0 bits (136), Expect = 3e-08
Identities = 35/96 (36%), Positives = 52/96 (53%), Gaps = 1/96 (1%)
Frame = +1
Query: 303 KIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKER 362
KI+ + K L++ EV+IK LQ++S+Y MSI + I + F W G+ R
Sbjct: 382 KINF*RNKCLSKVM*EVMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVGENR 561
Query: 363 -GIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQ 397
G+HW + K GG+GF DF+T N+ +L KQ
Sbjct: 562 KGMHWMS*EKLFVHKNYGGMGFTDFTTFNIPMLGKQ 669
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 52.8 bits (125), Expect = 6e-07
Identities = 35/128 (27%), Positives = 52/128 (40%), Gaps = 2/128 (1%)
Frame = -2
Query: 564 PIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHIAPDP--VCFICNQDEESVYHLL 621
P IW QVP K+ +F W+ + LP K++L R + P +C ES HL
Sbjct: 599 PYAEMIWHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGALESAQHLF 420
Query: 622 LGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLTQLVIRLQENSEDLSRGLTIVAYYLWGIW 681
L C + ++W + + + + + D Q V N S I W +W
Sbjct: 419 LSCSYFASLWSLVRDWIGFVGVDTNVLSDHFVQFVHSTGGNKASQSFLQLIWLLCAWVLW 240
Query: 682 KERNECHF 689
ERN F
Sbjct: 239 TERNNMCF 216
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 46.2 bits (108), Expect(2) = 9e-07
Identities = 31/111 (27%), Positives = 47/111 (41%), Gaps = 2/111 (1%)
Frame = +2
Query: 523 VPDKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFL 582
V DK W Y+VK Y+ S HI + + + +W +P K+ +F+
Sbjct: 512 VNDKWRWLLDPVNGYSVKVFYRYITSTG--------HISDRSLVDDVWHKHIPSKVSLFV 667
Query: 583 WKACNEALPVKASLFRRHI--APDPVCFICNQDEESVYHLLLGCPWTRAVW 631
W+ LP K +L R + A + C D ES HL L C ++W
Sbjct: 668 WRLLRNRLPTKDNLVHRGVLLATNAACVCGCVDSESTTHLFLHCNVFCSLW 820
Score = 33.1 bits (74), Expect(2) = 7e-04
Identities = 17/38 (44%), Positives = 20/38 (51%)
Frame = +1
Query: 380 GLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLY 417
GLG F NL+LL K WR+ + LW R LK Y
Sbjct: 31 GLGVGAF---NLSLLGKWCWRLLVDKEGLWHRVLKARY 135
Score = 28.5 bits (62), Expect(2) = 7e-04
Identities = 15/52 (28%), Positives = 26/52 (49%), Gaps = 8/52 (15%)
Frame = +3
Query: 431 RNSWGWQSILQGKEFLKEA-GLW-------IIGDGSEVKVWKDKWVSDQVLQ 474
++S W++I + +E + E G W ++GDG + W D W D L+
Sbjct: 171 QSSMWWKTICKVREGVGEGVGNWFEENIRMVVGDGRDAFFWYDTWAGDVPLR 326
Score = 25.4 bits (54), Expect(2) = 9e-07
Identities = 13/33 (39%), Positives = 15/33 (45%)
Frame = +1
Query: 672 IVAYYLWGIWKERNECHFQQRPPDPLGTTIRIR 704
I+ W IWKERN FQ P RI+
Sbjct: 928 ILQVNFWVIWKERNNRLFQNTVTAPFVLIDRIK 1026
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 50.4 bits (119), Expect = 3e-06
Identities = 25/65 (38%), Positives = 37/65 (56%), Gaps = 2/65 (3%)
Frame = -3
Query: 369 WHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFP--NSSFLQS 426
WH + + CKGGLG +D +N++LLAK WR+ +LW L+ +Y P +S +
Sbjct: 975 WHCLPR--CKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYGPRVSSRTMMV 802
Query: 427 GRTAR 431
GR R
Sbjct: 801 GRIGR 787
Score = 39.7 bits (91), Expect = 0.005
Identities = 24/101 (23%), Positives = 45/101 (43%), Gaps = 5/101 (4%)
Frame = -2
Query: 529 WPHTKDGEYTVKTGYQVAYSLK--EQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKAC 586
W K+G ++V + Y + +L+ E + + I+ ++W + P K+ F W
Sbjct: 436 WKPDKEGVFSVNSCYFLLQNLRLLEDRLSYEEEV----IFRELWKSKAPAKVLAFSWTLF 269
Query: 587 NEALPVKASLFRR---HIAPDPVCFICNQDEESVYHLLLGC 624
+ +P +L +R + C C +E+V HL L C
Sbjct: 268 LDRIPTMVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHC 146
>BG646730
Length = 799
Score = 49.7 bits (117), Expect = 5e-06
Identities = 30/117 (25%), Positives = 56/117 (47%), Gaps = 4/117 (3%)
Frame = +1
Query: 433 SWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQV---LQAPEAGDAELT-VNSL 488
S+ W+S+ K+ + W I +G V++WKD W+ +Q + +P E T ++ L
Sbjct: 400 SYAWRSMFNIKDVIDLGSRWSISNGQNVRIWKDDWLPNQTGFKVCSPMVDFEEATCISEL 579
Query: 489 ILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYTVKTGYQV 545
I + K + + F A ++ IPLS PD+ +D Y+V++ + +
Sbjct: 580 IDTNTKSSKRDLVRNVFSAFEAKQILNIPLSWRLWPDRRILYWERDRNYSVRSAHHL 750
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,964,970
Number of Sequences: 36976
Number of extensions: 513492
Number of successful extensions: 2458
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 2414
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2445
length of query: 863
length of database: 9,014,727
effective HSP length: 104
effective length of query: 759
effective length of database: 5,169,223
effective search space: 3923440257
effective search space used: 3923440257
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0111a.1


