

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.16
(1427 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
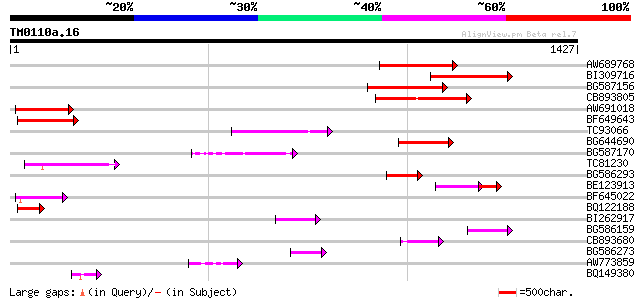
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 214 1e-55
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 210 3e-54
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 171 2e-42
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 169 7e-42
AW691018 weakly similar to GP|2462935|em open reading frame 1 {B... 145 1e-34
BF649643 similar to PIR|H86486|H86 protein Ty1/copia-element pol... 141 2e-33
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 129 6e-30
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 126 7e-29
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 125 9e-29
TC81230 101 2e-21
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 89 9e-18
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 88 3e-17
BF645022 weakly similar to GP|8777581|dbj retroelement pol polyp... 85 2e-16
BQ122188 weakly similar to GP|9758772|dbj| contains similarity t... 79 1e-14
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 75 1e-13
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 74 4e-13
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 72 2e-12
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 70 4e-12
AW773859 70 8e-12
BQ149380 65 2e-10
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 214 bits (546), Expect = 1e-55
Identities = 101/195 (51%), Positives = 136/195 (68%)
Frame = +1
Query: 931 PEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGY 990
P W QAM+TE A+ N+TWD+V LPP K IG KWVY++K +PDGS+++ KARLVAKG+
Sbjct: 91 PRWLQAMKTEYKALIDNKTWDLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKGF 270
Query: 991 TQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQG 1050
+Q G DY +TFSPV K T+R++L +A YKWE+ Q+DINNAFLNG L EEVYM PQG
Sbjct: 271 SQTLGCDYTETFSPVIKPVTIRLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQPQG 450
Query: 1051 YDVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVL 1110
++ ++L C+L KS+YGLKQA R W+ +S Q GF S D SL + L
Sbjct: 451 FEAANKSLVCKLNKSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLIYNQNGACIYL 630
Query: 1111 LVYVNDIIVSGPNAT 1125
+YV+DI+++G +A+
Sbjct: 631 XIYVDDILITGSSAS 675
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 210 bits (535), Expect = 3e-54
Identities = 106/205 (51%), Positives = 139/205 (67%)
Frame = +2
Query: 1060 CRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIV 1119
C L+KSIYGLKQASRQW++K S L G+ S D+SLFTK +F LLVYV+DI++
Sbjct: 23 CELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDIVL 202
Query: 1120 SGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGFT 1179
+G + +EI VK L FK+KDLG L++FLGLE+A S GI L+QR Y L LL D+G
Sbjct: 203 AGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDSGNL 382
Query: 1180 DCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSKP 1239
+ T P D +LKL + P D +QYRRLIG+L+YLT +RPDI+F + +LSQF+SKP
Sbjct: 383 AVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFVSKP 562
Query: 1240 TTTHLDALHHLLRYLKTTPGQGFFF 1264
H A +L+YLKT P +G F+
Sbjct: 563 QQVHYQAAIRVLQYLKTAPAKGLFY 637
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 171 bits (432), Expect = 2e-42
Identities = 85/202 (42%), Positives = 131/202 (64%), Gaps = 2/202 (0%)
Frame = -1
Query: 901 YDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEANQTWDVVSLPPGKH 960
+ + ++ + V + ++ P + +A++ EW++++ E AM N TW LP GK
Sbjct: 618 FSQYPEAHCAFMVNLNENHIPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKK 439
Query: 961 EIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAM 1020
+ S+W++ +K DGSI+R K RLVA+G+T G DY++TF+PVAK+ T+R++L+LA
Sbjct: 438 AVSSRWIFTIKYKADGSIERKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVN 259
Query: 1021 YKWELFQLDINNAFLNGDLFEEVYMKIPQGYD--VKGENLTCRLKKSIYGLKQASRQWFA 1078
W L+Q+D+ NAFL G+L +EVYM P G + VK N+ RLKK+IYGLKQ+ R W+
Sbjct: 258 LGWGLWQMDVKNAFLQGELEDEVYMYPPPGLEHLVKRGNV-LRLKKAIYGLKQSPRAWYN 82
Query: 1079 KFSSVLTQHGFQHSYHDYSLFT 1100
K S+ L GF+ S D++LFT
Sbjct: 81 KLSTTLNGRGFRKSELDHTLFT 16
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 169 bits (428), Expect = 7e-42
Identities = 87/245 (35%), Positives = 149/245 (60%), Gaps = 3/245 (1%)
Frame = +3
Query: 920 EPTYFHQAIQYPEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSID 979
+PT F +A++ +W+ +M E+ A E N TW++ L G IG KW++K KL+ +G I+
Sbjct: 39 DPTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKTKLNENGEIE 218
Query: 980 RHKARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDL 1039
++KARLVAKGY+QQ G+DY + F+PVA+ T+RM++ALAA K + + I+
Sbjct: 219 KYKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAAQIKRD--GVCIS*M*KAHSC 392
Query: 1040 FEEVYMK--IPQGYDVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYS 1097
E K + + ++ R+K+++YGLKQA R W+++ + T+ GF+ ++++
Sbjct: 393 MEN*MRKFLLINHRVM*RRVIS*RVKRALYGLKQAPRAWYSRIEAYFTKEGFEKCPYEHT 572
Query: 1098 LFTK-GSGDTFVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAY 1156
LF K G +++ +YV+D+I G + + K+ +K F + DLGK+ +FLG+E+
Sbjct: 573 LFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVTQ 752
Query: 1157 SATGI 1161
+ GI
Sbjct: 753 NEKGI 767
>AW691018 weakly similar to GP|2462935|em open reading frame 1 {Brassica
oleracea}, partial (12%)
Length = 639
Score = 145 bits (365), Expect = 1e-34
Identities = 68/145 (46%), Positives = 99/145 (67%), Gaps = 1/145 (0%)
Frame = +2
Query: 16 RIMENS-SNPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRP 74
+ ME S +P LHHSD PG+ LV+QPLNG NY W+RSM+++LS KNKLG I+G P
Sbjct: 128 KAMETSLPDPLLLHHSDTPGISLVNQPLNGGNYGEWSRSMLLSLSAKNKLGLIDGTVKAP 307
Query: 75 ADDDPLLQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQ 134
+ DDP L W R + +V++WIL+ + DI SVI +A +W DL DRF Q + +RIYQ
Sbjct: 308 SADDPKLPLWKRCNDLVLTWILHSIEPDIARSVIFSDTAAAVWSDLHDRFSQGDESRIYQ 487
Query: 135 LKKDLLNLQQESLTVTQFYTKHKAI 159
+++++ +Q SL ++ +YTK K++
Sbjct: 488 IRQEISECRQGSLLISDYYTKLKSL 562
>BF649643 similar to PIR|H86486|H86 protein Ty1/copia-element polyprotein
[imported] - Arabidopsis thaliana, partial (2%)
Length = 608
Score = 141 bits (356), Expect = 2e-33
Identities = 64/155 (41%), Positives = 104/155 (66%), Gaps = 2/155 (1%)
Frame = +1
Query: 20 NSSNPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPADDDP 79
+SSNP+++H SD+PG +LV LNG NY+SW+RSM+ AL+ KNK+GFING P++ D
Sbjct: 109 DSSNPYYIHPSDHPGHLLVPTKLNGTNYSSWSRSMVHALTAKNKVGFINGSIKTPSEVDQ 288
Query: 80 LLQS--WIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKK 137
+ W R + +++SW+ + V D+ VI ++A ++W+D +D+F Q N IYQ++K
Sbjct: 289 PAEYALWNRCNSMILSWLTHSVEPDLAKGVIHAKTACQVWEDFKDQFSQKNIPAIYQIQK 468
Query: 138 DLLNLQQESLTVTQFYTKHKAIWDELQDYLPQRSC 172
L +L Q +++ + ++TK K +WDEL+ Y +C
Sbjct: 469 SLASLSQGTVSASTYFTKIKGLWDELETYRTLPTC 573
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 129 bits (325), Expect = 6e-30
Identities = 83/260 (31%), Positives = 123/260 (46%), Gaps = 7/260 (2%)
Frame = +1
Query: 559 QKRLPFVSENHFANHSFDLIHCDVWGPFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSE 618
+K++ F + H D IH D+WGP V ++ G R+ +TI+DD R W + +++K+E
Sbjct: 49 RKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRYKNE 228
Query: 619 APLAIMAFFKMVQTQFGKVIKMVRTDNAKEL---QLTAFLEKQGTLHQFSCVERPQQNSV 675
+ +V+TQ GK +K + TDN E F G + PQQN V
Sbjct: 229 TFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQNGV 408
Query: 676 VERRHQQLLNVARSLLFQSHM--PLVFWGECISTATHLINLIPTPNLHNKSPSTLLYQQD 733
ER + LL AR +L + + W E STA HL+N P L K P +
Sbjct: 409 AERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWSGNL 588
Query: 734 PSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGYKGYKLYNLTTKT--FHISR 791
Y +LR+FGC ++ K +PRA C+ + Y KGY+L+ K+ +SR
Sbjct: 589 VDYSNLRIFGCPAYALV---NDGKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLILSR 759
Query: 792 DVVFNESIFPFQNTRPYVYS 811
DV FNE + +V S
Sbjct: 760 DVTFNEDALLSSGKQSFVSS 819
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 126 bits (316), Expect = 7e-29
Identities = 62/141 (43%), Positives = 96/141 (67%), Gaps = 1/141 (0%)
Frame = -2
Query: 978 IDRHKARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNG 1037
I R+K++LV +GY Q+ GIDY + FSPVA++ +R+L+A AA ++L+Q+D+ +AF+NG
Sbjct: 424 ITRNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFING 245
Query: 1038 DLFEEVYMKIPQGY-DVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDY 1096
DL EEV++K P G+ D + N RL K++YGLKQA R W+ + S L ++GF+ D
Sbjct: 244 DLKEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDN 65
Query: 1097 SLFTKGSGDTFVVLLVYVNDI 1117
+LF +++ VYV+DI
Sbjct: 64 TLFLLKRE*ELLIIQVYVDDI 2
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 125 bits (315), Expect = 9e-29
Identities = 96/271 (35%), Positives = 135/271 (49%), Gaps = 4/271 (1%)
Frame = -3
Query: 457 VIQENKLKTIGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTA 516
VI+ ++L IG G T LY L P+S N C+ + S NK
Sbjct: 713 VIESSQL--IGEGVTKGDLYMLE---------KLDPVS--NYKCSFTS-SSSLNKD---- 588
Query: 517 ALWHARLGHPSYNRLSLLSSTIHCKIPSSINENV-CPVCPLAKQKRLPFVSENHFANHSF 575
ALWHARLGHP L+L+ +P + EN C C L K + F + + F
Sbjct: 587 ALWHARLGHPHGRALNLM-------LPGVVFENKNCEACILGKHCKNVFPRTSTVYENCF 429
Query: 576 DLIHCDVWGPFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFG 635
DLI+ D+W + + H++F+T +D+ S++TW LI K A F V +
Sbjct: 428 DLIYTDLWTAPSL-SRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYH 252
Query: 636 KVIKMVRTDNAKELQLTAF---LEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLF 692
IK++R+DN E AF L+ G LHQ SC PQQN V +R+++ L+ VARSL+F
Sbjct: 251 AKIKILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMF 72
Query: 693 QSHMPLVFWGECISTATHLINLIPTPNLHNK 723
Q++ +STA +LIN IPT L K
Sbjct: 71 QAN---------VSTACYLINWIPTKVLRIK 6
>TC81230
Length = 958
Score = 101 bits (252), Expect = 2e-21
Identities = 70/248 (28%), Positives = 110/248 (44%), Gaps = 9/248 (3%)
Frame = +1
Query: 38 VSQPLNGENYNSWNRSMMIALSVKNKLGFINGD--FPRPADDDPL------LQSWIRNDH 89
+S LNG NYN W SM L + ++ GD P DD L+ W +H
Sbjct: 226 ISIILNGSNYNHWAESMCGFLKGRRLWRYVTGDKKCPTKGKDDTADAFADKLEEWDSKNH 405
Query: 90 VVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLLNLQQES-LT 148
+++W N I +AKE+W L+ R+ + + YQL KDL NL+Q+S
Sbjct: 406 QIITWFRNTSIPSIHMQFGRFENAKEVWDHLKQRYTISDLSHQYQLLKDLSNLKQQSGQP 585
Query: 149 VTQFYTKHKAIWDELQDYLPQRSCVCGENRVLLDYFQQEQIMHFLMSLSDSYNQIKSHVL 208
V +F + + IW++L P + + + + +++ FLM+L+D Y +++ L
Sbjct: 586 VYEFLAQMEVIWNQLTSCEPSLKDAT-DMKTYETHRNRVRLIQFLMALTDEYEPVRASSL 762
Query: 209 LMNPLPPMNRVFSMVLQEEKQREIAARSADLNSAFAAQATNSGKSGKKDRDRPLCSYCGK 268
NPLP + + EE + ++ ADL A AT C +C K
Sbjct: 763 HQNPLPTLENALPCLKSEETRLQLVPPKADLAFAVTNNATKP------------CRHCQK 906
Query: 269 LGHSVDRC 276
GHS C
Sbjct: 907 SGHSFSDC 930
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 89.4 bits (220), Expect = 9e-18
Identities = 43/90 (47%), Positives = 62/90 (68%)
Frame = +2
Query: 948 QTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAK 1007
QT +V P G IG +W+YK+K + DG++ ++KARLVAKGY +Q GID+ + F+PV +
Sbjct: 53 QTLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPVVR 232
Query: 1008 ITTVRMLLALAAMYKWELFQLDINNAFLNG 1037
I T+ +LLALAA + +D+ AFLNG
Sbjct: 233 IETI*LLLALAATNGC*IHHIDVKIAFLNG 322
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 87.8 bits (216), Expect = 3e-17
Identities = 47/122 (38%), Positives = 71/122 (57%), Gaps = 1/122 (0%)
Frame = +1
Query: 1071 QASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTF-VVLLVYVNDIIVSGPNATEIDS 1129
Q+ R WF +F+ V+ + G+ D+++F K S +L+VYV+DI ++G + I
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 1130 VKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSLPMD 1189
+K LL F++KDLG LK+FLG+E+A G S+SQR Y L LL +T C+ P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDPYG 360
Query: 1190 PN 1191
N
Sbjct: 361 CN 366
Score = 56.2 bits (134), Expect = 9e-08
Identities = 24/57 (42%), Positives = 39/57 (68%)
Frame = +2
Query: 1180 DCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFL 1236
D +P+ PMD +KL + L D +Y+RL+G+L+YL+ +RPDI+F + +SQF+
Sbjct: 332 DVKPSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQFM 502
>BF645022 weakly similar to GP|8777581|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (8%)
Length = 685
Score = 84.7 bits (208), Expect = 2e-16
Identities = 44/136 (32%), Positives = 72/136 (52%), Gaps = 7/136 (5%)
Frame = -2
Query: 16 RIMENSSNP-------FFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFIN 68
++ +NSSN FFL +DNPG ++ L G NY+ W+R++ + K K GF+
Sbjct: 423 KVDDNSSNSEKKVDLVFFLGSNDNPGNVITPIQLRGLNYDEWSRAIRTSFQAKRKYGFVE 244
Query: 69 GDFPRPADDDPLLQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPN 128
G P+P + L+ W ++++W+LN + + S++ A+ +W L+ RF N
Sbjct: 243 GKIPKPTTPEK-LEDWKAVQSMLIAWLLNTIEPSLRSTLSYYDDAESLWTHLKQRFCVVN 67
Query: 129 GTRIYQLKKDLLNLQQ 144
G RI QLK L +Q
Sbjct: 66 GARICQLKASLGECKQ 19
>BQ122188 weakly similar to GP|9758772|dbj| contains similarity to
retroelement pol polyprotein~gene_id:MNJ7.3 {Arabidopsis
thaliana}, partial (7%)
Length = 708
Score = 79.0 bits (193), Expect = 1e-14
Identities = 37/68 (54%), Positives = 48/68 (70%), Gaps = 1/68 (1%)
Frame = -3
Query: 20 NSSNPFFLHHSDNPGLILVSQPLNG-ENYNSWNRSMMIALSVKNKLGFINGDFPRPADDD 78
N+SNPFFLH S+NP + LV+ PL G +N+ SW RSM +AL+ KNK+ F++G F PA D
Sbjct: 205 NASNPFFLHPSENPAVELVTPPLEGNKNFQSWIRSMRLALTSKNKIAFVDGTFLTPAKSD 26
Query: 79 PLLQSWIR 86
L WIR
Sbjct: 25 ALYNQWIR 2
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 75.5 bits (184), Expect = 1e-13
Identities = 38/112 (33%), Positives = 58/112 (50%)
Frame = +1
Query: 670 PQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHLINLIPTPNLHNKSPSTLL 729
PQQN V ER ++ LL R++L + M FW E + TA ++IN P+ + K+P +
Sbjct: 88 PQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPMEMW 267
Query: 730 YQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGYKGYKLYN 781
+ Y L VFGC + R K P++ C+ +GY KGY L++
Sbjct: 268 KGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 73.9 bits (180), Expect = 4e-13
Identities = 35/113 (30%), Positives = 62/113 (53%)
Frame = +1
Query: 1152 LEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQYRRL 1211
+E+ + GI + QR Y LL G + P+ P KL D D+++Y+++
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 1212 IGRLLYLTISRPDIAFTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQGFFF 1264
+G L+YL +RPD+ + ++ +S+F++ PT H+ A+ +LRYL T G +
Sbjct: 181 VGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMY 339
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 72.0 bits (175), Expect = 2e-12
Identities = 39/108 (36%), Positives = 62/108 (57%)
Frame = -2
Query: 983 ARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEE 1042
+R++AKG + F P+ K+ T+ LL++ A+ L LD+ AFL GDL E+
Sbjct: 593 SRILAKGRI------LL*NFVPIVKLNTIMFLLSIVAIENLYLE*LDVKTAFLRGDLVED 432
Query: 1043 VYMKIPQGYDVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQ 1090
+YM P+G+ + + +LKKS+YGLKQ RQ ++ T+ GF+
Sbjct: 431 IYMHQPEGFS*EVGKMVGKLKKSMYGLKQGPRQCI*SLKALCTRKGFR 288
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 70.5 bits (171), Expect = 4e-12
Identities = 36/90 (40%), Positives = 52/90 (57%)
Frame = -2
Query: 708 ATHLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVL 767
A +LIN IPT L +++P +L Q+ PS ++RVFGCLC+ R+K R+ +
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 768 VGYKQGYKGYKLYNLTTKTFHISRDVVFNE 797
+GY KGYK Y+ + +SRDV F E
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIE 435
>AW773859
Length = 538
Score = 69.7 bits (169), Expect = 8e-12
Identities = 46/143 (32%), Positives = 73/143 (50%), Gaps = 5/143 (3%)
Frame = -3
Query: 449 IHFFH--KTFVIQENK-LKTIGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVF 505
+ +FH + +IQE + LK IG G+ GLY+L T ++ +S
Sbjct: 413 VRWFHTLQVILIQEKRSLKMIGSGELFEGLYFLTTQD-------TPSATIASSQVQPQPQ 255
Query: 506 PSHCNKSTCTAALWHARLGHPSYNRLSLLSSTIHCKIPS-SINEN-VCPVCPLAKQKRLP 563
P+ + ALWH RLGH S +L ++H P +I++N VC +C ++ K+LP
Sbjct: 254 PTFLPQE----ALWHFRLGHLSNRKLL----SLHSNFPFITIDQNSVCDICHYSRHKKLP 99
Query: 564 FVSENHFANHSFDLIHCDVWGPF 586
F + A+ ++L H D+WGPF
Sbjct: 98 FQLSTNRASKCYELFHFDIWGPF 30
>BQ149380
Length = 419
Score = 65.1 bits (157), Expect = 2e-10
Identities = 33/84 (39%), Positives = 49/84 (58%), Gaps = 9/84 (10%)
Frame = +3
Query: 157 KAIWDELQDYLPQRSCVCG---------ENRVLLDYFQQEQIMHFLMSLSDSYNQIKSHV 207
+ +WDEL Y P+ C C N+ L +++I+ FLM L+D Y + S V
Sbjct: 75 RGLWDELDQYRPKPQCTCPIQCTCFAMRNNKALT---AEDRIIKFLMGLNDDYQGVTSKV 245
Query: 208 LLMNPLPPMNRVFSMVLQEEKQRE 231
LLM PLP +N VFSMV+Q+E++ +
Sbjct: 246 LLMYPLPQINMVFSMVMQQERKMQ 317
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.136 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,723,418
Number of Sequences: 36976
Number of extensions: 934333
Number of successful extensions: 5644
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 5364
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5601
length of query: 1427
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1319
effective length of database: 5,021,319
effective search space: 6623119761
effective search space used: 6623119761
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0110a.16


