

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.15
(184 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
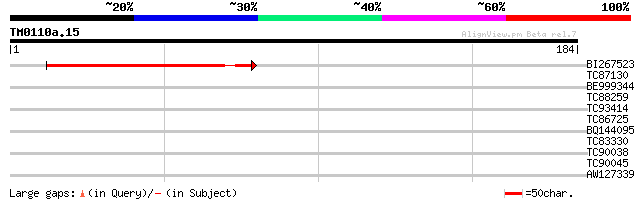
Score E
Sequences producing significant alignments: (bits) Value
BI267523 84 4e-17
TC87130 weakly similar to PIR|T04067|T04067 hypothetical protein... 37 0.004
BE999344 similar to PIR|T29150|T291 hypothetical protein F47B3.4... 33 0.084
TC88259 similar to GP|6729037|gb|AAF27033.1| unknown protein {Ar... 29 1.2
TC93414 homologue to GP|17907114|emb|CAD13249. tyrosine kinase {... 28 1.6
TC86725 similar to GP|9758909|dbj|BAB09485.1 NAM (no apical meri... 28 2.1
BQ144095 weakly similar to GP|12842949|dbj| data source:SPTR so... 27 3.5
TC83330 similar to GP|9758939|dbj|BAB09320.1 contains similarity... 27 4.6
TC90038 similar to PIR|A84451|A84451 probable AAA-type ATPase [i... 27 6.0
TC90045 similar to SP|P49160|CEMA_SOYBN Chloroplast envelope mem... 27 6.0
AW127339 similar to PIR|G84912|G849 probable acyl-CoA synthetase... 27 6.0
>BI267523
Length = 602
Score = 83.6 bits (205), Expect = 4e-17
Identities = 43/68 (63%), Positives = 49/68 (71%)
Frame = +2
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAP 72
HS FSHTVNNRTYTFLI+PPF+ FAIFD KS +TFLNRIRSSL E L+ ++ T
Sbjct: 377 HSFFSHTVNNRTYTFLIEPPFVLFAIFDTHLLKSHAITFLNRIRSSLIEPLNKNDNFT-- 550
Query: 73 PPLSLQTP 80
P LQ P
Sbjct: 551 -PFXLQPP 571
>TC87130 weakly similar to PIR|T04067|T04067 hypothetical protein F28M11.90
- Arabidopsis thaliana, partial (71%)
Length = 1510
Score = 37.0 bits (84), Expect = 0.004
Identities = 16/51 (31%), Positives = 25/51 (48%)
Frame = +1
Query: 11 STHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKE 61
S H + T+ RTY FL++ ++YF I D L FL +R ++
Sbjct: 514 SFHRWYFETIGKRTYGFLVEDGYVYFTIVDEGLGNLVVLRFLEHVRDEFRK 666
>BE999344 similar to PIR|T29150|T291 hypothetical protein F47B3.4 -
Caenorhabditis elegans, partial (7%)
Length = 596
Score = 32.7 bits (73), Expect = 0.084
Identities = 32/128 (25%), Positives = 55/128 (42%), Gaps = 7/128 (5%)
Frame = +3
Query: 21 NNRTYTFLIDPPFIYFAIFDHRHTKS--QTLTFLNRIRSSLKETLDSVNDSTA--PPPLS 76
++R TFL+ P ++ DH HT S Q ++R S L N+ T+ PPP+S
Sbjct: 30 HHRRLTFLLSSPSLF*LTLDHLHTLSLQQKTLLISRHLLSFLHLLTQSNNPTSPLPPPVS 209
Query: 77 L---QTPFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLA 133
+ D +L +L + +P A+ L K + + + + +SA L
Sbjct: 210 TSMRERDTDAVLLLLLFCETHKYNPEALTAPFSHWRLNLNK-IFSNTLLTLSYSSAPPLG 386
Query: 134 AAAATELV 141
A + L+
Sbjct: 387 ACFGSVLI 410
>TC88259 similar to GP|6729037|gb|AAF27033.1| unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 1757
Score = 28.9 bits (63), Expect = 1.2
Identities = 25/85 (29%), Positives = 43/85 (50%)
Frame = +3
Query: 100 AVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVEEALVGSAIQI 159
AV SHD G ++ ++ A + A ++K AAA+ ++ +AVE+ S++Q+
Sbjct: 699 AVANNSHDSGSRRFELERKVAELEAQLQTSKAEAAASIRSDLQRRLEAVEKE--NSSLQL 872
Query: 160 EGFRRVWLDLRSRLSLLHESRSEKD 184
E L+SRL L +E+D
Sbjct: 873 E--------LQSRLEELEFRIAERD 923
>TC93414 homologue to GP|17907114|emb|CAD13249. tyrosine kinase {Schistosoma
mansoni}, partial (1%)
Length = 578
Score = 28.5 bits (62), Expect = 1.6
Identities = 17/49 (34%), Positives = 24/49 (48%), Gaps = 6/49 (12%)
Frame = +1
Query: 13 HSLFSHTVNNRTYTF------LIDPPFIYFAIFDHRHTKSQTLTFLNRI 55
HS S TV++ T F L PP + + F H H K+ + FLN +
Sbjct: 247 HSPASSTVSHPTSLFPPSADPLQTPPLLLLSPFHHSHQKNSSPPFLNLV 393
>TC86725 similar to GP|9758909|dbj|BAB09485.1 NAM (no apical meristem)-like
protein {Arabidopsis thaliana}, partial (50%)
Length = 1645
Score = 28.1 bits (61), Expect = 2.1
Identities = 21/74 (28%), Positives = 36/74 (48%), Gaps = 1/74 (1%)
Frame = +3
Query: 8 SKGSTHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVN 67
S+ + L S ++NN + FLI P Y I +H + T+ +N S + L++
Sbjct: 810 SQSDGYYLPSFSINNNQHQFLIKPEDNYHRINYDQHEINPTM--MNYTSISNQSNLNNPI 983
Query: 68 DSTAP-PPLSLQTP 80
+T P P + +Q P
Sbjct: 984 GNTLPQPQIRIQNP 1025
>BQ144095 weakly similar to GP|12842949|dbj| data source:SPTR source
key:Q9Y2Y1 evidence:ISS~homolog to PUTATIVE
DNA-DIRECTED RNA POLYMERASE, partial (39%)
Length = 615
Score = 27.3 bits (59), Expect = 3.5
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = -3
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIF 39
HSLF + T + LI PP+IY I+
Sbjct: 541 HSLFLSASHYNTSSHLIHPPYIYIYIY 461
>TC83330 similar to GP|9758939|dbj|BAB09320.1 contains similarity to isoamyl
acetate-hydrolyzing esterase~gene_id:K15I22.12
{Arabidopsis thaliana}, partial (73%)
Length = 797
Score = 26.9 bits (58), Expect = 4.6
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = +1
Query: 25 YTFLIDPPFIYFAIFDHRHTKSQTLTF 51
Y +L+D PFI + H H K++ +F
Sbjct: 103 YIYLVDLPFILSFLHQHDHDKAKDFSF 183
>TC90038 similar to PIR|A84451|A84451 probable AAA-type ATPase [imported]
- Arabidopsis thaliana, partial (43%)
Length = 1000
Score = 26.6 bits (57), Expect = 6.0
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +2
Query: 16 FSHTVNNRTYTFLIDPPF 33
FSH+VNN+ F + PPF
Sbjct: 56 FSHSVNNQRRLFFLVPPF 109
>TC90045 similar to SP|P49160|CEMA_SOYBN Chloroplast envelope membrane
protein. [Soybean] {Glycine max}, partial (94%)
Length = 1076
Score = 26.6 bits (57), Expect = 6.0
Identities = 20/72 (27%), Positives = 30/72 (40%)
Frame = +2
Query: 10 GSTHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDS 69
GS + F T NN+ + L+ F + K +LNR+ SL S+ND
Sbjct: 494 GSVYKDFGFTQNNQIISGLVST----FPVILDTILKYWIFRYLNRVSPSLVVIYHSMND* 661
Query: 70 TAPPPLSLQTPF 81
P + L+ F
Sbjct: 662 KNDPLIQLENLF 697
>AW127339 similar to PIR|G84912|G849 probable acyl-CoA synthetase [imported]
- Arabidopsis thaliana, partial (8%)
Length = 350
Score = 26.6 bits (57), Expect = 6.0
Identities = 14/35 (40%), Positives = 18/35 (51%)
Frame = -3
Query: 43 HTKSQTLTFLNRIRSSLKETLDSVNDSTAPPPLSL 77
HTK+Q +T N S LK + T PP L+L
Sbjct: 267 HTKAQVVTRKNSSISELKAAPPDIISLTRPPSLAL 163
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,404,926
Number of Sequences: 36976
Number of extensions: 73225
Number of successful extensions: 441
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 438
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 441
length of query: 184
length of database: 9,014,727
effective HSP length: 90
effective length of query: 94
effective length of database: 5,686,887
effective search space: 534567378
effective search space used: 534567378
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0110a.15


