

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.13
(1389 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
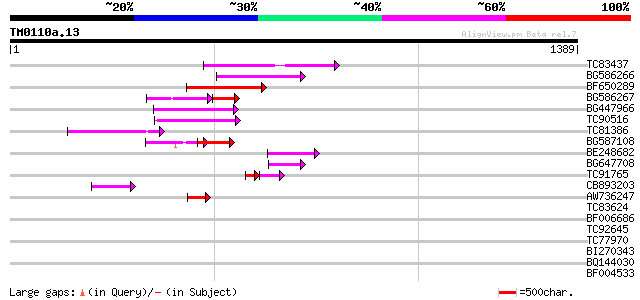
Score E
Sequences producing significant alignments: (bits) Value
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 202 7e-52
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 156 5e-38
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 146 6e-35
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 98 6e-32
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 125 1e-28
TC90516 107 4e-23
TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR re... 101 2e-21
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 86 8e-17
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 82 1e-15
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 71 3e-12
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 44 2e-07
CB893203 53 7e-07
AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase... 53 7e-07
TC83624 homologue to PIR|G84581|G84581 copia-like retroelement p... 37 0.053
BF006686 33 0.59
TC92645 33 1.0
TC77970 similar to GP|20197686|gb|AAM15203.1 expressed protein {... 31 3.8
BI270343 31 3.8
BQ144030 weakly similar to SP|P11592|VATA_ Vacuolar ATP synthase... 30 6.5
BF004533 similar to PIR|H86246|H862 hypothetical protein [import... 30 8.5
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 202 bits (514), Expect = 7e-52
Identities = 121/333 (36%), Positives = 184/333 (54%), Gaps = 1/333 (0%)
Frame = +2
Query: 476 KFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVV 535
+F +FH KL KG+N+TFI+LIPK + P ++DFRPISL+GS YK++ K+LA+RL+VV
Sbjct: 5 RFVSEFHRNRKLFKGINSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVV 184
Query: 536 TPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFD 595
+IS+ Q AF + RQI + + + + + F K + G ++
Sbjct: 185 IGSVISDAQSAFVKNRQILEMVFL*QMRLWMRLRN*RKIFCCLRWILKRLITLSIGLIWI 364
Query: 596 IL-KEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNG 654
+ M+F W WI+ CVSTAT S+LVNGS T + K L Q + + F +
Sbjct: 365 LF*VGMSFLVLWRKWIKECVSTATTSVLVNGSPTNV--LMKSLVQTQLFTRYSFGV---- 526
Query: 655 LSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSG 714
+ P + ++HLQFA+DTLL + I L L+ F SG
Sbjct: 527 ------------------VNP-VVVSHLQFANDTLLLETKNWANIRALRAALVIF*AMSG 649
Query: 715 LKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKL 774
LKVN HKS ++ N+ A + W +G++P LYLG P+ GN RR SFWEP+++++
Sbjct: 650 LKVNFHKSGLVCVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPIVNRI 829
Query: 775 RRKRRTWNSKFISLSGRLTLMKATLCSVPLFLM 807
+ + WNS+F+S GRL L+K+ L S+ ++ +
Sbjct: 830 KARLTGWNSRFLSFGGRLVLLKSVLTSLSVYAL 928
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 156 bits (395), Expect = 5e-38
Identities = 86/221 (38%), Positives = 128/221 (57%), Gaps = 4/221 (1%)
Frame = -3
Query: 508 VSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL 567
VS++R I+ + YK+I+K+L+ R+Q + +IS +Q AF GR ISD +LI ++++ +L
Sbjct: 778 VSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYL 599
Query: 568 NK---KQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVN 624
+ K+ +K D K YD I W FL ++L + F WI+WI CVST + S L+N
Sbjct: 598 RQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLIN 419
Query: 625 GSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQ 683
G G +GLRQGDPLSP+LF LC LS L Q L GV++ +NHL
Sbjct: 418 GGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINHLL 239
Query: 684 FADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTI 724
FADDT+ F +++ + +L + + + SG +N KS I
Sbjct: 238 FADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAI 116
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 146 bits (368), Expect = 6e-35
Identities = 72/196 (36%), Positives = 123/196 (62%), Gaps = 1/196 (0%)
Frame = +3
Query: 434 QFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNA 493
+F+ E+ L S D ++APG DG+N++FFK W I+ + V+ DF TG + K +N
Sbjct: 21 EFTAVEVKNALFSMDSSKAPGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPKIINC 200
Query: 494 TFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQI 553
T+++L+PK ++V +FRPI+ YK+ISK+L SR+Q V ++SENQ AF +GR I
Sbjct: 201 TYVTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVKGRVI 380
Query: 554 SDCILIANEVVDFLNKKQ-GGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRT 612
D I++++E+V ++K ++K+D K YD+ +W F+ ++ E+ F +++NW+
Sbjct: 381 FDNIILSHELVKSYSRKGISPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPYKFVNWVMA 560
Query: 613 CVSTATLSILVNGSST 628
++TA+ + NG T
Sbjct: 561 XLTTASYTFNXNGDLT 608
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 97.8 bits (242), Expect(2) = 6e-32
Identities = 56/166 (33%), Positives = 92/166 (54%), Gaps = 5/166 (3%)
Frame = +3
Query: 336 QEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGG----VSIEDPN 391
+EY EE W QK+R+ WLR GDRNT FFH+++K R+++ I L+ ED
Sbjct: 108 EEYHNEEIFWMQKSRLNWLRSGDRNTKFFHAVTKNRRAQNRILSLIDDDDKEWFVEEDLG 287
Query: 392 MIKDAIFSHFQSFFTSEN-RLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGN 450
+ D SHF+ ++SE+ + + + + +++ L Q S E+ E + + +
Sbjct: 288 RLAD---SHFKLLYSSEDVGITLEDWNSIPAIVTEEQNAQLMAQISREEVREAVFDINPH 458
Query: 451 RAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFI 496
+ PG DG N+ FF+QFW + +D+ ++F TGKL +G+N T I
Sbjct: 459 KCPGPDGMNVFFFQQFWDTMGDDLTSMAQEFLRTGKLEEGINKTNI 596
Score = 59.7 bits (143), Expect(2) = 6e-32
Identities = 32/64 (50%), Positives = 44/64 (68%)
Frame = +1
Query: 498 LIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCI 557
L+PK + +FRPISL AYK++SKVL+ RL+ V P +I+E Q AF + + ISD I
Sbjct: 601 LVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSVLPWIITETQAAFGRRQLISDNI 780
Query: 558 LIAN 561
LIA+
Sbjct: 781 LIAH 792
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 125 bits (313), Expect = 1e-28
Identities = 75/210 (35%), Positives = 116/210 (54%), Gaps = 2/210 (0%)
Frame = +2
Query: 353 WLREGDRNTSFFHSISKIRKSKKLISQLV-VGGVSIEDPNMIKDAIFSHFQSFFTSENRL 411
WL++GD+NT FFHS + R+ I +L G + ++ + ++F + FTS N
Sbjct: 5 WLKDGDKNTKFFHSKASQRRKVNEIKKLKDETGNWCKGEENVERLLITYFNNLFTSSNPT 184
Query: 412 RPKIRCANLS-RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIV 470
+ C + +LS + + EK+F+E E+ E + +APG DG FF+++W IV
Sbjct: 185 AIEETCEVVKGKLSHEHIVWCEKEFTEEEVLEAINQMHPVKAPGPDGLPALFFQKYWHIV 364
Query: 471 KEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLAS 530
++V + + + LN TFI LIPK P+T D+RPISL K+I+KV+A+
Sbjct: 365 GKEVQQMVLQVLNNSMETEELNKTFIVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIAN 544
Query: 531 RLQVVTPLLISENQFAFAQGRQISDCILIA 560
R++ P +I Q AF QGR I+D LIA
Sbjct: 545 RVKQTLPDVIDVEQSAFVQGRLITDNALIA 634
>TC90516
Length = 983
Score = 107 bits (266), Expect = 4e-23
Identities = 72/210 (34%), Positives = 111/210 (52%)
Frame = -2
Query: 355 REGDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIFSHFQSFFTSENRLRPK 414
REG NT++F + K R + IS V ++D K AI + F F+ RP
Sbjct: 754 REG-HNTTYFLACVKNRGMRNSISAPRVERW-MDDVAKNKQAIVNFFTHHFSDP*TYRPT 581
Query: 415 IRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDV 474
+ + +S + + L QF ++ V+ S DGN +P DGFNLNFF + ++K D+
Sbjct: 580 MGDIDFFHISNLDNVLLSAQFLVSKMELVVSSLDGNESPRPDGFNLNFFIRLRNMLKADI 401
Query: 475 VKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQV 534
FE F++ L+K + F++LI K P + DF +S +GS KL++KVLA RL
Sbjct: 400 EIMFEQFYTPANLLKIFS*YFLTLISKVEYPILLGDFSLMSFLGSL*KLMAKVLALRLAH 221
Query: 535 VTPLLISENQFAFAQGRQISDCILIANEVV 564
+ +I NQ F +GRQ D ++ NE++
Sbjct: 220 IMEKIIFVNQSTFVRGRQHVDGVVAINEII 131
>TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 798
Score = 101 bits (252), Expect = 2e-21
Identities = 76/247 (30%), Positives = 109/247 (43%), Gaps = 9/247 (3%)
Frame = -1
Query: 141 GDFNAILNEGERIGIQMGVESSFSDFVADS---NLIDLPLQSDKFTWYSSRAG--GLWSR 195
GDFN IL E G DF S NL LP + FTW + R G R
Sbjct: 729 GDFNTILGSHEHQGSHTPARLPMLDFQQWSDVNNLFHLPTRGSAFTWTNGRRGRNNTRKR 550
Query: 196 IDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFG-PKPFLFYNHWLLDKEFE 254
+DR +V++ I+ + +S SD PI+ L N F F W +
Sbjct: 549 LDRSIVNQLMIDNCESLSACTLTKLRSDHFPILFELQTQNIQFSSSFKFMKMWSAHPDCI 370
Query: 255 KLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEA---DLQVTM 311
++ W + VG FVL QKLK LK +K W +++ D+QV +
Sbjct: 369 NIIKQCWANQV-VGCPMFVLNQKLKNLKEVLKVWNKNTFGNVHSQVDNAYKELDDIQVKI 193
Query: 312 DRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIR 371
D + + ++ K LE EE W +K++V W EGDRNT+FFH ++KI+
Sbjct: 192 DSIGYSDVLMDQEKAAQLNLESA---LNIEEVFWHEKSKVNWHCEGDRNTAFFHRVAKIK 22
Query: 372 KSKKLIS 378
++ LIS
Sbjct: 21 RTSSLIS 1
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 86.3 bits (212), Expect = 8e-17
Identities = 45/91 (49%), Positives = 58/91 (63%)
Frame = +3
Query: 460 LNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGS 519
L+FF+ W I+K D++K F ++GKL LN T I LIPK P+ +++ RPISL
Sbjct: 405 LSFFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNV 584
Query: 520 AYKLISKVLASRLQVVTPLLISENQFAFAQG 550
YK+ISKVL RL+V P LISE Q AF G
Sbjct: 585 GYKIISKVLCQRLKVCLPSLISETQSAFVHG 677
Score = 48.1 bits (113), Expect = 2e-05
Identities = 38/162 (23%), Positives = 70/162 (42%), Gaps = 6/162 (3%)
Frame = +2
Query: 332 EGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSK-KLISQLVVGGVSIEDP 390
E L + Y+ EE W+QK+R W GD N F+H+++K R ++ +++ G I +
Sbjct: 14 EKLQEAYKDEEDYWQQKSRNMWHISGDLNKKFYHALTKQRHARNRIVGLYDYDGNWITEE 193
Query: 391 NMIKDAIFSHFQSFF-----TSENRLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLK 445
++ +F+ F T + +I + +++Q L + +E E+ L
Sbjct: 194 QGVEKVAVDYFEDLFQRTTPTGFDGFLDEITSSITPQMNQ----RLLRLATEEEVRLALF 361
Query: 446 SCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKL 487
+APG DG F K ++ + F GK+
Sbjct: 362 IMHPEKAPGPDGMTTLLFSTLMAYNKNGSIENGQ*FSGIGKI 487
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 82.4 bits (202), Expect = 1e-15
Identities = 48/127 (37%), Positives = 69/127 (53%), Gaps = 1/127 (0%)
Frame = +3
Query: 633 MEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRI-GPGLALNHLQFADDTLLF 691
+++GL+QGDPL+PFLF L G+S L+ ++ + F G + G ++HLQ+ADDTL
Sbjct: 18 VQRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADDTLCI 197
Query: 692 CENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPIL 751
+ + L L F SGLKVN HKS++IG NV + +P +
Sbjct: 198 GMPTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAACRFLNCREESIPFI 377
Query: 752 YLGAPLG 758
YLG P G
Sbjct: 378 YLGLPGG 398
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 70.9 bits (172), Expect = 3e-12
Identities = 40/92 (43%), Positives = 53/92 (57%), Gaps = 1/92 (1%)
Frame = +1
Query: 634 EKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIG-PGLALNHLQFADDTLLFC 692
EKGLRQGDPLSP+LF LC N LS LL + + N G+++ + HL FADD+LLF
Sbjct: 7 EKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLFA 186
Query: 693 ENDENQIDLLCNTLLSFLFTSGLKVNLHKSTI 724
+ + + L S+ SG VN KS +
Sbjct: 187 RANLTEAATIMQVLHSYQSASGQLVNFEKSEV 282
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 43.9 bits (102), Expect(2) = 2e-07
Identities = 22/61 (36%), Positives = 33/61 (54%)
Frame = +2
Query: 613 CVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVR 672
CV + +LVN + +GL+QGD LSP++F +CV GLS L+ + + G
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKERGDTHGTS 283
Query: 673 I 673
I
Sbjct: 284 I 286
Score = 30.8 bits (68), Expect(2) = 2e-07
Identities = 12/32 (37%), Positives = 21/32 (65%)
Frame = +1
Query: 579 LDFAKDYDNIDWGFLFDILKEMNFGDRWINWI 610
L +K Y+ +D +L +I+ +M F +RWI W+
Sbjct: 1 LYISKVYNRVD*DYLKEIMIKMGFNNRWIYWM 96
>CB893203
Length = 800
Score = 53.1 bits (126), Expect = 7e-07
Identities = 33/107 (30%), Positives = 50/107 (45%)
Frame = -3
Query: 201 VSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFGPKPFLFYNHWLLDKEFEKLVSDW 260
VSE K+ + Q GLSDR PI+L++ N+GP+ W + V D
Sbjct: 798 VSESWCIKWPNMIQRALFRGLSDRCPIMLTIDEGNWGPRLHHMLKCWADLPGYHLFVKDK 619
Query: 261 WCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEADL 307
W S W +VL++K K ++N +K+W + RIN I +
Sbjct: 618 WNSFQVSRWGGYVLKEK*KMVRNNLKDWHHNHTRNLDARINDIRESI 478
>AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (4%)
Length = 305
Score = 53.1 bits (126), Expect = 7e-07
Identities = 23/58 (39%), Positives = 36/58 (61%), Gaps = 2/58 (3%)
Frame = +1
Query: 435 FSEGEIWEVLKSCDGNRA--PGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKG 490
FSE E+ + + CD + + PG DG N F K++W ++K D ++ +FH+ GKL KG
Sbjct: 121 FSEEEVRKAVWDCDSSHS*NPGPDGVNFTFVKEYWELIKVDFLRVVMEFHTNGKLGKG 294
>TC83624 homologue to PIR|G84581|G84581 copia-like retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(1%)
Length = 831
Score = 37.0 bits (84), Expect = 0.053
Identities = 13/32 (40%), Positives = 24/32 (74%)
Frame = +1
Query: 744 SIGQLPILYLGAPLGGNPRRASFWEPMLDKLR 775
++ ++P +LG P+G NP+R+S +P+LD L+
Sbjct: 331 NVNEVPFCFLGLPIGANPKRSSTRKPVLDSLQ 426
>BF006686
Length = 325
Score = 33.5 bits (75), Expect = 0.59
Identities = 10/29 (34%), Positives = 20/29 (68%)
Frame = +3
Query: 767 WEPMLDKLRRKRRTWNSKFISLSGRLTLM 795
WEP+L+ + + ++W +K +S GR+ L+
Sbjct: 237 WEPLLEHVNKMLKSWGNKLLSFGGRIVLL 323
>TC92645
Length = 561
Score = 32.7 bits (73), Expect = 1.0
Identities = 28/94 (29%), Positives = 41/94 (42%), Gaps = 5/94 (5%)
Frame = +3
Query: 422 RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDF 481
++ N L FS EIW + ++A G DG NL + +F +V D + +
Sbjct: 279 KIGDQNNCMLTAPFSVDEIWTAIFQMHHDKAFGLDGLNLAVYYRF*NLVGGDT-NYVGLY 455
Query: 482 HSTGK-----LVKGLNATFISLIPKHNCPSTVSD 510
G L G N I L+PK + PST+ D
Sbjct: 456 SLVG*R*VA*LYWGTN---IVLVPKCDSPSTMWD 548
>TC77970 similar to GP|20197686|gb|AAM15203.1 expressed protein {Arabidopsis
thaliana}, partial (97%)
Length = 2062
Score = 30.8 bits (68), Expect = 3.8
Identities = 26/83 (31%), Positives = 38/83 (45%), Gaps = 5/83 (6%)
Frame = -2
Query: 419 NLSRLSQA-----NKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKED 473
NLS LSQA NK+SL +++ K C + T L F + FWG+ E
Sbjct: 1971 NLSFLSQACT*LTNKISLTQEY---------KKCPSHCFCYT-ARRLKFKRWFWGVRVES 1822
Query: 474 VVKFFEDFHSTGKLVKGLNATFI 496
V + +FH V N+T++
Sbjct: 1821 EVHMYGEFHHHAYAVLHCNSTWM 1753
>BI270343
Length = 619
Score = 30.8 bits (68), Expect = 3.8
Identities = 11/41 (26%), Positives = 26/41 (62%)
Frame = +1
Query: 4 VSWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRV 44
+SWN RGLG + + + + K ++V++ E+ ++S+++
Sbjct: 484 ISWNCRGLGNVKAVPCIKDLIRVYKSDVVILIETLVESNKI 606
>BQ144030 weakly similar to SP|P11592|VATA_ Vacuolar ATP synthase catalytic
subunit A (EC 3.6.3.14) (V-ATPase A subunit), partial
(3%)
Length = 779
Score = 30.0 bits (66), Expect = 6.5
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = -3
Query: 107 MNVYGPNVDSERYDFFNLLSEALSLNNCF 135
++ P++ YDF NL S +SL NCF
Sbjct: 126 LSTSSPDISPSCYDFMNLKSPVISLCNCF 40
>BF004533 similar to PIR|H86246|H862 hypothetical protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 571
Score = 29.6 bits (65), Expect = 8.5
Identities = 27/98 (27%), Positives = 44/98 (44%), Gaps = 1/98 (1%)
Frame = +1
Query: 349 ARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIFSHFQSFFTSE 408
A + W R + +T +HSI RK+ K QL G + ++ N D +F SE
Sbjct: 238 AYIMWRRTSNSSTKLWHSIKSTRKTNKKDFQLFNKGGTSDENN--SDDVFGGL-----SE 396
Query: 409 NRLRPKIRCANLSRLSQA-NKMSLEKQFSEGEIWEVLK 445
RL+ ++ + +L+ A N L + +G V K
Sbjct: 397 VRLQ-ELLLFDFEKLATATNNFHLSNKLGQGGFGPVYK 507
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.334 0.147 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,482,036
Number of Sequences: 36976
Number of extensions: 737335
Number of successful extensions: 5597
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 3051
Number of HSP's successfully gapped in prelim test: 247
Number of HSP's that attempted gapping in prelim test: 2402
Number of HSP's gapped (non-prelim): 3528
length of query: 1389
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1281
effective length of database: 5,021,319
effective search space: 6432309639
effective search space used: 6432309639
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0110a.13


