

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.10
(1408 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
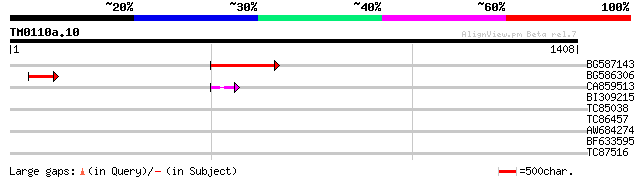
BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F2... 176 4e-44
BG586306 weakly similar to GP|15128241|db helicase-like protein ... 79 2e-14
CA859513 50 5e-06
BI309215 SP|P03749|VH Host specificity protein J. {Bacteriophage... 33 0.60
TC85038 weakly similar to PIR|G86449|G86449 hypothetical protein... 32 1.7
TC86457 similar to SP|Q06364|PSD3_DAUCA Probable 26S proteasome ... 31 3.9
AW684274 similar to PIR|T17000|T170 oxygenase ARRO-1 2-oxoacid ... 30 5.1
BF633595 30 6.6
TC87516 PIR|T09591|T09591 probable cdc2-like protein kinase cdc2... 30 8.6
>BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F20D21.24
[imported] - Arabidopsis thaliana, partial (12%)
Length = 717
Score = 176 bits (447), Expect = 4e-44
Identities = 84/175 (48%), Positives = 117/175 (66%), Gaps = 3/175 (1%)
Frame = -3
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
++ IEFQKRGL HAHILLW + S +D +I AE+P+ K P + V+ +M+HG
Sbjct: 589 LHRIEFQKRGLRHAHILLWFGNSSRTPSSEEVDEIISAELPNKKQDPEAYNLVTKHMIHG 410
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRR-NTGVSTNRRGVDLDNGF 617
PCG+ +SPCM+N C K +P+ Y T+ D GY +YRRR N + G L+N F
Sbjct: 409 PCGVINPKSPCMENNVCTKKYPRPYNGSTSIDKSGYVLYRRR*NETEHVVKNGAILNNTF 230
Query: 618 VVPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGR--TTQD 670
+VP+N KLL KY+AHIN+E+CN+++++KYLFKYI KGVDRV++ + G TT D
Sbjct: 229 IVPHNIKLLKKYEAHINMEWCNRTSAVKYLFKYITKGVDRVSVVIEKGNPATTSD 65
>BG586306 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (2%)
Length = 667
Score = 78.6 bits (192), Expect = 2e-14
Identities = 38/76 (50%), Positives = 52/76 (68%)
Frame = +2
Query: 46 RHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFA 105
R F D IR YNS+ AFTSIG K+D + + G I GQ +HRI SL+P +G P++
Sbjct: 5 RWFRDTIRVYNSVLAFTSIGMKMDYSVVNAPGRYTIRIQGQTHHRIDSLIPRQGRPPEYL 184
Query: 106 QLYIYDTKNEVQNRMS 121
Q+YI+DT NEV+NR++
Sbjct: 185 QIYIFDTGNEVRNRLN 232
>CA859513
Length = 363
Score = 50.4 bits (119), Expect = 5e-06
Identities = 26/73 (35%), Positives = 38/73 (51%)
Frame = +1
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+Y EF+K LPHAH+L + Y D+ + AE+ DP L++ V + M+HG
Sbjct: 136 IYVAEFEKCDLPHAHMLFHRADNY--------DTFVSAELSDPVEQLRLYQTVVSVMIHG 291
Query: 559 PCGLSKRESPCMK 571
P G PCM+
Sbjct: 292 PYGPFNNNVPCMR 330
>BI309215 SP|P03749|VH Host specificity protein J. {Bacteriophage lambda},
partial (15%)
Length = 545
Score = 33.5 bits (75), Expect = 0.60
Identities = 30/90 (33%), Positives = 41/90 (45%), Gaps = 3/90 (3%)
Frame = +3
Query: 765 PELFVYDP--KSKSWHPRKQGESIGRMSFVPPGTGE-LYYLRMLLNVQRGCTKYEDLRSV 821
P L VYDP + + W KQ I ++ G LY++ +N++ G Y +RSV
Sbjct: 99 PHLAVYDPTVQFEFWFSEKQIADIRQVETSTRYLGTALYWIAASINIKPGHDYYFYIRSV 278
Query: 822 NNNVHDTFRGACEALGLLKDDREFIDGILD 851
N F EA+G DD E G LD
Sbjct: 279 NTVGKSAF---VEAVGRASDDAE---GYLD 350
>TC85038 weakly similar to PIR|G86449|G86449 hypothetical protein AAF81343.1
[imported] - Arabidopsis thaliana, partial (14%)
Length = 652
Score = 32.0 bits (71), Expect = 1.7
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = +2
Query: 1016 NFCSI*G*WWVFLCQWTWWYGENILV 1041
N CS *G WW +LCQ T +Y +I+V
Sbjct: 533 NLCSS*G-WWHYLCQLTCFYSNHIVV 607
>TC86457 similar to SP|Q06364|PSD3_DAUCA Probable 26S proteasome non-ATPase
regulatory subunit 3 (26S proteasome subunit S3), partial
(96%)
Length = 1913
Score = 30.8 bits (68), Expect = 3.9
Identities = 20/58 (34%), Positives = 33/58 (56%), Gaps = 2/58 (3%)
Frame = +1
Query: 1273 LYLLQ--LWMLLILLIDTRCLYSMPRRGLI*ALIQYGVLMKRLPLKRIGLQLNFLMIS 1328
LYLL +++LL+L + CL PRR ++++ ++ LP K + LQLN +S
Sbjct: 325 LYLLFSIMFLLLVLNLMLNCLRIFPRR----MIMRWRWTLQHLPFKPLSLQLNTCCLS 486
>AW684274 similar to PIR|T17000|T170 oxygenase ARRO-1 2-oxoacid dependent -
apple tree, partial (29%)
Length = 524
Score = 30.4 bits (67), Expect = 5.1
Identities = 18/47 (38%), Positives = 26/47 (55%), Gaps = 4/47 (8%)
Frame = -3
Query: 865 FVSFLLPNS--MCN--PLHVWNETWHVLADGIACDLQRKHNNRGMCL 907
FV F LPN +C+ PL ++E WH + I L + N+RG+ L
Sbjct: 444 FVLF*LPNHVLLCHMPPLFPFDEVWHSIESRIHAQLVKNLNHRGLKL 304
>BF633595
Length = 313
Score = 30.0 bits (66), Expect = 6.6
Identities = 13/27 (48%), Positives = 18/27 (66%)
Frame = +1
Query: 136 IYVDMCLIVFSYLCEIVVILVANVHLK 162
IY+ + L +F Y+C V V+NVHLK
Sbjct: 184 IYIAVSLFIFYYICLHVWFKVSNVHLK 264
>TC87516 PIR|T09591|T09591 probable cdc2-like protein kinase cdc2MsF -
alfalfa, complete
Length = 1251
Score = 29.6 bits (65), Expect = 8.6
Identities = 11/48 (22%), Positives = 28/48 (57%)
Frame = +2
Query: 1244 LLIL*VTLYPQCIQISNKMLVVCSIFKTRLYLLQLWMLLILLIDTRCL 1291
L++L + YP + N ++ S++ +L+++++W+ L L+ C+
Sbjct: 1040 LIMLLSSAYPSLF*VDNICRLIFSLYSIKLWIVRIWIKLNSLVRKECM 1183
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.353 0.158 0.566
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,460,904
Number of Sequences: 36976
Number of extensions: 795861
Number of successful extensions: 10128
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 3184
Number of HSP's successfully gapped in prelim test: 518
Number of HSP's that attempted gapping in prelim test: 6657
Number of HSP's gapped (non-prelim): 4323
length of query: 1408
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1300
effective length of database: 5,021,319
effective search space: 6527714700
effective search space used: 6527714700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0110a.10


