

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.3
(169 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
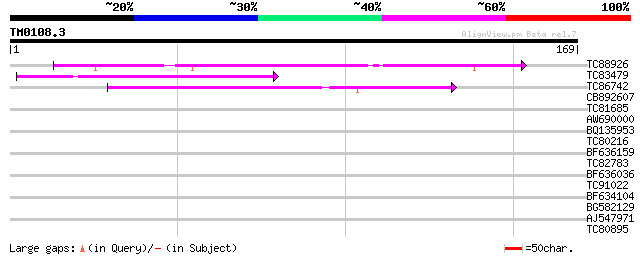
TC88926 similar to GP|7290507|gb|AAF45960.1| peb gene product {D... 55 2e-08
TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protei... 51 2e-07
TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG00... 42 2e-04
CB892607 similar to GP|8927657|gb| EST gb|N38213 comes from this... 37 0.004
TC81685 35 0.019
AW690000 31 0.27
BQ135953 29 1.0
TC80216 similar to GP|10176796|dbj|BAB09935. contains similarity... 29 1.0
BF636159 similar to GP|22136764|gb unknown protein {Arabidopsis ... 27 3.9
TC82783 similar to GP|22136764|gb|AAM91701.1 unknown protein {Ar... 27 3.9
BF636036 similar to GP|6069471|dbj S-locus protein 3 {Brassica r... 27 5.1
TC91022 27 5.1
BF634104 27 5.1
BG582129 26 6.7
AJ547971 similar to PIR|T08399|T083 hypothetical protein F18B3.6... 26 8.7
TC80895 similar to GP|10178031|dbj|BAB11514. gene_id:MUG13.23~un... 26 8.7
>TC88926 similar to GP|7290507|gb|AAF45960.1| peb gene product {Drosophila
melanogaster}, partial (1%)
Length = 1073
Score = 54.7 bits (130), Expect = 2e-08
Identities = 49/149 (32%), Positives = 70/149 (46%), Gaps = 8/149 (5%)
Frame = +3
Query: 14 NVDGSVWQTRE---AGCGGVLRDCDGNWVQGFCRRL*SSNPLL----AELWAIRTAVDLA 66
NVDGS+ + RE AGCGGVL D G W+ GF ++L NP L E AI +
Sbjct: 351 NVDGSLLREREVPSAGCGGVLSDSSGKWLCGFAQKL---NPNLKVDETEKEAILRGLLWV 521
Query: 67 VQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMV 126
++G I++++D+ +V P +R + S PH + I +N V
Sbjct: 522 KEKGKRKILVKSDNEGVVYSVNCGGRSNDPLVCGIRDLLNS-PH-WEATLTCIHGRSNAV 695
Query: 127 ADCLAQNAHVF-NFGVHLLHHPLMECRYL 154
AD LA AH F +F + +P C L
Sbjct: 696 ADRLAHKAHSFTSFDLCQFDYPPENCTSL 782
>TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 1222
Score = 51.2 bits (121), Expect = 2e-07
Identities = 30/78 (38%), Positives = 40/78 (50%)
Frame = +2
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTA 62
W P +G K N DGSV AG GG+LRD G + F + + L ELWAI
Sbjct: 977 WKKPEIGWTKLNTDGSV-NKETAGFGGLLRDYRGEPICAFVSKAPQGDTFLVELWAIWRG 1153
Query: 63 VDLAVQRGCSPIIIETDS 80
+ L++ G I +E+DS
Sbjct: 1154 LVLSLGLGIKSIWVESDS 1207
>TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG005I10.16 -
Arabidopsis thaliana, partial (19%)
Length = 2073
Score = 41.6 bits (96), Expect = 2e-04
Identities = 33/105 (31%), Positives = 51/105 (48%), Gaps = 1/105 (0%)
Frame = -1
Query: 30 VLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHAAVTS 89
V+RD ++ + + PL AE A A++ A++ G + I +ETDS + A
Sbjct: 468 VIRDSQFGFLGALSCNIGHATPLEAEFCACMIAIEKAMELGLNNICLETDSLKVVNAFHK 289
Query: 90 AQPLIPPYSEIVR-HIRRSIPHGMKVVFDLIPREANMVADCLAQN 133
+ P+ VR H H + V IPRE N+VAD LA++
Sbjct: 288 IVGI--PWQMRVRWHNCIRFCHSIACVCVHIPREGNLVADALARH 160
>CB892607 similar to GP|8927657|gb| EST gb|N38213 comes from this gene.
{Arabidopsis thaliana}, partial (1%)
Length = 782
Score = 37.0 bits (84), Expect = 0.004
Identities = 19/50 (38%), Positives = 29/50 (58%)
Frame = +1
Query: 25 AGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPI 74
A CGG+++D G++V + +L S + L AELW I + + RG S I
Sbjct: 622 AACGGLIQDDQGHFVFHYANKLGSCSVLQAELWGI*HGLSIDWNRGYSKI 771
>TC81685
Length = 1706
Score = 34.7 bits (78), Expect = 0.019
Identities = 17/44 (38%), Positives = 22/44 (49%)
Frame = -2
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRRL 46
W LP K N DG + AGCGG++ G W+ GF + L
Sbjct: 172 WTLPQSDWDKINSDGMCQGSLRAGCGGLI*GDGGEWLCGFSKFL 41
>AW690000
Length = 652
Score = 30.8 bits (68), Expect = 0.27
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Frame = +3
Query: 6 PPVGSW-KFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAEL 56
PP+ W K N DGS ++ + CGG+ R+ D + + F N AEL
Sbjct: 417 PPIPHWIKCNTDGSS-RSHSSACGGIFRNHDTDLLLCFAENTGECNAFHAEL 569
>BQ135953
Length = 850
Score = 28.9 bits (63), Expect = 1.0
Identities = 15/38 (39%), Positives = 21/38 (54%)
Frame = +2
Query: 84 HAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ + TS Q I PY ++ I + HG K +FDLI R
Sbjct: 116 YLSTTSQQGFIYPYPMLIYRIDKLHFHGRKNLFDLIQR 229
>TC80216 similar to GP|10176796|dbj|BAB09935. contains similarity to unknown
protein~gb|AAF26170.1~gene_id:K5F14.3 {Arabidopsis
thaliana}, partial (59%)
Length = 1162
Score = 28.9 bits (63), Expect = 1.0
Identities = 13/41 (31%), Positives = 20/41 (48%)
Frame = +1
Query: 69 RGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIP 109
RG S I+ S + + Q L P Y+ ++ HIR +P
Sbjct: 880 RGGSSAIVANTSEKPNVIAAKLQALSPKYNSVMNHIRIHLP 1002
>BF636159 similar to GP|22136764|gb unknown protein {Arabidopsis thaliana},
partial (21%)
Length = 683
Score = 26.9 bits (58), Expect = 3.9
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = +1
Query: 105 RRSIPHGMKVVFDLIPREANMVADCLAQNAHVFNFGVHLL 144
RR + H V+F + R + C + H+++FG H+L
Sbjct: 157 RRVLLH*CLVIFSVY*RMSVFSVICKSTYVHIYDFGFHIL 276
>TC82783 similar to GP|22136764|gb|AAM91701.1 unknown protein {Arabidopsis
thaliana}, partial (36%)
Length = 823
Score = 26.9 bits (58), Expect = 3.9
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = -2
Query: 105 RRSIPHGMKVVFDLIPREANMVADCLAQNAHVFNFGVHLL 144
RR + H V+F + R + C + H+++FG H+L
Sbjct: 585 RRVLLH*CLVIFSVY*RMSVFSVICKSTYVHIYDFGFHIL 466
>BF636036 similar to GP|6069471|dbj S-locus protein 3 {Brassica rapa},
partial (50%)
Length = 674
Score = 26.6 bits (57), Expect = 5.1
Identities = 9/17 (52%), Positives = 13/17 (75%)
Frame = +2
Query: 140 GVHLLHHPLMECRYLLY 156
G +LLH+PL+ C Y+ Y
Sbjct: 56 GSNLLHYPLLRCIYICY 106
>TC91022
Length = 731
Score = 26.6 bits (57), Expect = 5.1
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = -3
Query: 31 LRDCDGNWVQGFCRRL 46
LRD D W++ FCRRL
Sbjct: 279 LRDLDNLWLRQFCRRL 232
>BF634104
Length = 299
Score = 26.6 bits (57), Expect = 5.1
Identities = 10/31 (32%), Positives = 16/31 (51%)
Frame = +3
Query: 11 WKFNVDGSVWQTREAGCGGVLRDCDGNWVQG 41
W F+++G + GC +L +C G W G
Sbjct: 123 WWFHIEGL--GVTDRGCRALLAECSGQWKPG 209
>BG582129
Length = 534
Score = 26.2 bits (56), Expect = 6.7
Identities = 12/19 (63%), Positives = 13/19 (68%), Gaps = 1/19 (5%)
Frame = -3
Query: 1 LC-WLLPPVGSWKFNVDGS 18
LC W PPVG K+NVD S
Sbjct: 274 LCTWQPPPVGRLKYNVDAS 218
>AJ547971 similar to PIR|T08399|T083 hypothetical protein F18B3.60 -
Arabidopsis thaliana, partial (15%)
Length = 507
Score = 25.8 bits (55), Expect = 8.7
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = +2
Query: 105 RRSIPHGMKVVFDLIPREANMVAD 128
RR+I HG+ ++FD+ P E + D
Sbjct: 224 RRTI*HGITLIFDIWPNECSAPPD 295
>TC80895 similar to GP|10178031|dbj|BAB11514. gene_id:MUG13.23~unknown
protein {Arabidopsis thaliana}, partial (57%)
Length = 861
Score = 25.8 bits (55), Expect = 8.7
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = -3
Query: 145 HHPLMECRYLLYQDSMTVVASLSL 168
H P + R +LYQ S+T++ASL L
Sbjct: 382 HPPTLL*RGILYQHSITILASLPL 311
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.141 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,060,903
Number of Sequences: 36976
Number of extensions: 110509
Number of successful extensions: 648
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 648
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 648
length of query: 169
length of database: 9,014,727
effective HSP length: 89
effective length of query: 80
effective length of database: 5,723,863
effective search space: 457909040
effective search space used: 457909040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0108.3


