

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
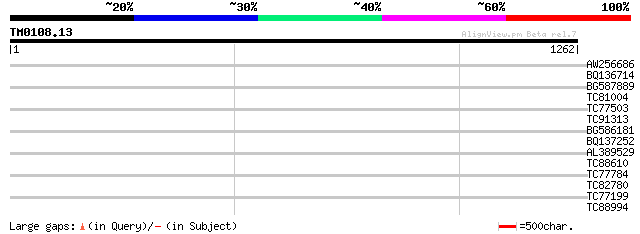
Query= TM0108.13
(1262 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW256686 similar to GP|7269000|emb| contains EST gb:AI998867.1 {... 35 0.18
BQ136714 similar to GP|22795875|emb maturase {Boopis graminea}, ... 34 0.41
BG587889 similar to GP|22136974|gb putative villin 2 protein {Ar... 33 0.91
TC81004 weakly similar to SP|Q39529|LEC2_CLALU Agglutinin II pre... 32 1.6
TC77503 weakly similar to GP|16930417|gb|AAL31894.1 At2g24420/T2... 32 1.6
TC91313 similar to GP|15292869|gb|AAK92805.1 unknown protein {Ar... 31 3.5
BG586181 similar to GP|4689316|gb| peroxisomal targeting signal ... 31 3.5
BQ137252 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 31 3.5
AL389529 31 3.5
TC88610 homologue to PIR|G96755|G96755 developmental protein hom... 31 3.5
TC77784 similar to GP|14272369|emb|CAC40022. bZIP transcription ... 30 4.5
TC82780 similar to PIR|T00713|T00713 helicase homolog F22O13.8 -... 30 4.5
TC77199 similar to SP|O04379|AGO1_ARATH Argonaute protein. [Mous... 30 5.9
TC88994 similar to GP|20260620|gb|AAM13208.1 unknown protein {Ar... 30 7.7
>AW256686 similar to GP|7269000|emb| contains EST gb:AI998867.1 {Arabidopsis
thaliana}, partial (3%)
Length = 632
Score = 35.0 bits (79), Expect = 0.18
Identities = 28/119 (23%), Positives = 54/119 (44%), Gaps = 1/119 (0%)
Frame = +1
Query: 898 ENASESKITPVLNRCPRIKSLKHAG-SPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFL 956
E A+E + R +L+H SP +R + + + +CA+VN H +H++ +
Sbjct: 31 EAANEYAMKQECQRAEERMTLRHKQKSPWLQRKMQRGLDKQDIGTCAAVNEHFQRHEELI 210
Query: 957 DQTPLLPISDSDLRATSGGDSNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSP 1015
+ + S SD +T D+ ++ + AN+ ++ K+ L +D P S P
Sbjct: 211 RKMYNMDSSSSD-DSTYEEDNENTADSDLDKANNILRKAKQKTVEVLEEDDKMPNSGLP 384
>BQ136714 similar to GP|22795875|emb maturase {Boopis graminea}, partial (4%)
Length = 1268
Score = 33.9 bits (76), Expect = 0.41
Identities = 25/77 (32%), Positives = 37/77 (47%), Gaps = 4/77 (5%)
Frame = +2
Query: 7 LRESASAMEETLISRKRSNPSSS---PPGALTRSRSQLFV-HRNRSGQRRPDSGPRQGGL 62
+R+ S E T + K + SS+ P T R+ + HR R + RP GP +GG
Sbjct: 125 MRQDTSITERTNKTEKTARDSST*TVPRRPPTHPRTPTPLGHRQRPPRPRP--GPERGGG 298
Query: 63 YSGLGGSSGNDEKREPP 79
+ G +G +K EPP
Sbjct: 299 ACSVAGEAGRCQKEEPP 349
>BG587889 similar to GP|22136974|gb putative villin 2 protein {Arabidopsis
thaliana}, partial (15%)
Length = 799
Score = 32.7 bits (73), Expect = 0.91
Identities = 25/81 (30%), Positives = 40/81 (48%), Gaps = 3/81 (3%)
Frame = -1
Query: 375 PLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGECL---SVPSG 431
PL S+S + S L S + + E + S L+S L + + L EVSG+C +V +
Sbjct: 655 PLLSESGETSSTSLASVASSTSEYSLSLLASEMLLSGTGLSLATGEVSGDCFFFSAVRTD 476
Query: 432 DDKCFNESKCDPAADSDGVKV 452
+ + CDP +DG+ V
Sbjct: 475 ERAATAAALCDP*PFTDGLGV 413
>TC81004 weakly similar to SP|Q39529|LEC2_CLALU Agglutinin II precursor
(ClAII) (LecClAII). [Yellow wood] {Cladrastis lutea},
partial (24%)
Length = 610
Score = 32.0 bits (71), Expect = 1.6
Identities = 17/37 (45%), Positives = 24/37 (63%)
Frame = +1
Query: 1042 PESSSGTPISVRRIDSPSTPLRPIVNEVMAREEATFA 1078
PESSSG + + SP T +PI+N+++A E TFA
Sbjct: 139 PESSSGGYLG---LFSPETAFKPIINQIVAVEFDTFA 240
>TC77503 weakly similar to GP|16930417|gb|AAL31894.1 At2g24420/T28I24.15
{Arabidopsis thaliana}, partial (54%)
Length = 1787
Score = 32.0 bits (71), Expect = 1.6
Identities = 24/104 (23%), Positives = 50/104 (48%), Gaps = 3/104 (2%)
Frame = +2
Query: 738 IKHMQNSISSLQERGPSPKDQSM--LYVNVGVSNQTLEHHSSEVKGFSVNNGEVKQLLNM 795
I+ +QN ++SLQ++G ++ + Y G + ++ SE+ + N
Sbjct: 353 IQSLQNEVASLQKKGSLDAEEQVGKAYARAGELQKQVDKLKSEL-----------EAQNS 499
Query: 796 EKRKYVSRYPSEGQSSSQLDTDMQDLEE-NERSNGAIRKSDDAV 838
EK + SR + L++ ++D+++ NE IRK++ A+
Sbjct: 500 EKVNWESRVAKLEKKIHDLNSKLEDVQKINEEQKKQIRKTERAL 631
>TC91313 similar to GP|15292869|gb|AAK92805.1 unknown protein {Arabidopsis
thaliana}, partial (54%)
Length = 806
Score = 30.8 bits (68), Expect = 3.5
Identities = 14/46 (30%), Positives = 22/46 (47%)
Frame = +2
Query: 1130 FKKGILKRNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEV 1175
F+KG+ + C+C+ C RL+ R F+Y R +L V
Sbjct: 503 FQKGLKQNIHSATCSSCSCVQCVK*RLNLGRVFQYMRVRLTQGNRV 640
>BG586181 similar to GP|4689316|gb| peroxisomal targeting signal type 2
receptor {Arabidopsis thaliana}, partial (56%)
Length = 804
Score = 30.8 bits (68), Expect = 3.5
Identities = 17/59 (28%), Positives = 27/59 (44%), Gaps = 7/59 (11%)
Frame = -1
Query: 905 ITPVLNRCPRIKSLKHAGSPTYKRLLPLLSNAKKYNSCASVNRHD-------PKHQKFL 956
+ P L+RCPRI S+ P + +P ++ K +N +N + P HQ L
Sbjct: 561 LVPSLSRCPRISSM----LPNINKFIPTSTSKKPFNKNTHINPNSKLSMMIIPTHQSIL 397
>BQ137252 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (10%)
Length = 1067
Score = 30.8 bits (68), Expect = 3.5
Identities = 22/74 (29%), Positives = 29/74 (38%)
Frame = +1
Query: 30 PPGALTRSRSQLFVHRNRSGQRRPDSGPRQGGLYSGLGGSSGNDEKREPPEEDDGDTLLV 89
PP + R R R G+ D PR+G S GG G +R EE +
Sbjct: 19 PPDSTGRRDPAATEKREREGKGNEDDAPRRGKKNSAGGGREGRGAERGRGEESE------ 180
Query: 90 KYLRKKAKRDDDGG 103
RK+A+ GG
Sbjct: 181 --RRKRARGGGAGG 216
>AL389529
Length = 421
Score = 30.8 bits (68), Expect = 3.5
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = -3
Query: 259 SKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRN 298
+K+D + G L+ PFT E S E L+RK +KRN
Sbjct: 146 AKIDNFDDNGRXSLVTLXTPFTRAHEFSXELLARKFTKRN 27
>TC88610 homologue to PIR|G96755|G96755 developmental protein homolog DG1118
[imported] - Arabidopsis thaliana, partial (79%)
Length = 689
Score = 30.8 bits (68), Expect = 3.5
Identities = 25/116 (21%), Positives = 52/116 (44%), Gaps = 6/116 (5%)
Frame = +3
Query: 643 SNALGNILFNSANELSNGNLPKLTSSQDLPE-----LPMQSDVKE-VVQECLSAPSVEEP 696
S ++GNI+ + + L+ GNL K++ + D E + +Q++ E + S + E
Sbjct: 159 SKSMGNIVKSLESSLATGNLQKMSETMDQFEKQFVNMEVQAEFMESAMAGSTSLSTPEGE 338
Query: 697 IEKAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDGHDVDIKHMQNSISSLQERG 752
+ + D+ E + +L + H+ + D VD + ++ L+ RG
Sbjct: 339 VNSLMQQVADDYGLEVSVNL--PVAAAHAVPVKEKDAEKVDEDDLSRRLAELKARG 500
>TC77784 similar to GP|14272369|emb|CAC40022. bZIP transcription factor
{Arabidopsis thaliana}, partial (72%)
Length = 2052
Score = 30.4 bits (67), Expect = 4.5
Identities = 21/66 (31%), Positives = 32/66 (47%), Gaps = 2/66 (3%)
Frame = +3
Query: 346 NWPDNFVGTKKLGHCDKDEKGMGGQDFQLPLSSQSEKPSEPE--LKSESFTMRETNGSKL 403
+WP F T +G +K GG+ LSS++E P EPE L S+ + ++ +
Sbjct: 552 SWPMRFHQTSTVGGGNKS----GGESSDSALSSKNENPFEPESPLSSKKASSFSSDHNNN 719
Query: 404 SSSQNL 409
QNL
Sbjct: 720 MDQQNL 737
>TC82780 similar to PIR|T00713|T00713 helicase homolog F22O13.8 -
Arabidopsis thaliana, partial (5%)
Length = 563
Score = 30.4 bits (67), Expect = 4.5
Identities = 20/97 (20%), Positives = 43/97 (43%)
Frame = +2
Query: 767 VSNQTLEHHSSEVKGFSVNNGEVKQLLNMEKRKYVSRYPSEGQSSSQLDTDMQDLEENER 826
++ +L+H G S ++ ++ LL ++++ Y + + + EE +
Sbjct: 128 LAGDSLKHTVPHSNGSSYSDKLMESLLGKHHPRWIANYHLHESLLQENEEEKLSKEEQDM 307
Query: 827 SNGAIRKSDDAVVLNHVGVSNEVPRPPPENIISKTQD 863
+ RKS + + V + VP P PE +SK ++
Sbjct: 308 AWEVYRKSLEWEEVQRVPIGESVPDPKPE--VSKAEE 412
>TC77199 similar to SP|O04379|AGO1_ARATH Argonaute protein. [Mouse-ear
cress] {Arabidopsis thaliana}, partial (74%)
Length = 2714
Score = 30.0 bits (66), Expect = 5.9
Identities = 15/62 (24%), Positives = 26/62 (41%)
Frame = -2
Query: 685 QECLSAPSVEEPIEKAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDGHDVDIKHMQNS 744
+EC + S+ A+ + NFD + D HS I G D+ +K + N
Sbjct: 568 REC*FSGSLGSQTSNAIFSPHIATMCDFNFDTTKCLFDLHSISIRQRPGRDIPVKQLGNE 389
Query: 745 IS 746
++
Sbjct: 388 LN 383
>TC88994 similar to GP|20260620|gb|AAM13208.1 unknown protein {Arabidopsis
thaliana}, partial (16%)
Length = 888
Score = 29.6 bits (65), Expect = 7.7
Identities = 23/87 (26%), Positives = 43/87 (48%), Gaps = 3/87 (3%)
Frame = +2
Query: 696 PIEKAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDG---HDVDIKHMQNSISSLQERG 752
P+++A+V E ++EPNF+L + D + + +DG DV + + +Q +
Sbjct: 308 PLQEAMVKDTLERTKEPNFNLKDFLHDAYIHPVFNDDGDTDSDVMSQEWKEEPVIVQTKR 487
Query: 753 PSPKDQSMLYVNVGVSNQTLEHHSSEV 779
S K+ + G S QT H +++V
Sbjct: 488 QSRKNTPAPSKHSGGSLQTSMHGTTDV 568
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.131 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,528,702
Number of Sequences: 36976
Number of extensions: 493227
Number of successful extensions: 1979
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1954
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1978
length of query: 1262
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1155
effective length of database: 5,058,295
effective search space: 5842330725
effective search space used: 5842330725
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0108.13


