

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0106.15
(120 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
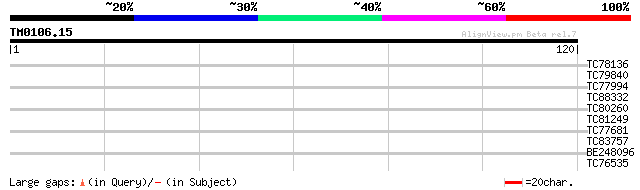
Score E
Sequences producing significant alignments: (bits) Value
TC78136 similar to GP|22652297|gb|AAN03675.1 putative nucleopori... 34 0.015
TC79840 similar to PIR|A86211|A86211 hypothetical protein [impor... 28 0.82
TC77994 weakly similar to PIR|T39903|T39903 serine-rich protein ... 26 3.1
TC88332 similar to GP|10177796|dbj|BAB11287. gb|AAF31027.1~gene_... 26 4.1
TC80260 similar to GP|5059352|gb|AAD38983.1| syntaxin of plants ... 26 4.1
TC81249 similar to GP|14517556|gb|AAK62668.1 AT3g61710/F15G16_10... 25 5.3
TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone d... 25 5.3
TC83757 similar to GP|10177454|dbj|BAB10845. contains similarity... 25 6.9
BE248096 similar to GP|20160913|dbj contains EST C28023(C53723)~... 25 6.9
TC76535 similar to SP|Q9BBN6|YCF1_LOTJA Hypothetical 214.8 kDa p... 25 6.9
>TC78136 similar to GP|22652297|gb|AAN03675.1 putative nucleoporin PRECOZ
{Arabidopsis thaliana}, partial (48%)
Length = 2532
Score = 33.9 bits (76), Expect = 0.015
Identities = 25/96 (26%), Positives = 47/96 (48%), Gaps = 14/96 (14%)
Frame = +3
Query: 13 AEGDHIKVDRVKEVDNEVDD-------MKEWGHILKED-----EDLMNAIDDMNALSHVV 60
A+ + ++ +KEVDN+VD E+ + ++E ++ +++++D + L HV
Sbjct: 606 ADNEEDMMEDIKEVDNDVDQEALPLIRRAEFSYWMRESVSYHVQNQISSLNDSHYLQHVF 785
Query: 61 MKFQNRQVFDWAQLVGKNNDFVVIMIRSDYG--TLN 94
RQ+ + QL N D + + S G TLN
Sbjct: 786 TLLTGRQLDEAVQLAVSNGDVRLACLLSQAGGSTLN 893
>TC79840 similar to PIR|A86211|A86211 hypothetical protein [imported] -
Arabidopsis thaliana, partial (22%)
Length = 841
Score = 28.1 bits (61), Expect = 0.82
Identities = 17/61 (27%), Positives = 30/61 (48%), Gaps = 9/61 (14%)
Frame = +1
Query: 1 MSHVLIEKVD---------NVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAID 51
M L EKVD + G H +++ ++ E + +W H LKE+ ++M+ ID
Sbjct: 193 MHGYLTEKVDVYSFGVVALEIVSGKHNTMNKPRD---ECFSLVDWVHFLKEEGNIMDLID 363
Query: 52 D 52
+
Sbjct: 364 E 366
>TC77994 weakly similar to PIR|T39903|T39903 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (5%)
Length = 1571
Score = 26.2 bits (56), Expect = 3.1
Identities = 21/84 (25%), Positives = 34/84 (40%)
Frame = -1
Query: 20 VDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVFDWAQLVGKNN 79
V++ KE +NE +W EDED + + ++ DW + VG +
Sbjct: 1379 VEKAKECENE-----DWP---PEDEDEVEK----------ERSEERKRAEDWKKTVGGSR 1254
Query: 80 DFVVIMIRSDYGTLNGKESLTLGC 103
DF M+R+ + LGC
Sbjct: 1253 DFYYFMLRASMIRAEIRAEGLLGC 1182
>TC88332 similar to GP|10177796|dbj|BAB11287.
gb|AAF31027.1~gene_id:MXA21.11~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (57%)
Length = 698
Score = 25.8 bits (55), Expect = 4.1
Identities = 12/38 (31%), Positives = 19/38 (49%)
Frame = +3
Query: 51 DDMNALSHVVMKFQNRQVFDWAQLVGKNNDFVVIMIRS 88
DD + HV+ +N +FDW + KN I++ S
Sbjct: 540 DDYDGHRHVIKVVKNTPLFDWFKDSLKNEGMDEILVNS 653
>TC80260 similar to GP|5059352|gb|AAD38983.1| syntaxin of plants 42
{Arabidopsis thaliana}, partial (83%)
Length = 995
Score = 25.8 bits (55), Expect = 4.1
Identities = 9/36 (25%), Positives = 20/36 (55%)
Frame = +2
Query: 7 EKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKE 42
E++ + + +HI +R +E+D + E I+K+
Sbjct: 782 EQMTKLKKNEHISAEREREIDQVAKSVHELAQIMKD 889
>TC81249 similar to GP|14517556|gb|AAK62668.1 AT3g61710/F15G16_100
{Arabidopsis thaliana}, partial (41%)
Length = 813
Score = 25.4 bits (54), Expect = 5.3
Identities = 12/37 (32%), Positives = 17/37 (45%)
Frame = +1
Query: 81 FVVIMIRSDYGTLNGKESLTLGCEHGGKFKPYKGKFD 117
FVV+ G SL+ G +HGG P+ F+
Sbjct: 316 FVVVYKSESASDGGGGNSLSPGVDHGGHLPPHNSGFN 426
>TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone deacetylase
HD2-P39 {Glycine max}, partial (47%)
Length = 1233
Score = 25.4 bits (54), Expect = 5.3
Identities = 14/42 (33%), Positives = 22/42 (52%)
Frame = +2
Query: 11 NVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDD 52
N A H+KV K+ D++ DD + I D+++ NA D
Sbjct: 527 NAATAKHVKVVDPKDEDSDDDDESD-DEIGSSDDEMENADSD 649
>TC83757 similar to GP|10177454|dbj|BAB10845. contains similarity to nuclear
protein ZAP~gene_id:MQB2.8 {Arabidopsis thaliana},
partial (10%)
Length = 995
Score = 25.0 bits (53), Expect = 6.9
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = +3
Query: 15 GDHIKVDRVKEVDNEVDD 32
GD +K R++EVD ++DD
Sbjct: 705 GDDLKESRIQEVDMDMDD 758
>BE248096 similar to GP|20160913|dbj contains EST C28023(C53723)~similar to
sperm protein homolog~unknown protein, partial (49%)
Length = 318
Score = 25.0 bits (53), Expect = 6.9
Identities = 18/59 (30%), Positives = 28/59 (46%), Gaps = 1/59 (1%)
Frame = +3
Query: 52 DMNALSH-VVMKFQNRQVFDWAQLVGKNNDFVVIMIRSDYGTLNGKESLTLGCEHGGKF 109
+ N+L H +++++ Q+FD V +DFV RS+ N S E G KF
Sbjct: 42 EANSLIHSLIIRYTPEQLFDVVAAVDYYHDFVPWCQRSEIVKRNPDGSFDAELEIGFKF 218
>TC76535 similar to SP|Q9BBN6|YCF1_LOTJA Hypothetical 214.8 kDa protein ycf1.
{Lotus japonicus}, partial (16%)
Length = 1658
Score = 25.0 bits (53), Expect = 6.9
Identities = 11/42 (26%), Positives = 24/42 (56%), Gaps = 3/42 (7%)
Frame = +3
Query: 10 DNVAEGDHIKVDRVK---EVDNEVDDMKEWGHILKEDEDLMN 48
D + E D + D + E+D E+D +E ++++ED+++
Sbjct: 993 DEILEYDEKQKDEINGETEIDVEIDSEQEQNGSIEDEEDILS 1118
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.139 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,945,808
Number of Sequences: 36976
Number of extensions: 29508
Number of successful extensions: 129
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 128
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 129
length of query: 120
length of database: 9,014,727
effective HSP length: 84
effective length of query: 36
effective length of database: 5,908,743
effective search space: 212714748
effective search space used: 212714748
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0106.15


