

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0105.5
(231 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
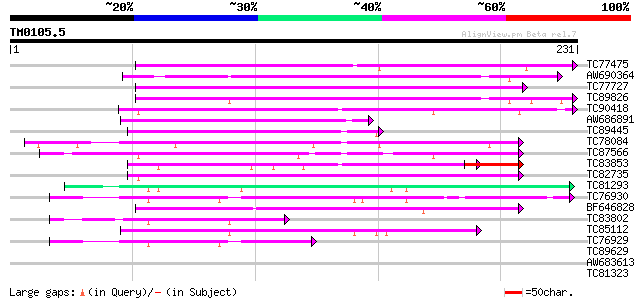
TC77475 similar to GP|6041819|gb|AAF02134.1| unknown protein {Ar... 125 2e-29
AW690364 weakly similar to GP|9759067|dbj contains similarity to... 109 1e-24
TC77727 weakly similar to GP|6041819|gb|AAF02134.1| unknown prot... 106 9e-24
TC89826 weakly similar to PIR|T49138|T49138 hypothetical protein... 92 2e-19
TC90418 similar to GP|11762174|gb|AAG40365.1 At1g17620 {Arabidop... 76 9e-15
AW686891 weakly similar to GP|10177234|db gb|AAF18257.1~gene_id:... 65 2e-11
TC89445 similar to GP|7417006|gb|AAF62403.1| harpin inducing pro... 64 5e-11
TC78084 weakly similar to GP|13877665|gb|AAK43910.1 Unknown prot... 62 2e-10
TC87566 weakly similar to GP|21554256|gb|AAM63331.1 unknown {Ara... 61 4e-10
TC83853 weakly similar to GP|13877665|gb|AAK43910.1 Unknown prot... 50 3e-08
TC82735 weakly similar to GP|6686408|gb|AAF23842.1| F1E22.7 {Ara... 52 1e-07
TC81293 similar to GP|19911581|dbj|BAB86894. syringolide-induced... 50 1e-06
TC76930 similar to GP|19911581|dbj|BAB86894. syringolide-induced... 48 3e-06
BF646828 similar to GP|19911581|db syringolide-induced protein B... 44 7e-05
TC83802 similar to GP|7417006|gb|AAF62403.1| harpin inducing pro... 44 7e-05
TC85112 similar to GP|19911581|dbj|BAB86894. syringolide-induced... 43 1e-04
TC76929 similar to GP|19911581|dbj|BAB86894. syringolide-induced... 40 8e-04
TC89629 weakly similar to GP|11994147|dbj|BAB01168. non-race spe... 40 0.001
AW683613 weakly similar to GP|21536796|gb| unknown {Arabidopsis ... 35 0.032
TC81323 similar to GP|11762174|gb|AAG40365.1 At1g17620 {Arabidop... 34 0.054
>TC77475 similar to GP|6041819|gb|AAF02134.1| unknown protein {Arabidopsis
thaliana}, partial (83%)
Length = 1128
Score = 125 bits (313), Expect = 2e-29
Identities = 67/188 (35%), Positives = 104/188 (54%), Gaps = 8/188 (4%)
Frame = +3
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKR 111
I +FL V +T+L++W V +P KP F + VYA N T P L++ Q V RNPN +
Sbjct: 237 IIIFLFIVLVTILIIWAVLKPSKPSFILQDVTVYAFNATVPNFLTSNFQVTVSSRNPNDK 416
Query: 112 VSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVANGLAM 170
+ IYYDR ++V+YR+Q IT + +PP + + H +V + SP + GT +PV + L+
Sbjct: 417 IGIYYDRLDSYVTYRSQQITYRTAIPPSY-QGHKEVDVWSPFVYGTNVPVAPYNSVALSQ 593
Query: 171 DESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLV-------GLKKGFVGQVPLLQ 223
D+ G V + + F GRV+WK G + H+ ++V+C + G+ G G V
Sbjct: 594 DQDNGNVLVVVKFDGRVRWKVGAFISGHYHIFVRCPAFITFGPRSNGISVGDSGAVKYQI 773
Query: 224 AQACDVDL 231
Q C V +
Sbjct: 774 VQRCTVSV 797
>AW690364 weakly similar to GP|9759067|dbj contains similarity to
harpin-induced protein~gene_id:MGN6.8 {Arabidopsis
thaliana}, partial (48%)
Length = 684
Score = 109 bits (272), Expect = 1e-24
Identities = 63/182 (34%), Positives = 99/182 (53%), Gaps = 3/182 (1%)
Frame = +3
Query: 47 AACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIR 106
A TF T LL + L+++ + +P KP+F++ +Y LN S P+L++++Q +L +
Sbjct: 99 AFSTFFTTILLLI----LLIYFILKPSKPQFSLQELDIYQLNL-SGPILNSSIQLTLLSK 263
Query: 107 NPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVAN 166
NPN++VSIYYD F + +Y+NQ IT +PP + LS + G +PV +
Sbjct: 264 NPNQKVSIYYDEFQVYATYKNQQITSDSFVPPFYQGTQESNFLSSSLIGNGLPVAPSIGY 443
Query: 167 GLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLY---VKCDLLVGLKKGFVGQVPLLQ 223
L+ D+ G +GL L G+++WK G TW G Y V CD +V G VP L
Sbjct: 444 ELSRDQVSGRLGLSLKANGKLRWKIG---TWVSGRYRFNVNCDSIVAFGIGPTLNVPPLN 614
Query: 224 AQ 225
++
Sbjct: 615 SK 620
>TC77727 weakly similar to GP|6041819|gb|AAF02134.1| unknown protein
{Arabidopsis thaliana}, partial (93%)
Length = 1078
Score = 106 bits (264), Expect = 9e-24
Identities = 51/160 (31%), Positives = 89/160 (54%)
Frame = +3
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKR 111
I VFL V +T+L++W + RP KP F ++ +YA N + P L++ Q + RNPN
Sbjct: 243 IMVFLFLVLLTILLIWAILRPTKPTFILLDVTLYAFNASQPNFLTSNFQVTLSSRNPNDN 422
Query: 112 VSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVANGLAMD 171
+ +YYDR +++YR+Q IT + +PP + + SP + G +PV + L+ D
Sbjct: 423 IGVYYDRLDTYMTYRSQQITLRTAIPPSYQGHNEYDVWSPFVYGNDIPVAPFNSVSLSQD 602
Query: 172 ESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGL 211
E+ G + + + GRV++K G + + ++V+C + L
Sbjct: 603 ENNGNIFVTVRVDGRVRFKVGAFISGRYHIHVRCPAYISL 722
>TC89826 weakly similar to PIR|T49138|T49138 hypothetical protein T10D17.10
- Arabidopsis thaliana, partial (59%)
Length = 1130
Score = 92.0 bits (227), Expect = 2e-19
Identities = 59/202 (29%), Positives = 96/202 (47%), Gaps = 22/202 (10%)
Frame = +2
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALN-----------TTSPPLLSAALQ 100
I F++ + + ++W++ RP KPRF + A V+A N T +P ++ +Q
Sbjct: 152 ILSFIILILFVIFLIWIILRPTKPRFILQDATVFAFNLSSTGETPSSTTPTPNTITLTIQ 331
Query: 101 FNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPV 160
+ NPN ++ IYY + A+ SYR Q I+ LP + SP++ G +PV
Sbjct: 332 VTLSSFNPNSKIGIYYHKLDAYASYRGQQISLATELPQTYQGHRDVAVWSPILYGAAVPV 511
Query: 161 TVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLY---VKCDLLVGL-----K 212
+ ++ L D++ G V + + GRVKWK G TW G Y V C + +
Sbjct: 512 SPYLSEILRQDQTSGGVLVNVKVNGRVKWKVG---TWVSGRYHIDVNCPAFIKVAGDKGD 682
Query: 213 KGFVGQVPLLQ---AQACDVDL 231
GF P ++ Q+C VD+
Sbjct: 683 DGFGVSAPAVKFQFLQSCVVDV 748
>TC90418 similar to GP|11762174|gb|AAG40365.1 At1g17620 {Arabidopsis
thaliana}, partial (9%)
Length = 835
Score = 76.3 bits (186), Expect = 9e-15
Identities = 57/201 (28%), Positives = 93/201 (45%), Gaps = 14/201 (6%)
Frame = +3
Query: 45 KRAACTF-------ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSA 97
+R CTF + + LL +G+T +L+YRPH+P FTV + LN TS L++
Sbjct: 183 RRCCCTFFFWFFLILLILLLLLGVTGTAFYLIYRPHRPSFTVTSLKLSYLNLTSSSTLNS 362
Query: 98 ALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTP 157
N+ +NPNK ++ Y + + N I + +P K + L I T
Sbjct: 363 KFNVNITAKNPNKHITFVYQPTTITILSNNINIGEGT-IPSFKHGKKNTTLLKSSISSTG 539
Query: 158 MPVTVEVANGLAMD-ESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCD------LLVG 210
+ + E + L + +S + LK+ +VK K G ++T + G+ V CD +VG
Sbjct: 540 LALESEASTELKKNMKSKNGLALKVKLDTKVKAKMGKMKTPNVGIRVTCDGVRVHVPVVG 719
Query: 211 LKKGFVGQVPLLQAQACDVDL 231
KK ++ CDVD+
Sbjct: 720 SKKPVTASTSNVK---CDVDV 773
>AW686891 weakly similar to GP|10177234|db
gb|AAF18257.1~gene_id:MRN17.10~similar to unknown
protein {Arabidopsis thaliana}, partial (64%)
Length = 482
Score = 65.1 bits (157), Expect = 2e-11
Identities = 32/103 (31%), Positives = 52/103 (50%)
Frame = +3
Query: 46 RAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLI 105
R I ++ VG+ +L+ WLV RP +++V A+++ N T L A F +
Sbjct: 84 RYIAIIILALIIIVGLAVLISWLVVRPKNLQYSVEDASIHNFNLTDANHLYANFDFTIRS 263
Query: 106 RNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVS 148
+NPN +VS+YYD V Y +Q + + P F + H V+
Sbjct: 264 KNPNSKVSLYYDSIEVSVRYEDQTLATNAVQP--FFQPHKNVT 386
>TC89445 similar to GP|7417006|gb|AAF62403.1| harpin inducing protein
{Nicotiana tabacum}, partial (36%)
Length = 651
Score = 63.9 bits (154), Expect = 5e-11
Identities = 36/105 (34%), Positives = 50/105 (47%), Gaps = 1/105 (0%)
Frame = +1
Query: 49 CTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNP 108
C IT ++ + I + WL+ RP+ +FTV A + N L L NV +RNP
Sbjct: 232 CKIITTIIVIIVILGFIFWLIVRPNVVKFTVNDATLTQFNFNETNTLHYDLALNVTVRNP 411
Query: 109 NKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVS-LSPV 152
N+RV IYYD Y++ Q L F + H S L+PV
Sbjct: 412 NRRVGIYYDTIETMAFYKDVQFANQTL--GRFFQHHKNTSFLNPV 540
>TC78084 weakly similar to GP|13877665|gb|AAK43910.1 Unknown protein
{Arabidopsis thaliana}, partial (46%)
Length = 832
Score = 62.0 bits (149), Expect = 2e-10
Identities = 58/222 (26%), Positives = 95/222 (42%), Gaps = 19/222 (8%)
Frame = +1
Query: 7 PPKG-------QNQHHHLPPPNKIQMN-NHKASPSPYGGGGGGSSPKRAACTFITVFLLA 58
PP G ++Q + +PPP Q N+ S +R C + +
Sbjct: 76 PPSGTYVIQLPKDQTYRIPPPENAQRYANYTRRKS-----------RRCGCCCCLCWFIG 222
Query: 59 VGITLLVL--------WLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNK 110
+ TL+VL +L+ RP P++ + +V +N TS +S +V N NK
Sbjct: 223 ILFTLIVLMAIAAGVFYLIVRPEAPKYAIDRVSVKGMNLTSLSTISPEFDVSVKADNGNK 402
Query: 111 RVSIYYDRFSAF-VSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVANGL 168
++ IYY+ S+ + YR + + VL P F + + V++ V+ G + + L
Sbjct: 403 KIGIYYESDSSVEMFYRGVSLCKGVL--PAFYQPSNNVTVFQTVLKGNGIELVRSDQRAL 576
Query: 169 AMDESYGVVGLKLVFLGRVKWKAGGL-RTWHHGLYVKCDLLV 209
+ G V L L VK K G + +TW + V CDL V
Sbjct: 577 VNAVTKGSVPLTLKLRAPVKIKVGSVKKTWKVRVKVDCDLTV 702
>TC87566 weakly similar to GP|21554256|gb|AAM63331.1 unknown {Arabidopsis
thaliana}, partial (44%)
Length = 1042
Score = 60.8 bits (146), Expect = 4e-10
Identities = 54/212 (25%), Positives = 94/212 (43%), Gaps = 15/212 (7%)
Frame = +3
Query: 13 QHHHLPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACTF-------ITVFLLAVGITLLV 65
+ H PPP + + P P + C F + + ++A+GIT+ +
Sbjct: 219 EQQHPPPPFP---STQQTLPIPINQSKPPKKRRSCCCKFFCWIFSILIILIIALGITIGI 389
Query: 66 LWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSIYY---DRFSAF 122
L+LV+RP P+++V V + ++ L+ + RNPNK++ I Y R SA+
Sbjct: 390 LFLVFRPKIPKYSVDELRVTQFDLSNNNSLAVTFNLTITARNPNKKIGIDYRGGSRISAW 569
Query: 123 VSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVANGLAMD-----ESYGVV 177
Y + + + L P F + H V++ + P+ + A GL+ +S G V
Sbjct: 570 --YIDTKLCEGSL--PKFYQGHKNVTVLSI----PLTGQTQNATGLSSSLQQQFQSEGSV 725
Query: 178 GLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLV 209
LKL V+ K G L+ V+C ++V
Sbjct: 726 PLKLNVKQPVRIKLGKLKLPKINFRVRCTIVV 821
>TC83853 weakly similar to GP|13877665|gb|AAK43910.1 Unknown protein
{Arabidopsis thaliana}, partial (49%)
Length = 815
Score = 50.4 bits (119), Expect(2) = 3e-08
Identities = 36/151 (23%), Positives = 72/151 (46%), Gaps = 7/151 (4%)
Frame = +1
Query: 49 CTFITVFLLAV---GITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSA-ALQFNVL 104
C FI + + + GI + +LV+RP P +T+ + +N TSP + + +F+V
Sbjct: 274 CWFIGIIFILIALLGIAAGIFYLVFRPKAPNYTIENITIRGINITSPSSTTGISPEFDVT 453
Query: 105 IR--NPNKRVSIYYDR-FSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVT 161
++ NPN ++ I Y++ SA + Y++ + + LP + ++ ++ G + ++
Sbjct: 454 VKADNPNDKIGISYEKDSSAEIFYKDMRLCNGI-LPSFYQPSNNVTVFKTMLKGNGVKMS 630
Query: 162 VEVANGLAMDESYGVVGLKLVFLGRVKWKAG 192
E L ++ V L + VK K G
Sbjct: 631 SEDQRALVKAQTKQEVQLMVKLRAPVKIKVG 723
Score = 23.9 bits (50), Expect(2) = 3e-08
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = +2
Query: 186 RVKWKAGGLRTWHHGLYVKCDLLV 209
R K K G ++TW + V CDL+V
Sbjct: 704 R*K*KWGSVKTWKITVKVDCDLMV 775
>TC82735 weakly similar to GP|6686408|gb|AAF23842.1| F1E22.7 {Arabidopsis
thaliana}, partial (74%)
Length = 965
Score = 52.4 bits (124), Expect = 1e-07
Identities = 39/163 (23%), Positives = 69/163 (41%), Gaps = 2/163 (1%)
Frame = +1
Query: 49 CTF--ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIR 106
CTF + + ++A+GIT+ L+L +RP +++V + N + L V R
Sbjct: 259 CTFTILLILIIAIGITVGALYLAFRPKLTKYSVDRLRITQFNLSDDNSLFVTFNVTVTAR 438
Query: 107 NPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVAN 166
NPNK++ IYY S ++ + + LP + + L+ + G T
Sbjct: 439 NPNKKIGIYYVSGSHISAWYKETGLCEGSLPKFYQGHRNTTVLNLPLTGQTQDATGLFNT 618
Query: 167 GLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLV 209
+ G + L + V+ K G L+ + V+C L V
Sbjct: 619 LQQQLQEAGNIPLDIKVNQNVRVKLGKLKIFRVKFRVRCSLQV 747
>TC81293 similar to GP|19911581|dbj|BAB86894. syringolide-induced protein
B13-1-9 {Glycine max}, partial (60%)
Length = 799
Score = 49.7 bits (117), Expect = 1e-06
Identities = 52/220 (23%), Positives = 87/220 (38%), Gaps = 12/220 (5%)
Frame = +1
Query: 23 IQMNNHKASPSPYGGGGGGSSPKRAACTFITVF---LLAV----GITLLVLWLVYRPHKP 75
++ N A P+ G +R C + LLA+ G+ +L+ WLV +P
Sbjct: 31 LKYRN*MAEAQPHNRG------RRCCCCLFGIIWKLLLAIIVLAGLIILIFWLVVQPRTF 192
Query: 76 RFTVVGAAVYALNTTSPP-LLSAALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQV 134
+F+V A + N + L L N RNPNK+++IYYD SY
Sbjct: 193 KFSVKEAKLTKFNYSDDTNTLHYNLVLNFTARNPNKKLNIYYDVIEGHASYEGTRFASTK 372
Query: 135 LLPPLFLEKHSQVSLSPVIG---GTPMPV-TVEVANGLAMDESYGVVGLKLVFLGRVKWK 190
++ L + S +P+ G G + V + + DE GV + + ++++
Sbjct: 373 VITWLNSFRQYTKSTNPMSGVFSGQRVVVFDRDQVSNFERDEKDGVFHINVKLNFDMRFR 552
Query: 191 AGGLRTWHHGLYVKCDLLVGLKKGFVGQVPLLQAQACDVD 230
G H +KC L V + + CDV+
Sbjct: 553 LGDYIGPHTKGNIKCRLDVPFVANGTKVMKAFEPTTCDVN 672
>TC76930 similar to GP|19911581|dbj|BAB86894. syringolide-induced protein
B13-1-9 {Glycine max}, partial (87%)
Length = 996
Score = 48.1 bits (113), Expect = 3e-06
Identities = 57/232 (24%), Positives = 96/232 (40%), Gaps = 18/232 (7%)
Frame = +2
Query: 17 LPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACTFITVF-------LLAVGITLLVLWLV 69
+PPP + + +H++ + C + F ++ G+ +L+ +LV
Sbjct: 185 IPPPAQPRPRSHRS--------------RSCCCCLFSFFWKLLIALIVLAGLAVLIFYLV 322
Query: 70 YRPHKPRFTVVGAAV----YALNTTSPPLLSAALQFNVLIRNPNKRVSIYYDRFSAFVSY 125
+P +F V A + Y+ NT L+ + N RNPNK+++IYYD+ A Y
Sbjct: 323 VQPRGFKFYVDEANLTEFDYSNNT-----LNYNMVLNFTARNPNKKLNIYYDKVEARAFY 487
Query: 126 RNQPITQQVLLPPL--FLE-KHSQVSLSPVIGGTPMPV----TVEVANGLAMDESYGVVG 178
++ + F + K S +S V G + V V N DE Y +
Sbjct: 488 EGSRFANVDVITHMNSFRQYKKSSNPMSAVFSGQHVFVLDSDQVSEFNKDKSDEVYDIY- 664
Query: 179 LKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKKGFVGQVPLLQAQACDVD 230
++L F R++++ G L + + VKCD V L CDVD
Sbjct: 665 VRLYF--RIRFRLGDLISGDYKPKVKCDFKVPLSSKISNAT--FTRTKCDVD 808
>BF646828 similar to GP|19911581|db syringolide-induced protein B13-1-9
{Glycine max}, partial (60%)
Length = 648
Score = 43.5 bits (101), Expect = 7e-05
Identities = 35/162 (21%), Positives = 70/162 (42%), Gaps = 4/162 (2%)
Frame = +1
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKR 111
I ++ G+ +L+ +L+ +P +F V A + + T+ L + N NPNK+
Sbjct: 163 IIALIVLFGLIILIFFLIVQPRAFKFYVTKAELTQFDHTNNTLYYNMV-LNFTSHNPNKK 339
Query: 112 VSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVANG---- 167
+ IYYD+ A Y + ++ + + + + +P+ G + + N
Sbjct: 340 LGIYYDKVEAQAFYEGSRFSNVDVITHMNSFRQDKKNSNPMTGVFSGQKLLMLDNDQISE 519
Query: 168 LAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLV 209
D++ GV + + +++K G + VKCDL V
Sbjct: 520 FNKDKNVGVYDIYVKLYFXIRFKLGDSISRKFKPKVKCDLTV 645
>TC83802 similar to GP|7417006|gb|AAF62403.1| harpin inducing protein
{Nicotiana tabacum}, partial (19%)
Length = 435
Score = 43.5 bits (101), Expect = 7e-05
Identities = 30/105 (28%), Positives = 48/105 (45%), Gaps = 7/105 (6%)
Frame = +2
Query: 17 LPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACTFITVF------LLAVGITLLVLWLVY 70
+PPP + + P GGG C F +F ++ +GI + + WL+
Sbjct: 149 IPPPTR-------SYNRPSRGGGFDCC---CGCIFNLIFKLILTVIIIIGIAVFLFWLIV 298
Query: 71 RPHKPRFTVVGAAVYALN-TTSPPLLSAALQFNVLIRNPNKRVSI 114
RP+ + V A + N T + L+ L N+ IRNPN+R+ I
Sbjct: 299 RPNAVKVHVTDATLTQFNYTNNNSNLNYNLALNITIRNPNRRLGI 433
>TC85112 similar to GP|19911581|dbj|BAB86894. syringolide-induced protein
B13-1-9 {Glycine max}, partial (27%)
Length = 597
Score = 42.7 bits (99), Expect = 1e-04
Identities = 36/152 (23%), Positives = 67/152 (43%), Gaps = 5/152 (3%)
Frame = +1
Query: 46 RAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALN-TTSPPLLSAALQFNVL 104
R + ++ V + +LV++++ P +F V A + N T + L+ L N
Sbjct: 127 RVLWIILVTIIILVSLIILVIYIIITPRSFKFHVNQAKLTQFNFTNNNTTLNYNLVLNFT 306
Query: 105 IRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPL-FLEKHSQVS--LSPV-IGGTPMPV 160
+NPNK++ IYYD A Y+ + + PL L++ V +S V IG M +
Sbjct: 307 AQNPNKKLKIYYDVVEANAFYKGYNFSATDMNKPLRTLQETKSVDYRMSAVFIGQHVMML 486
Query: 161 TVEVANGLAMDESYGVVGLKLVFLGRVKWKAG 192
+ + D G+ G+ + +++ G
Sbjct: 487 DRDEVDEFQEDYXNGIFGIDIXIYFXIRFXLG 582
>TC76929 similar to GP|19911581|dbj|BAB86894. syringolide-induced protein
B13-1-9 {Glycine max}, partial (79%)
Length = 767
Score = 40.0 bits (92), Expect = 8e-04
Identities = 30/120 (25%), Positives = 54/120 (45%), Gaps = 11/120 (9%)
Frame = +2
Query: 17 LPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACTFITVF-------LLAVGITLLVLWLV 69
+PPP + + +H++ + C + F ++ G+ +L+ +LV
Sbjct: 161 IPPPAQPRPRSHRS--------------RSCCCCLFSFFWKLLIALIVLAGLAVLIFYLV 298
Query: 70 YRPHKPRFTVVGAAV----YALNTTSPPLLSAALQFNVLIRNPNKRVSIYYDRFSAFVSY 125
+P +F V A + Y+ NT L+ + N RNPNK+++IYYD+ A Y
Sbjct: 299 VQPRGFKFYVDEANLTEFDYSNNT-----LNYNMVLNFTARNPNKKLNIYYDKVEARAFY 463
>TC89629 weakly similar to GP|11994147|dbj|BAB01168. non-race specific
disease resistance protein {Arabidopsis thaliana},
partial (25%)
Length = 864
Score = 39.7 bits (91), Expect = 0.001
Identities = 25/89 (28%), Positives = 42/89 (47%), Gaps = 4/89 (4%)
Frame = +3
Query: 33 SPYGGGGGGSSPKRAACTFITVFLLAVGITLLVLWLVYRPHKPRFTV----VGAAVYALN 88
+P SS + C+ + F++ +G+ L +WL R +P+ + + A ALN
Sbjct: 27 NPTASNQMASSKPNSCCSCCSGFIITIGLMALFIWLSLRVDEPKLYIDKIYLPALNKALN 206
Query: 89 TTSPPLLSAALQFNVLIRNPNKRVSIYYD 117
TT+ + F + + NPNK I YD
Sbjct: 207 TTAKS--NTTFTFLLKLVNPNKDKGIQYD 287
>AW683613 weakly similar to GP|21536796|gb| unknown {Arabidopsis thaliana},
partial (20%)
Length = 503
Score = 34.7 bits (78), Expect = 0.032
Identities = 29/140 (20%), Positives = 52/140 (36%), Gaps = 21/140 (15%)
Frame = +2
Query: 1 MEEDPNPPKGQNQHHHL--------PPPNKIQMNNHKAS---PSPYGGGGGGSSPKRAAC 49
M E P+ P G + + P P K Q+ N P Y + C
Sbjct: 83 MTEPPSKPNGNGAINGVAATNGNGNPAPVKSQLYNPNRQVYRPQQYNRRRRSNRSLCCCC 262
Query: 50 TFITVFLLA-----VGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNV- 103
F T+ + V I V +++Y PH+P+F++ + +N +P + + +
Sbjct: 263 CFWTILTVLAAAFLVAIVGAVFYVLYHPHQPQFSITNLRIAKMNLKTPA*FTFTSHYTLA 442
Query: 104 ----LIRNPNKRVSIYYDRF 119
+NPN + + F
Sbjct: 443 TSHSCAKNPNNHLVLLLRAF 502
>TC81323 similar to GP|11762174|gb|AAG40365.1 At1g17620 {Arabidopsis
thaliana}, partial (7%)
Length = 1014
Score = 33.9 bits (76), Expect = 0.054
Identities = 18/65 (27%), Positives = 31/65 (47%)
Frame = +1
Query: 56 LLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSIY 115
L+ +G+ V++ +Y P +P FTV TS L S+ + NPN++++
Sbjct: 277 LILLGVAGTVVYFLYHPQRPSFTV----------TSLKLTSSEFDLTLSTTNPNEKITFS 426
Query: 116 YDRFS 120
Y S
Sbjct: 427 YQPIS 441
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,905,887
Number of Sequences: 36976
Number of extensions: 152264
Number of successful extensions: 1539
Number of sequences better than 10.0: 78
Number of HSP's better than 10.0 without gapping: 1384
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1497
length of query: 231
length of database: 9,014,727
effective HSP length: 93
effective length of query: 138
effective length of database: 5,575,959
effective search space: 769482342
effective search space used: 769482342
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0105.5


