

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0105.17
(488 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
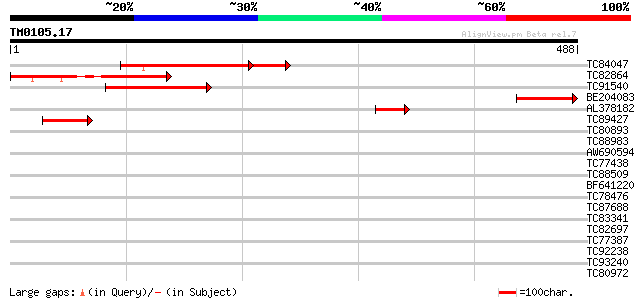
Sequences producing significant alignments: (bits) Value
TC84047 homologue to GP|3695063|gb|AAC62626.1| rac GTPase activa... 170 2e-51
TC82864 similar to GP|3695061|gb|AAC62625.1| rac GTPase activati... 153 1e-37
TC91540 similar to GP|9757821|dbj|BAB08339.1 rac GTPase activati... 100 2e-21
BE204083 similar to GP|3695061|gb|A rac GTPase activating protei... 92 4e-19
AL378182 similar to GP|3695063|gb|A rac GTPase activating protei... 50 1e-06
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 42 5e-04
TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {... 41 0.001
TC88983 similar to SP|P93231|VP41_LYCES Vacuolar assembly protei... 40 0.002
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 40 0.002
TC77438 homologue to PIR|T06807|T06807 nucleosome assembly prote... 40 0.003
TC88509 similar to PIR|T01826|T01826 microfibril-associated prot... 39 0.003
BF641220 weakly similar to PIR|G86203|G86 probable N-arginine di... 38 0.010
TC78476 homologue to SP|O22437|CHLD_PEA Magnesium-chelatase subu... 37 0.013
TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~un... 37 0.022
TC83341 weakly similar to GP|8978076|dbj|BAA98104.1 gene_id:K14A... 37 0.022
TC82697 similar to GP|23497724|gb|AAN37262.1 hypothetical protei... 36 0.029
TC77387 similar to GP|15983396|gb|AAL11566.1 At1g51440/F5D21_19 ... 36 0.029
TC92238 weakly similar to GP|14456138|emb|CAC41653. hypothetical... 36 0.038
TC93240 weakly similar to GP|14335170|gb|AAK59865.1 AT5g51070/K3... 36 0.038
TC80972 weakly similar to GP|23496972|gb|AAN36523.1 hypothetical... 35 0.050
>TC84047 homologue to GP|3695063|gb|AAC62626.1| rac GTPase activating
protein 3 {Lotus japonicus}, partial (34%)
Length = 630
Score = 170 bits (430), Expect(2) = 2e-51
Identities = 85/121 (70%), Positives = 99/121 (81%), Gaps = 6/121 (4%)
Frame = +3
Query: 96 ALKKSLVTCSVDR-EDVSS-----LDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEVPQ 149
ALKKS+V CSV+ +DV S ++I WPT V+HV+HVTFDRFNGFLGLP EL+ VP
Sbjct: 156 ALKKSMVACSVESPDDVISAVHHPMEIGWPTNVKHVNHVTFDRFNGFLGLPLELEVHVPA 335
Query: 150 KVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQE 209
VPSAS VFGVSA+SMQCSYD +GNSVPTILL+MQ RLYS+GGLKAEGIFRIN +N +E
Sbjct: 336 PVPSASVSVFGVSAESMQCSYDSKGNSVPTILLLMQERLYSQGGLKAEGIFRINPENGEE 515
Query: 210 E 210
E
Sbjct: 516 E 518
Score = 51.2 bits (121), Expect(2) = 2e-51
Identities = 21/35 (60%), Positives = 28/35 (80%)
Frame = +1
Query: 207 SQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFREL 241
++ +R QLN G+VP ++VHCL+GLIKAWFREL
Sbjct: 508 AKRNILREQLNSGIVPNDIDVHCLAGLIKAWFREL 612
>TC82864 similar to GP|3695061|gb|AAC62625.1| rac GTPase activating protein
2 {Lotus japonicus}, partial (14%)
Length = 583
Score = 153 bits (387), Expect = 1e-37
Identities = 86/145 (59%), Positives = 105/145 (72%), Gaps = 6/145 (4%)
Frame = +2
Query: 1 MTRLFRSKSCGLVEFNPS---PPSPFFHNVKDEEDDEDDEEEEEEE---EELYNDDEDDY 54
MTRLFRSKSC + F+ S P + +N K EE E +EE+EEEE ++ NDD+D+
Sbjct: 182 MTRLFRSKSCNIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 361
Query: 55 WSNPVSTPLINPAGSKDKGNWRGQSHNQNQNQFAILDILVAALKKSLVTCSVDREDVSSL 114
+S+ N + KG S+N +QNQFAILDI++AALKKS+VTCSV+REDVSSL
Sbjct: 362 YSSN------NASRQFTKG-----SNNNHQNQFAILDIVMAALKKSIVTCSVEREDVSSL 508
Query: 115 DISWPTEVRHVSHVTFDRFNGFLGL 139
DISWPTEVRHVSHVTFDRFNGFLGL
Sbjct: 509 DISWPTEVRHVSHVTFDRFNGFLGL 583
>TC91540 similar to GP|9757821|dbj|BAB08339.1 rac GTPase activating protein
{Arabidopsis thaliana}, partial (13%)
Length = 645
Score = 99.8 bits (247), Expect = 2e-21
Identities = 50/92 (54%), Positives = 66/92 (71%), Gaps = 1/92 (1%)
Frame = +1
Query: 83 NQNQFAILDILVAALKKSLVTC-SVDREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPT 141
+Q F + ++LV +KSL S +D+ ++DIS PT VRHV+HVTFDRFNGFLGLP
Sbjct: 370 SQPHFPLFELLVTLFRKSLFPFKSSGNKDLCNMDISPPTNVRHVAHVTFDRFNGFLGLPD 549
Query: 142 ELQPEVPQKVPSASAKVFGVSAKSMQCSYDER 173
E +P+ P++ PSASA VF VS KS+Q SYD +
Sbjct: 550 EFEPDFPRRPPSASATVFEVSTKSIQFSYDSK 645
>BE204083 similar to GP|3695061|gb|A rac GTPase activating protein 2 {Lotus
japonicus}, partial (12%)
Length = 468
Score = 92.0 bits (227), Expect = 4e-19
Identities = 42/52 (80%), Positives = 45/52 (85%)
Frame = -2
Query: 437 YRGIYDSEHWLRLRKGVRRLCQHPVFQLSKSTKKRADLGIVNTREGGGEAWA 488
YR YDSEHW RLR GVR+LC+HPVFQLSK +KK A LGIVNTREGGGEAWA
Sbjct: 455 YRRRYDSEHWSRLRNGVRKLCRHPVFQLSKPSKKPASLGIVNTREGGGEAWA 300
>AL378182 similar to GP|3695063|gb|A rac GTPase activating protein 3 {Lotus
japonicus}, partial (12%)
Length = 218
Score = 50.4 bits (119), Expect = 1e-06
Identities = 23/29 (79%), Positives = 28/29 (96%)
Frame = +3
Query: 316 PLTALIHAVQVMNFLKTLILKTLRERDES 344
P+TAL+HAVQVMN LKTLI+KTLRER+E+
Sbjct: 3 PVTALMHAVQVMNLLKTLIMKTLREREET 89
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 42.0 bits (97), Expect = 5e-04
Identities = 17/43 (39%), Positives = 29/43 (66%)
Frame = +1
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKD 71
DE+DD+D +E+++ +E ++ DEDD ++P P+ N AG D
Sbjct: 322 DEDDDDDVNDEDDDNDEDFSGDEDDEDADPEDDPVPNGAGGSD 450
Score = 34.7 bits (78), Expect = 0.085
Identities = 16/47 (34%), Positives = 28/47 (59%)
Frame = +1
Query: 14 EFNPSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVS 60
E +P P + DE+DD+DD++ ++ E+E +++EDD P S
Sbjct: 412 EDDPVPNGAGGSDDDDEDDDDDDDDNDDGEDEDEDEEEDDDEDQPPS 552
>TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {Arabidopsis
thaliana}, partial (39%)
Length = 883
Score = 41.2 bits (95), Expect = 0.001
Identities = 20/52 (38%), Positives = 31/52 (59%), Gaps = 4/52 (7%)
Frame = +3
Query: 26 NVKDEEDDEDD----EEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKG 73
+ +D+EDD+DD +E+++ EEE Y+ DE + +P P N AG D G
Sbjct: 318 DTEDDEDDDDDDDVQDEDDDGEEEDYSGDEGEEEGDPEDDPEANGAGGSDDG 473
Score = 33.9 bits (76), Expect = 0.14
Identities = 13/30 (43%), Positives = 22/30 (73%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNP 58
D++ DE+DEE+ E+EE+ +++EDD P
Sbjct: 483 DDDGDEEDEEDGEDEEDEEDEEEDDETPQP 572
Score = 32.3 bits (72), Expect = 0.42
Identities = 13/25 (52%), Positives = 19/25 (76%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
D EDD+DD +EE+EE+ +DE+D
Sbjct: 468 DGEDDDDDGDEEDEEDGEDEEDEED 542
Score = 32.0 bits (71), Expect = 0.55
Identities = 12/18 (66%), Positives = 17/18 (93%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEE 45
+DEED ED+E+EE+EEE+
Sbjct: 501 EDEEDGEDEEDEEDEEED 554
Score = 31.6 bits (70), Expect = 0.72
Identities = 12/24 (50%), Positives = 19/24 (79%)
Frame = +3
Query: 30 EEDDEDDEEEEEEEEELYNDDEDD 53
E+DD+D +EE+EE+ E D+ED+
Sbjct: 474 EDDDDDGDEEDEEDGEDEEDEEDE 545
Score = 28.5 bits (62), Expect = 6.1
Identities = 10/23 (43%), Positives = 18/23 (77%)
Frame = +3
Query: 31 EDDEDDEEEEEEEEELYNDDEDD 53
+D EDD+++ +EE+E +DE+D
Sbjct: 465 DDGEDDDDDGDEEDEEDGEDEED 533
>TC88983 similar to SP|P93231|VP41_LYCES Vacuolar assembly protein VPS41
homolog. [Tomato] {Lycopersicon esculentum}, partial
(17%)
Length = 668
Score = 40.4 bits (93), Expect = 0.002
Identities = 16/41 (39%), Positives = 26/41 (63%)
Frame = +2
Query: 13 VEFNPSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
+ P PP + E+DE+DEEEE+E+EE+ DD+++
Sbjct: 104 IRMAPIPPENGVDGDDEREEDEEDEEEEDEDEEVEEDDDEE 226
Score = 35.4 bits (80), Expect = 0.050
Identities = 14/24 (58%), Positives = 21/24 (87%)
Frame = +2
Query: 30 EEDDEDDEEEEEEEEELYNDDEDD 53
EED+ED+EEE+E+EE +DDE++
Sbjct: 158 EEDEEDEEEEDEDEEVEEDDDEEE 229
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 40.4 bits (93), Expect = 0.002
Identities = 17/25 (68%), Positives = 22/25 (88%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDED 52
+DE DDE++EEEEEEEEE +DDE+
Sbjct: 162 EDEHDDEEEEEEEEEEEEEEDDDEE 236
Score = 39.7 bits (91), Expect = 0.003
Identities = 16/26 (61%), Positives = 22/26 (84%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EED+ DDEEEEEEEEE +++DD
Sbjct: 153 EEEEDEHDDEEEEEEEEEEEEEEDDD 230
Score = 39.3 bits (90), Expect = 0.003
Identities = 17/25 (68%), Positives = 20/25 (80%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
+EE+DE DEEEEEEEEE D+ DD
Sbjct: 105 EEEEDEHDEEEEEEEEEEEEDEHDD 179
Score = 38.9 bits (89), Expect = 0.004
Identities = 16/26 (61%), Positives = 22/26 (84%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE+DE D+EEEEEEEE ++EDD
Sbjct: 150 EEEEEDEHDDEEEEEEEEEEEEEEDD 227
Score = 38.5 bits (88), Expect = 0.006
Identities = 14/26 (53%), Positives = 24/26 (91%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+DE D+E++EEEEEEEE+ ++D+E++
Sbjct: 114 EDEHDEEEEEEEEEEEEDEHDDEEEE 191
Score = 37.4 bits (85), Expect = 0.013
Identities = 15/26 (57%), Positives = 23/26 (87%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+++E DE++EEEEEEEEE +DDE++
Sbjct: 111 EEDEHDEEEEEEEEEEEEDEHDDEEE 188
Score = 35.8 bits (81), Expect = 0.038
Identities = 14/26 (53%), Positives = 22/26 (83%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EED+ D+EEEEEEEEE ++ +D+
Sbjct: 105 EEEEDEHDEEEEEEEEEEEEDEHDDE 182
Score = 35.0 bits (79), Expect = 0.065
Identities = 13/26 (50%), Positives = 22/26 (84%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+++E D+++EEEEEEEEE DD+++
Sbjct: 159 EEDEHDDEEEEEEEEEEEEEEDDDEE 236
Score = 35.0 bits (79), Expect = 0.065
Identities = 13/26 (50%), Positives = 22/26 (84%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++E++ +D+EEEEEEEEE +D+D+
Sbjct: 156 EEEDEHDDEEEEEEEEEEEEEEDDDE 233
Score = 34.7 bits (78), Expect = 0.085
Identities = 13/29 (44%), Positives = 24/29 (81%)
Frame = +3
Query: 25 HNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
H+ ++EE++E++EEEEE+++E +DE D
Sbjct: 171 HDDEEEEEEEEEEEEEEDDDEEGEEDEID 257
Score = 34.7 bits (78), Expect = 0.085
Identities = 14/25 (56%), Positives = 20/25 (80%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DEE++++ +EEEEEEEE +DE D
Sbjct: 102 DEEEEDEHDEEEEEEEEEEEEDEHD 176
Score = 33.5 bits (75), Expect = 0.19
Identities = 13/25 (52%), Positives = 21/25 (84%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
+ +D+E++EEEEEEEEE +D+E +
Sbjct: 168 EHDDEEEEEEEEEEEEEEDDDEEGE 242
Score = 33.1 bits (74), Expect = 0.25
Identities = 15/34 (44%), Positives = 25/34 (73%), Gaps = 5/34 (14%)
Frame = +3
Query: 25 HNVKDEEDDEDDEE-----EEEEEEELYNDDEDD 53
H+ ++EE++E++EE EEEEEEE ++E+D
Sbjct: 123 HDEEEEEEEEEEEEDEHDDEEEEEEEEEEEEEED 224
Score = 30.8 bits (68), Expect = 1.2
Identities = 11/25 (44%), Positives = 21/25 (84%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
+E ++E+DE E EEEE+ ++++E+D
Sbjct: 45 EEHNEEEDEGEVEEEEDEHDEEEED 119
Score = 30.4 bits (67), Expect = 1.6
Identities = 11/29 (37%), Positives = 21/29 (71%)
Frame = +3
Query: 25 HNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
HN +++E + ++EE+E +EEE DE++
Sbjct: 51 HNEEEDEGEVEEEEDEHDEEEEDEHDEEE 137
Score = 29.6 bits (65), Expect = 2.7
Identities = 11/36 (30%), Positives = 26/36 (71%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLI 64
+EE++E++EEEEEE+++ ++++ + + PL+
Sbjct: 180 EEEEEEEEEEEEEEDDDEEGEEDEIDRISTLPDPLL 287
Score = 29.6 bits (65), Expect = 2.7
Identities = 11/28 (39%), Positives = 21/28 (74%)
Frame = +3
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDEDDY 54
V++ ++ED+ E EEEE+E ++ED++
Sbjct: 42 VEEHNEEEDEGEVEEEEDEHDEEEEDEH 125
Score = 28.1 bits (61), Expect = 7.9
Identities = 12/26 (46%), Positives = 20/26 (76%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
K EE +E+++E E EEEE +D+E++
Sbjct: 39 KVEEHNEEEDEGEVEEEEDEHDEEEE 116
>TC77438 homologue to PIR|T06807|T06807 nucleosome assembly protein 1 - garden
pea, partial (93%)
Length = 1607
Score = 39.7 bits (91), Expect = 0.003
Identities = 15/30 (50%), Positives = 25/30 (83%)
Frame = +2
Query: 24 FHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
F ++ DE+DDEDD+ E+++E+E ++DEDD
Sbjct: 1025 FGDLDDEDDDEDDDAEDDDEDEDEDEDEDD 1114
Score = 36.2 bits (82), Expect = 0.029
Identities = 13/25 (52%), Positives = 22/25 (88%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE+DD +D++E+E+E+E +DDED+
Sbjct: 1052 DEDDDAEDDDEDEDEDEDEDDDEDE 1126
Score = 29.3 bits (64), Expect = 3.6
Identities = 9/17 (52%), Positives = 17/17 (99%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEE 45
DE++DED++E+++E+EE
Sbjct: 1079 DEDEDEDEDEDDDEDEE 1129
Score = 28.9 bits (63), Expect = 4.7
Identities = 9/17 (52%), Positives = 17/17 (99%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEE 44
+DE++DED++++E+EEE
Sbjct: 1082 EDEDEDEDEDDDEDEEE 1132
>TC88509 similar to PIR|T01826|T01826 microfibril-associated protein homolog
T15F16.8 - Arabidopsis thaliana, partial (83%)
Length = 1306
Score = 39.3 bits (90), Expect = 0.003
Identities = 15/44 (34%), Positives = 28/44 (63%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKD 71
++EE++E++EEEEEEE + D +++Y + P+ P +D
Sbjct: 499 QEEEEEEEEEEEEEEESDYETDSDEEYTGVAMVKPVFVPKSERD 630
Score = 32.7 bits (73), Expect = 0.32
Identities = 14/27 (51%), Positives = 19/27 (69%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDY 54
+D+E+ EEEEEEEEE ++E DY
Sbjct: 475 RDQEEALPQEEEEEEEEEEEEEEESDY 555
>BF641220 weakly similar to PIR|G86203|G86 probable N-arginine dibasic
convertase [imported] - Arabidopsis thaliana, partial
(5%)
Length = 634
Score = 37.7 bits (86), Expect = 0.010
Identities = 16/46 (34%), Positives = 27/46 (57%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKGN 74
DEE+D++DEE++EE+++ DDED+ + + G K N
Sbjct: 368 DEEEDDEDEEDDEEDDDEGEDDEDEEXEDEDEXXVXGREGGKGAAN 505
Score = 35.4 bits (80), Expect = 0.050
Identities = 13/26 (50%), Positives = 22/26 (84%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+DEEDDE+D++E E++E+ +DED+
Sbjct: 386 EDEEDDEEDDDEGEDDEDEEXEDEDE 463
Score = 35.0 bits (79), Expect = 0.065
Identities = 13/25 (52%), Positives = 22/25 (88%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE+D+E+D+E+EE++EE ++ EDD
Sbjct: 359 DEDDEEEDDEDEEDDEEDDDEGEDD 433
Score = 34.7 bits (78), Expect = 0.085
Identities = 16/26 (61%), Positives = 21/26 (80%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
KD DEDDEEE++E+EE +D+EDD
Sbjct: 344 KDGSIDEDDEEEDDEDEE--DDEEDD 415
>TC78476 homologue to SP|O22437|CHLD_PEA Magnesium-chelatase subunit chlD
chloroplast precursor (Mg- protoporphyrin IX chelatase),
partial (68%)
Length = 1596
Score = 37.4 bits (85), Expect = 0.013
Identities = 17/37 (45%), Positives = 25/37 (66%)
Frame = +2
Query: 17 PSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
P PP P +E+++E+D+EEEEEEE DD+D+
Sbjct: 503 PPPPPPQNQESNEEQNEEEDKEEEEEEE----DDKDE 601
Score = 30.0 bits (66), Expect = 2.1
Identities = 10/37 (27%), Positives = 26/37 (70%)
Frame = +2
Query: 17 PSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
P P + + ++EE+D+++EEEEE++++ +++ +
Sbjct: 512 PPPQNQESNEEQNEEEDKEEEEEEEDDKDEEKEEQQE 622
Score = 29.3 bits (64), Expect = 3.6
Identities = 10/19 (52%), Positives = 18/19 (94%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEEL 46
++EE+D+ DEE+EE++E+L
Sbjct: 572 EEEEEDDKDEEKEEQQEQL 628
>TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~unknown
protein {Arabidopsis thaliana}, partial (46%)
Length = 2077
Score = 36.6 bits (83), Expect = 0.022
Identities = 18/44 (40%), Positives = 27/44 (60%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDK 72
D+EDD+DD+EE+EE E++ N +D++ N GSK K
Sbjct: 716 DDEDDDDDDEEDEEAEDMGNVRYEDFFGGK------NEKGSKRK 829
Score = 31.2 bits (69), Expect = 0.94
Identities = 18/56 (32%), Positives = 26/56 (46%), Gaps = 15/56 (26%)
Frame = +2
Query: 13 VEFNPSPPSPFFHNVKDEEDD---------------EDDEEEEEEEEELYNDDEDD 53
VE F + D++DD EDD EEE++E+E ++DEDD
Sbjct: 365 VELEEEESDDFDEELDDDDDDFEGVEKKKAKGGSEGEDDFEEEDDEDEEGSEDEDD 532
Score = 30.0 bits (66), Expect = 2.1
Identities = 12/25 (48%), Positives = 19/25 (76%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE+ +EDDE E+ E E+ ++D+DD
Sbjct: 662 DEDSEEDDELEKAGEFEMDDEDDDD 736
Score = 29.6 bits (65), Expect = 2.7
Identities = 13/26 (50%), Positives = 20/26 (76%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+D+ ++EDDE+EE E+E DDE+D
Sbjct: 473 EDDFEEEDDEDEEGSEDE---DDEED 541
Score = 28.5 bits (62), Expect = 6.1
Identities = 10/16 (62%), Positives = 15/16 (93%)
Frame = +2
Query: 30 EEDDEDDEEEEEEEEE 45
E DDEDD++++EE+EE
Sbjct: 710 EMDDEDDDDDDEEDEE 757
>TC83341 weakly similar to GP|8978076|dbj|BAA98104.1 gene_id:K14A3.4~unknown
protein {Arabidopsis thaliana}, partial (80%)
Length = 1253
Score = 36.6 bits (83), Expect = 0.022
Identities = 23/72 (31%), Positives = 35/72 (47%), Gaps = 2/72 (2%)
Frame = +1
Query: 19 PPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPL--INPAGSKDKGNWR 76
P S + N +E++E++EEEEEEEEE +D E D + P ++ G+ D R
Sbjct: 550 PRSEWIGNEGLQEEEEEEEEEEEEEEEDDDDVEKDETVEQIHVPQPDVSDNGTSDPARMR 729
Query: 77 GQSHNQNQNQFA 88
N + A
Sbjct: 730 HDVTNNGTSDAA 765
>TC82697 similar to GP|23497724|gb|AAN37262.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (0%)
Length = 614
Score = 36.2 bits (82), Expect = 0.029
Identities = 16/26 (61%), Positives = 22/26 (84%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
KDE+D+EDDEE++EE+EE D+E D
Sbjct: 156 KDEDDEEDDEEDDEEDEE--EDEEVD 227
Score = 29.6 bits (65), Expect = 2.7
Identities = 12/23 (52%), Positives = 19/23 (82%)
Frame = +3
Query: 31 EDDEDDEEEEEEEEELYNDDEDD 53
+ DEDDEE++EE++E D+E+D
Sbjct: 153 DKDEDDEEDDEEDDE--EDEEED 215
Score = 28.9 bits (63), Expect = 4.7
Identities = 9/23 (39%), Positives = 20/23 (86%)
Frame = +3
Query: 30 EEDDEDDEEEEEEEEELYNDDED 52
++D++D+E++EE++EE +DE+
Sbjct: 153 DKDEDDEEDDEEDDEEDEEEDEE 221
>TC77387 similar to GP|15983396|gb|AAL11566.1 At1g51440/F5D21_19
{Arabidopsis thaliana}, partial (75%)
Length = 1793
Score = 36.2 bits (82), Expect = 0.029
Identities = 20/53 (37%), Positives = 31/53 (57%)
Frame = +3
Query: 25 HNVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKGNWRG 77
H ++EE++ED++EEEEEEEE ++E++ P+S G D W G
Sbjct: 321 HEEEEEEEEEDEDEEEEEEEE---EEEEESPLPPLSEVWREIQGEND---WEG 461
>TC92238 weakly similar to GP|14456138|emb|CAC41653. hypothetical protein
{Ustilago maydis}, partial (5%)
Length = 641
Score = 35.8 bits (81), Expect = 0.038
Identities = 18/43 (41%), Positives = 26/43 (59%), Gaps = 1/43 (2%)
Frame = -3
Query: 17 PSPPSPFFHNVKDEEDDE-DDEEEEEEEEELYNDDEDDYWSNP 58
P+ S F N KD+ED+E D+ EE+ +EE + + DD NP
Sbjct: 495 PNDESIFLQNGKDDEDEEYYDDHEEDSDEEFFEEYFDDNELNP 367
>TC93240 weakly similar to GP|14335170|gb|AAK59865.1 AT5g51070/K3K7_27
{Arabidopsis thaliana}, partial (8%)
Length = 700
Score = 35.8 bits (81), Expect = 0.038
Identities = 15/44 (34%), Positives = 26/44 (59%)
Frame = -3
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSK 70
VK+EE++E++EEEEEEEE D+ ++ + ++ K
Sbjct: 254 VKEEEEEEEEEEEEEEEERFL*DERNEREGGEYTVAVVEDGSGK 123
Score = 30.4 bits (67), Expect = 1.6
Identities = 12/22 (54%), Positives = 19/22 (85%)
Frame = -3
Query: 24 FHNVKDEEDDEDDEEEEEEEEE 45
F +V + +++E++EEEEEEEEE
Sbjct: 272 FVDVSEVKEEEEEEEEEEEEEE 207
>TC80972 weakly similar to GP|23496972|gb|AAN36523.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (9%)
Length = 662
Score = 35.4 bits (80), Expect = 0.050
Identities = 21/45 (46%), Positives = 27/45 (59%)
Frame = -2
Query: 3 RLFRSKSCGLVEFNPSPPSPFFHNVKDEEDDEDDEEEEEEEEELY 47
R+ RSK +E + + + KDEE EDDE EEEE+EELY
Sbjct: 661 RMNRSKKSWWIEEDDDDDADDDNEEKDEE--EDDEVEEEEQEELY 533
Score = 30.0 bits (66), Expect = 2.1
Identities = 8/27 (29%), Positives = 23/27 (84%)
Frame = -2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYW 55
D+ DD+++E++EEE++E+ +++++ +
Sbjct: 613 DDADDDNEEKDEEEDDEVEEEEQEELY 533
Score = 30.0 bits (66), Expect = 2.1
Identities = 11/22 (50%), Positives = 17/22 (77%)
Frame = -2
Query: 30 EEDDEDDEEEEEEEEELYNDDE 51
EEDD+DD +++ EE++ DDE
Sbjct: 628 EEDDDDDADDDNEEKDEEEDDE 563
Score = 29.6 bits (65), Expect = 2.7
Identities = 12/38 (31%), Positives = 24/38 (62%)
Frame = -2
Query: 16 NPSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
N S S + D++ D+D+EE++EEE++ ++E +
Sbjct: 655 NRSKKSWWIEEDDDDDADDDNEEKDEEEDDEVEEEEQE 542
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,342,188
Number of Sequences: 36976
Number of extensions: 221758
Number of successful extensions: 4020
Number of sequences better than 10.0: 173
Number of HSP's better than 10.0 without gapping: 1717
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2772
length of query: 488
length of database: 9,014,727
effective HSP length: 100
effective length of query: 388
effective length of database: 5,317,127
effective search space: 2063045276
effective search space used: 2063045276
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0105.17


