

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0102b.4
(278 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
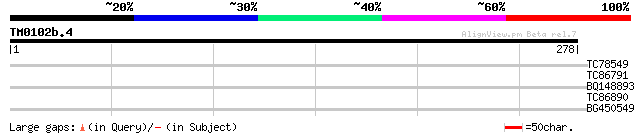
TC78549 similar to GP|15027268|emb|CAC44902. replication licensi... 31 0.46
TC86791 similar to GP|9828617|gb|AAG00240.1| F1N21.15 {Arabidops... 28 3.9
BQ148893 similar to GP|19578315|emb transcription factor E2Fe {A... 27 6.6
TC86890 weakly similar to GP|4406801|gb|AAD20110.1| unknown prot... 27 6.6
BG450549 similar to GP|19578315|emb transcription factor E2Fe {A... 27 6.6
>TC78549 similar to GP|15027268|emb|CAC44902. replication licensing factor
MCM7 homologue {Zea mays}, partial (73%)
Length = 1900
Score = 31.2 bits (69), Expect = 0.46
Identities = 11/23 (47%), Positives = 14/23 (60%)
Frame = +3
Query: 197 NCCCREESSVEDDVFSGWSLNGV 219
NCCC E S + D F GWS+ +
Sbjct: 1251 NCCCSERSGDK*DGFRGWSIGAI 1319
>TC86791 similar to GP|9828617|gb|AAG00240.1| F1N21.15 {Arabidopsis thaliana},
partial (70%)
Length = 1157
Score = 28.1 bits (61), Expect = 3.9
Identities = 15/54 (27%), Positives = 30/54 (54%)
Frame = +2
Query: 159 LLVRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPNCCCREESSVEDDVFS 212
+++ L F+ A ILI + +++ +I++ + +CCCRE S V ++S
Sbjct: 953 VMILLLVLVFVN*ALIILIMASDFSLFLIKDEWW**YKHCCCRE*SVVPLQIYS 1114
>BQ148893 similar to GP|19578315|emb transcription factor E2Fe {Arabidopsis
thaliana}, partial (17%)
Length = 545
Score = 27.3 bits (59), Expect = 6.6
Identities = 15/40 (37%), Positives = 19/40 (47%)
Frame = +2
Query: 224 ELSESGGEDDDGLWPVETKSQHDLQGSPNHSLGDNKNPRS 263
++ S ED+D L T SQ + P S DN NP S
Sbjct: 53 DVKVSDEEDEDELLSQTTGSQGESLSQPTGSQNDNLNPNS 172
>TC86890 weakly similar to GP|4406801|gb|AAD20110.1| unknown protein
{Arabidopsis thaliana}, partial (54%)
Length = 2446
Score = 27.3 bits (59), Expect = 6.6
Identities = 17/67 (25%), Positives = 27/67 (39%)
Frame = +3
Query: 212 SGWSLNGVTVTPELSESGGEDDDGLWPVETKSQHDLQGSPNHSLGDNKNPRSNTGAVNRL 271
S + +NG E E G ++GL + + +HS D+ N N V +
Sbjct: 1887 SDFLINGYNEDLEYVEESGNSEEGLGMMHMARGSTITNGDSHSHSDHANGNGNGHVVINV 2066
Query: 272 LSSRLNS 278
S+ NS
Sbjct: 2067 DSNSTNS 2087
>BG450549 similar to GP|19578315|emb transcription factor E2Fe {Arabidopsis
thaliana}, partial (25%)
Length = 679
Score = 27.3 bits (59), Expect = 6.6
Identities = 15/40 (37%), Positives = 19/40 (47%)
Frame = +2
Query: 224 ELSESGGEDDDGLWPVETKSQHDLQGSPNHSLGDNKNPRS 263
++ S ED+D L T SQ + P S DN NP S
Sbjct: 113 DVKVSDEEDEDELLSQTTGSQGESLSQPTGSQNDNLNPNS 232
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,753,151
Number of Sequences: 36976
Number of extensions: 145180
Number of successful extensions: 781
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 779
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 781
length of query: 278
length of database: 9,014,727
effective HSP length: 95
effective length of query: 183
effective length of database: 5,502,007
effective search space: 1006867281
effective search space used: 1006867281
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0102b.4


