

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.3
(244 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
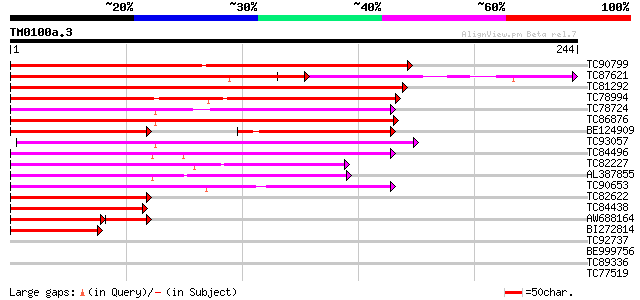
Score E
Sequences producing significant alignments: (bits) Value
TC90799 similar to GP|7672991|gb|AAF66690.1| MADS-box transcript... 144 2e-35
TC87621 similar to SP|Q9FVC1|SVP_ARATH SHORT VEGETATIVE PHASE pr... 125 2e-29
TC81292 homologue to PIR|T09700|T09700 MADS-box protein - alfalf... 122 1e-28
TC78994 similar to SP|O64645|SOC1_ARATH SUPPRESSOR OF CONSTANS O... 117 5e-27
TC78724 similar to GP|4322475|gb|AAD16052.1| putative MADS box t... 114 4e-26
TC86876 similar to GP|16973296|emb|CAC80857. C-type MADS box pro... 112 1e-25
BE124909 similar to GP|4322475|gb| putative MADS box transcripti... 84 8e-25
TC93057 homologue to GP|23194453|gb|AAN15183.1 MADS box protein ... 107 3e-24
TC84496 similar to GP|1483232|emb|CAA67969.1 MADS5 protein {Betu... 103 8e-23
TC82227 similar to GP|1483232|emb|CAA67969.1 MADS5 protein {Betu... 97 7e-21
AL387855 similar to GP|1483230|emb MADS4 protein {Betula pendula... 94 4e-20
TC90653 similar to GP|10835358|gb|AAC13695.2 PTD protein {Populu... 87 8e-18
TC82622 similar to GP|21739238|emb|CAD40993. OSJNBa0072F16.17 {O... 84 4e-17
TC84438 homologue to GP|1483232|emb|CAA67969.1 MADS5 protein {Be... 83 8e-17
AW688164 similar to GP|3114588|gb|A MADS box protein {Eucalyptus... 51 1e-09
BI272814 PIR|S71757|S71 MADS box protein DEFH200 - garden snapdr... 54 4e-08
TC92737 weakly similar to GP|15022157|gb|AAK77938.1 MADS box pro... 38 0.003
BE999756 weakly similar to GP|9758603|dbj| gene_id:MDF20.13~unkn... 37 0.009
TC89336 weakly similar to GP|21554135|gb|AAM63215.1 unknown {Ara... 30 1.1
TC77519 similar to PIR|H86398|H86398 protein F17L21.10 [imported... 29 1.4
>TC90799 similar to GP|7672991|gb|AAF66690.1| MADS-box transcription factor
{Canavalia lineata}, partial (76%)
Length = 653
Score = 144 bits (364), Expect = 2e-35
Identities = 78/173 (45%), Positives = 113/173 (65%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDN ++RQVTFSKRR+G+FKKA+ELS LCDA++ L++FS T KL+EYASS
Sbjct: 71 MARQKIKIKKIDNATARQVTFSKRRRGIFKKAEELSILCDAEVGLVIFSTTGKLYEYASS 250
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
+ ++ S K P +Q E + L K+V ++T +LR + ED +GL L
Sbjct: 251 NMKDIITRYGQQSHHITKLDKPL-QVQVEKNMPAELNKEVADRTQQLRGMKSEDFEGLNL 427
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSL 173
LQ+LE+ L+ L V +K++K + EI L+ KE+ L EN+ LKQ + L
Sbjct: 428 EGLQQLEKSLESXLKRVIEMKEKKILNEIKALRMKEIMLEXENKHLKQKMAML 586
>TC87621 similar to SP|Q9FVC1|SVP_ARATH SHORT VEGETATIVE PHASE protein.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (65%)
Length = 1322
Score = 125 bits (313), Expect = 2e-29
Identities = 67/130 (51%), Positives = 91/130 (69%), Gaps = 1/130 (0%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQIKKI+N ++RQVTFSKRR+GL KKA+ELS LCDAD+ALI+FS+T KLFEY++
Sbjct: 437 MAREKIQIKKIENSTARQVTFSKRRRGLIKKAEELSVLCDADVALIIFSSTGKLFEYSNL 616
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSN-DTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L +S K PS +Q +SN L K+V +K+H+LRQ+ GEDLQG
Sbjct: 617 SMREILERHHLHSKNLAKLEEPSLELQLVENSNCSRLSKEVAQKSHQLRQMRGEDLQGFE 796
Query: 120 LHQLQKLEEV 129
++ + EV
Sbjct: 797 SGRVTTIGEV 826
Score = 62.4 bits (150), Expect = 2e-10
Identities = 43/130 (33%), Positives = 72/130 (55%), Gaps = 1/130 (0%)
Frame = +3
Query: 116 QGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFK 175
+GL+L +LQ+LE+ L+ L V K EK M EI+ L+ K +L+EEN +LK+ + +F
Sbjct: 786 KGLSLEELQQLEKSLEIGLGRVIETKGEKIMMEINELQTKGRQLMEENNRLKRHVSGMFN 965
Query: 176 LLFNEYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQR-QSLESTISSSSYLLEEDGS 234
+GG++S++++ T+ GQ +S+ + +S+ + + S
Sbjct: 966 GKM----------FGGVESENMV-----------TEEGQSSESVTNVYNSTGPPQDYESS 1082
Query: 235 DTSLKLGLVY 244
DTSLKLGL Y
Sbjct: 1083DTSLKLGLPY 1112
>TC81292 homologue to PIR|T09700|T09700 MADS-box protein - alfalfa
(fragment), partial (91%)
Length = 800
Score = 122 bits (306), Expect = 1e-28
Identities = 68/171 (39%), Positives = 107/171 (61%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +IQI++IDN +SRQVTFSKRR GL KKA+EL+ LCDA++ +++FS+T KL+++AS+
Sbjct: 71 MGRGKIQIRRIDNSTSRQVTFSKRRNGLLKKAKELAILCDAEVGVMIFSSTAKLYDFAST 250
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S ++ + ++ + ++F LR+++ RQ+ GE+L GLT+
Sbjct: 251 SMRSVIDRYNKTKEEHNQLGSSTSEIKFWQREAAMLRQQLHNLQESHRQIMGEELSGLTV 430
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLY 171
+LQ LE L+ SL V K++ FM EI L RK + +EN +L + +Y
Sbjct: 431 KELQGLENQLEISLRGVRMKKEQLFMDEIQELNRKGDIIHQENVELYRKVY 583
>TC78994 similar to SP|O64645|SOC1_ARATH SUPPRESSOR OF CONSTANS
OVEREXPRESSION 1 protein (Agamous-like MADS box protein
AGL20)., partial (83%)
Length = 946
Score = 117 bits (292), Expect = 5e-27
Identities = 71/173 (41%), Positives = 106/173 (61%), Gaps = 5/173 (2%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R + Q+K+I+N +SRQVTFSKRR GL KKA ELS LCDA++ALIVFS +L+E+ASS
Sbjct: 54 MVRGKTQMKRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALIVFSPRGRLYEFASS 233
Query: 61 SRGGLLPLPDANSAT*DKTTPPSP-----TMQFESDSNDTLRKKVEEKTHELRQLNGEDL 115
S L + S T TP + T Q + ++ + + KK++ R+L GE L
Sbjct: 234 SI--LETIERYRSHTRINNTPTTSESVENTQQLKEEA-ENMMKKIDLLETSKRKLLGEGL 404
Query: 116 QGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
++ +LQK+E+ L++S+ + K + F ++I LK KE L+ EN +L +
Sbjct: 405 GSCSIDELQKIEQQLEKSINKIRVKKTKVFREQIDQLKEKEKALVAENVRLSE 563
>TC78724 similar to GP|4322475|gb|AAD16052.1| putative MADS box
transcription factor ETL {Eucalyptus globulus subsp.
globulus}, partial (71%)
Length = 1089
Score = 114 bits (284), Expect = 4e-26
Identities = 71/174 (40%), Positives = 103/174 (58%), Gaps = 8/174 (4%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R + Q+K+I+N S+RQVTFSKRR GL KKA ELS LCDA++ALIVFS T KL+E++SS
Sbjct: 226 MVRGKTQMKRIENESNRQVTFSKRRNGLLKKAFELSVLCDAEVALIVFSTTGKLYEFSSS 405
Query: 61 S--------RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNG 112
S +G + L + + T Q + + KK+E R+L G
Sbjct: 406 SISKTVERYQGKVKELVLSTKGIQENT-------QHLKECDIDTTKKLEHLELSKRKLLG 564
Query: 113 EDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL 166
E+L +LQ++E L+RSL+ + K++ ++I LK KE L+EEN++L
Sbjct: 565 EELGSCAFDELQQIENQLERSLSKIRARKNQLLKEQIEKLKDKERLLLEENKRL 726
>TC86876 similar to GP|16973296|emb|CAC80857. C-type MADS box protein {Malus
x domestica}, partial (85%)
Length = 2050
Score = 112 bits (281), Expect = 1e-25
Identities = 63/168 (37%), Positives = 102/168 (60%), Gaps = 1/168 (0%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++AL+VFS +L+EYA++
Sbjct: 89 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVVFSTRGRLYEYANN 268
Query: 61 S-RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S R + A +A+ + + QF + LR+++ + + R + GE L L+
Sbjct: 269 SVRATIERYKKACAASTNAESVSEANTQFYQQESSKLRRQIRDIQNLNRHILGEALGSLS 448
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
L +L+ LE L++ L+ V K E ++ ++++E+EL N L+
Sbjct: 449 LKELKNLEGRLEKGLSRVRSRKHETLFADVEFMQKREIELQNHNNYLR 592
>BE124909 similar to GP|4322475|gb| putative MADS box transcription factor
ETL {Eucalyptus globulus subsp. globulus}, partial (60%)
Length = 608
Score = 84.0 bits (206), Expect(2) = 8e-25
Identities = 41/61 (67%), Positives = 51/61 (83%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R + Q+K+I+N S+RQVTFSKRR GL KKA ELS LCDA++ALIVFS T KL+E++SS
Sbjct: 148 MVRGKTQMKRIENESNRQVTFSKRRNGLLKKAFELSVLCDAEVALIVFSTTGKLYEFSSS 327
Query: 61 S 61
S
Sbjct: 328 S 330
Score = 46.6 bits (109), Expect(2) = 8e-25
Identities = 26/68 (38%), Positives = 42/68 (61%)
Frame = +2
Query: 99 KVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVE 158
K +K H R+L GE+L +LQ++E L+RSL+ + K++ ++I LK KE
Sbjct: 389 KESKKIH--RKLLGEELGSCAFDELQQIENQLERSLSKIRARKNQLLKEQIEKLKDKERL 562
Query: 159 LIEENQKL 166
L+EEN++L
Sbjct: 563 LLEENKRL 586
>TC93057 homologue to GP|23194453|gb|AAN15183.1 MADS box protein GHMADS-2
{Gossypium hirsutum}, partial (95%)
Length = 858
Score = 107 bits (268), Expect = 3e-24
Identities = 63/174 (36%), Positives = 103/174 (58%), Gaps = 1/174 (0%)
Frame = +3
Query: 4 KRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSS-R 62
+R++ K I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS+ +L+EY++++ R
Sbjct: 39 ERLR*KGIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNNNIR 218
Query: 63 GGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLHQ 122
+ A S TT Q+ + LR++++ + R L G+ L LT+ +
Sbjct: 219 STIDRYKKACSDHSSTTTTTEINAQYYQQESAKLRQQIQMLQNSNRHLMGDALSTLTVKE 398
Query: 123 LQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKL 176
L++LE L+R + + K E + EI +++E+EL EN L+ + + +L
Sbjct: 399 LKQLENRLERGITRIRSKKHEMLLAEIEYFQKREIELENENLCLRTKINDVERL 560
>TC84496 similar to GP|1483232|emb|CAA67969.1 MADS5 protein {Betula
pendula}, partial (74%)
Length = 681
Score = 103 bits (256), Expect = 8e-23
Identities = 66/168 (39%), Positives = 95/168 (56%), Gaps = 2/168 (1%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+Q+KKI+N +RQVTFSKRR GL KKA E+S LCDA++ALIVFS KL+EY+S
Sbjct: 26 MGRGRVQLKKIENKINRQVTFSKRRSGLLKKAHEISVLCDAEVALIVFSNKGKLYEYSSD 205
Query: 61 -SRGGLLPLPDANS-AT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGL 118
+L + S A P + + L+ ++E R GE+L GL
Sbjct: 206 PCMEKILERYERYSYAERQHVANDQPQNENWIIEHARLKTRLEVIQKNQRNFMGEELDGL 385
Query: 119 TLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL 166
++ +LQ LE L +L + K++ + IS L +K+ L E+N+ L
Sbjct: 386 SMKELQHLEHQLDSALKQIRSRKNQLMYESISELSKKDKALQEKNKLL 529
>TC82227 similar to GP|1483232|emb|CAA67969.1 MADS5 protein {Betula
pendula}, partial (68%)
Length = 681
Score = 96.7 bits (239), Expect = 7e-21
Identities = 58/149 (38%), Positives = 88/149 (58%), Gaps = 3/149 (2%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+Q+K+I+N +RQVTFSKRR GL KKAQE+S LCDA++ALI+FS KLFEY+S
Sbjct: 130 MGRGRVQLKRIENKINRQVTFSKRRSGLLKKAQEISVLCDAEVALIIFSTKGKLFEYSSD 309
Query: 61 SRGGLLPLPDANSAT*DK---TTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQG 117
+ + ++ T+ SP + + + L+ ++E R GEDL G
Sbjct: 310 PCMEKILERYERCSYMERQLVTSEQSPNENWVLE-HAKLKARMEVLQRNQRNFMGEDLDG 486
Query: 118 LTLHQLQKLEEVLKRSLASVSRVKDEKFM 146
L L +LQ LE+ L +L ++ ++ ++
Sbjct: 487 LGLKELQSLEQQLDSALKQITITEEPSYV 573
>AL387855 similar to GP|1483230|emb MADS4 protein {Betula pendula}, partial
(51%)
Length = 493
Score = 94.4 bits (233), Expect = 4e-20
Identities = 61/148 (41%), Positives = 85/148 (57%), Gaps = 1/148 (0%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+Q+K+I+N +S+QVTF KRR GL KKA E+S LCDA +ALI+FS KLFEY+S+
Sbjct: 53 MGRGRVQLKRIENKTSQQVTFFKRRTGLLKKANEISVLCDAQVALIMFSTKGKLFEYSSA 232
Query: 61 -SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L + + T + T + T S L KV+ LR G DL L+
Sbjct: 233 PSMEDILERYERQNHT-ELTGATNETQGNWSFEYMKLTAKVQVLERNLRNFVGNDLDPLS 409
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQ 147
+ +LQ LE+ L SL + K++ Q
Sbjct: 410 VKELQSLEQQLDTSLKRIRTRKNQVMNQ 493
>TC90653 similar to GP|10835358|gb|AAC13695.2 PTD protein {Populus
balsamifera subsp. trichocarpa}, partial (59%)
Length = 845
Score = 86.7 bits (213), Expect = 8e-18
Identities = 58/172 (33%), Positives = 86/172 (49%), Gaps = 6/172 (3%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK I+N ++RQVT+SKRR G+FKKA ELS LCDA ++LI+FS NK+ EY +
Sbjct: 44 MGRGKIEIKLIENPTNRQVTYSKRRNGIFKKAHELSVLCDAKVSLIMFSKNNKMHEYITP 223
Query: 61 SRGGLLPLPDANSAT*DKTTPPS------PTMQFESDSNDTLRKKVEEKTHELRQLNGED 114
+ D S ++ D N LR+++ + E G +
Sbjct: 224 GLSTKKIIDQYQKTLGDIDLWRSHYEKMLENLKKLKDINHKLRRQIRHRIGE----GGME 391
Query: 115 LQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL 166
L L+ QL+ LEE + S+A + K T ++K L + N L
Sbjct: 392 LDDLSFQQLRSLEEDMNSSIAKIRERKFHVIKTRTDTCRKKVRSLEQMNGNL 547
>TC82622 similar to GP|21739238|emb|CAD40993. OSJNBa0072F16.17 {Oryza
sativa}, partial (78%)
Length = 847
Score = 84.3 bits (207), Expect = 4e-17
Identities = 39/61 (63%), Positives = 52/61 (84%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I I++IDN +SRQVTFSKRRKGL KKA+EL+ LCDA + L++FS+T KL+EYA++
Sbjct: 138 MGRGKIVIRRIDNCTSRQVTFSKRRKGLIKKAKELAILCDAQVGLVIFSSTGKLYEYANT 317
Query: 61 S 61
S
Sbjct: 318 S 320
>TC84438 homologue to GP|1483232|emb|CAA67969.1 MADS5 protein {Betula
pendula}, partial (24%)
Length = 702
Score = 83.2 bits (204), Expect = 8e-17
Identities = 39/59 (66%), Positives = 49/59 (82%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYAS 59
M R R+Q+K+I+N +RQVTFSKRR GL KKAQE+S LCDA++ALI+FS KLFEY+S
Sbjct: 223 MGRGRVQLKRIENKINRQVTFSKRRSGLLKKAQEISVLCDAEVALIIFSTKGKLFEYSS 399
>AW688164 similar to GP|3114588|gb|A MADS box protein {Eucalyptus grandis},
partial (31%)
Length = 549
Score = 51.2 bits (121), Expect(2) = 1e-09
Identities = 25/41 (60%), Positives = 31/41 (74%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDA 41
M R R+ +++I+N +RQVTFSKRR GL KKA EL LCDA
Sbjct: 7 MGRGRVVLERIENKINRQVTFSKRRSGLLKKAFELCVLCDA 129
Score = 28.1 bits (61), Expect(2) = 1e-09
Identities = 10/20 (50%), Positives = 18/20 (90%)
Frame = +2
Query: 42 DIALIVFSATNKLFEYASSS 61
++ALI+FS+ KLF+Y+S++
Sbjct: 131 EVALIIFSSRGKLFQYSSTT 190
>BI272814 PIR|S71757|S71 MADS box protein DEFH200 - garden snapdragon,
partial (16%)
Length = 320
Score = 54.3 bits (129), Expect = 4e-08
Identities = 25/40 (62%), Positives = 32/40 (79%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCD 40
M R R+++K+I+N +RQVTF+KRR GL KKA ELS LCD
Sbjct: 199 MGRGRVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCD 318
>TC92737 weakly similar to GP|15022157|gb|AAK77938.1 MADS box protein-like
protein NGL9 {Medicago sativa}, partial (33%)
Length = 541
Score = 38.1 bits (87), Expect = 0.003
Identities = 25/77 (32%), Positives = 39/77 (50%)
Frame = +3
Query: 94 DTLRKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLK 153
D ++K+ + +LR L GED+ L +L LEE L+ L V +K M+ K
Sbjct: 21 DRIKKENDSMQIDLRHLKGEDITSLNYKELMALEESLENGLTGVR----DKKMEVHRMFK 188
Query: 154 RKEVELIEENQKLKQVL 170
R L +EN++L +L
Sbjct: 189 RNGKILEDENKELNFLL 239
>BE999756 weakly similar to GP|9758603|dbj| gene_id:MDF20.13~unknown
protein {Arabidopsis thaliana}, partial (15%)
Length = 534
Score = 36.6 bits (83), Expect = 0.009
Identities = 17/48 (35%), Positives = 28/48 (57%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVF 48
M ++++ I N ++R+ T+ KR + L KK E++TLC IVF
Sbjct: 13 MAGNKVKLAFITNHTARRATYKKRVQSLMKKLNEITTLCGVKACGIVF 156
>TC89336 weakly similar to GP|21554135|gb|AAM63215.1 unknown {Arabidopsis
thaliana}, partial (7%)
Length = 1241
Score = 29.6 bits (65), Expect = 1.1
Identities = 24/88 (27%), Positives = 39/88 (44%)
Frame = +1
Query: 86 MQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKF 145
++ +D LRK++E E RQL +D + + KLE L + ++D
Sbjct: 736 LKISNDEVARLRKELESNLAETRQL--QDQLEVAQENVTKLEWQLDSGRKQIRELED--- 900
Query: 146 MQEISTLKRKEVELIEENQKLKQVLYSL 173
I+ K E L E QKLK ++ +
Sbjct: 901 --RITWFKTNETNLEVEVQKLKDEMHDV 978
>TC77519 similar to PIR|H86398|H86398 protein F17L21.10 [imported] -
Arabidopsis thaliana, partial (98%)
Length = 1037
Score = 29.3 bits (64), Expect = 1.4
Identities = 14/32 (43%), Positives = 22/32 (68%)
Frame = +2
Query: 156 EVELIEENQKLKQVLYSLFKLLFNEYLCSIIL 187
E +LI++ LKQVLY +F + +Y CS++L
Sbjct: 884 EDDLIKK*ANLKQVLY*MFITVLVKYYCSVML 979
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.136 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,846,708
Number of Sequences: 36976
Number of extensions: 67636
Number of successful extensions: 382
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 377
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 377
length of query: 244
length of database: 9,014,727
effective HSP length: 94
effective length of query: 150
effective length of database: 5,538,983
effective search space: 830847450
effective search space used: 830847450
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0100a.3


