

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.12
(405 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
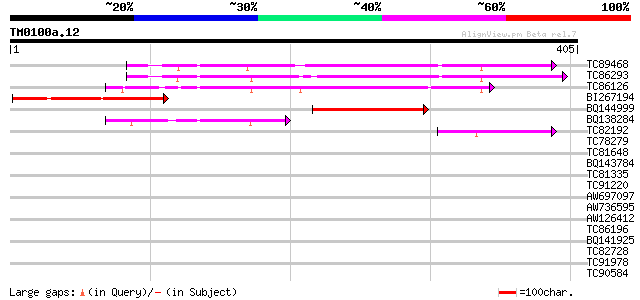
TC89468 similar to GP|5305145|emb|CAB46084.1 fructose-1 6-bispho... 231 3e-61
TC86293 homologue to GP|5305145|emb|CAB46084.1 fructose-1 6-bisp... 227 5e-60
TC86126 homologue to GP|14318171|gb|AAK59929.1 fructose-1 6-bisp... 206 1e-53
BI267194 similar to GP|17063187|gb| fructose-bisphosphatase-like... 112 2e-25
BQ144999 homologue to GP|14318171|gb| fructose-1 6-bisphosphatas... 79 4e-15
BQ138284 similar to PIR|JC7375|JC7 fructose-bisphosphatase (EC 3... 71 9e-13
TC82192 similar to GP|15451178|gb|AAK96860.1 sedoheptulose-bisph... 43 2e-04
TC78279 similar to SP|O20252|S17P_SPIOL Sedoheptulose-1 7-bispho... 41 0.001
TC81648 similar to PIR|T00840|T00840 probable senescence-related... 33 0.15
BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antig... 30 0.21
TC81335 33 0.26
TC91220 31 0.75
AW697097 similar to PIR|T48399|T483 heat shock transcription fac... 31 0.75
AW736595 similar to PIR|T48399|T483 heat shock transcription fac... 31 0.75
AW126412 similar to GP|11994418|db gb|AAF04863.1~gene_id:MBK21.1... 30 2.2
TC86196 similar to GP|22652125|gb|AAN03626.1 BEL1-related homeot... 30 2.2
BQ141925 homologue to GP|10727150|gb| Rm62 gene product {alt 1} ... 29 2.9
TC82728 similar to GP|7292941|gb|AAF48332.1| CG11071-PA {Drosoph... 29 2.9
TC91978 similar to GP|16508407|emb|CAD10175. unnamed protein pro... 29 3.7
TC90584 similar to GP|22136810|gb|AAM91749.1 unknown protein {Ar... 28 4.9
>TC89468 similar to GP|5305145|emb|CAB46084.1 fructose-1 6-bisphosphatase
{Pisum sativum}, partial (94%)
Length = 1161
Score = 231 bits (590), Expect = 3e-61
Identities = 133/320 (41%), Positives = 190/320 (58%), Gaps = 13/320 (4%)
Frame = +1
Query: 84 KDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGV----GAGGGGGSSGRDAPKPLDIV 139
+ D +L++HI CK F CS + G+ G G G + K LD++
Sbjct: 37 RGDFTILINHIVLGCK---------FVCSSVNKAGLAKILGLAGDTNVQGEEQKK-LDVI 186
Query: 140 SNEIILSSLKKSGKVAVMASEENDAPTWI--IDDGPYVVVTDPLDGSRNIDASIPTGTIF 197
SNE+ + +L SG+ ++ SEE + + G Y+VV DPLDGS NID + GTIF
Sbjct: 187 SNEVFVKALISSGRTCLLVSEEVEDAIIVPPAQRGKYIVVFDPLDGSSNIDCGVSIGTIF 366
Query: 198 GIYNRLEELDNLPTEEKAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHST 257
GIY + T E + ++LQ GS L+AAGY +Y S+ +S G+G FTLD S
Sbjct: 367 GIYTVKD------TAEVSVEDALQPGSSLVAAGYCMYGSSCTFVLSTGNGVNGFTLDPSL 528
Query: 258 GDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVR-QGKGRYPKKYSARYICSL 316
G+FILT+P+IKIP +G+IYSVN+ +W E +Y++ + +G PK S RYI S+
Sbjct: 529 GEFILTHPNIKIPKKGKIYSVNEGNAKNWDEPTAKYVENCKFPQEGSAPK--SLRYIGSM 702
Query: 317 VADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPV 370
VAD+HRTL+YGG+ M P D LR++YE P+S+L+EQAGG+ GK R L L P
Sbjct: 703 VADVHRTLLYGGIFMYPADKKSPNGKLRVLYEVFPMSYLMEQAGGQAFTGKQRALDLVPK 882
Query: 371 KLHQRLPLFLGSLEDMEELE 390
K+H R P+FLGS +++E ++
Sbjct: 883 KIHDRSPIFLGSYDEIEHIK 942
>TC86293 homologue to GP|5305145|emb|CAB46084.1 fructose-1 6-bisphosphatase
{Pisum sativum}, complete
Length = 1377
Score = 227 bits (579), Expect = 5e-60
Identities = 134/327 (40%), Positives = 187/327 (56%), Gaps = 12/327 (3%)
Frame = +1
Query: 84 KDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTG----VGAGGGGGSSGRDAPKPLDIV 139
+ D +LL++I CK F CS + G +G G G + K LD++
Sbjct: 256 RGDFTILLNNIVLGCK---------FVCSAVNKAGLAKLIGLAGETNVQGEEQKK-LDVL 405
Query: 140 SNEIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIF 197
SNE+ +L SG+ ++ SEE++ ++ G Y VV DPLDGS NID + GTIF
Sbjct: 406 SNEVFCKALISSGRTCILVSEEDEDAIFVEPKQRGKYCVVFDPLDGSSNIDCGVSIGTIF 585
Query: 198 GIYNRLEELDNLPTEEKAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHST 257
GIY + + P E A LQ G ++AAGY +Y S+ ++ GSG FTLD S
Sbjct: 586 GIYMMKDNHE--PVIEDA----LQPGKNMLAAGYCMYGSSCTFVITTGSGVNGFTLDPSL 747
Query: 258 GDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLV 317
G+FILT+P IKIP +G+IYSVN+ +W Y++ + P K S RYI S+V
Sbjct: 748 GEFILTHPDIKIPKKGKIYSVNEGNAKNWDGPTAAYVEKCKFPTDGSPAK-SLRYIGSMV 924
Query: 318 ADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVK 371
AD+HRTL+YGG + P D LR++YE P+SFL+EQAGG+ GK R L L P K
Sbjct: 925 ADVHRTLLYGGTFLYPADKKSPNGKLRVLYEVFPMSFLMEQAGGQAFTGKERALDLIPTK 1104
Query: 372 LHQRLPLFLGSLEDMEELESFGDVQQK 398
LH+R P+FLGS +D+EE+++ + K
Sbjct: 1105LHERSPIFLGSYDDIEEIKALYAAEAK 1185
>TC86126 homologue to GP|14318171|gb|AAK59929.1 fructose-1 6-bisphosphatase
{Pisum sativum}, partial (88%)
Length = 1205
Score = 206 bits (524), Expect = 1e-53
Identities = 127/300 (42%), Positives = 180/300 (59%), Gaps = 22/300 (7%)
Frame = +1
Query: 69 VVTLIEYLGKE---GIDLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGG 125
++TL +L K+ G+ + +L ++L+ I ACK+IA++V +I TGV G
Sbjct: 328 LITLTSWLLKQEQTGV-IDAELTIVLNSISLACKQIASLVQR---ANISNLTGVQ--GAV 489
Query: 126 GSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDG 183
G D K LD+VSNE+ + L+ SG+ ++ASEE D P + + G Y+VV DPLDG
Sbjct: 490 NIQGEDQKK-LDVVSNEVFSNCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVFDPLDG 666
Query: 184 SRNIDASIPTGTIFGIYNRLEEL----------DNLPTEE-KAQLNSLQSGSKLIAAGYV 232
S NIDA++ TG+IFGIY+ +E L TEE + +N Q GS L+AAGY
Sbjct: 667 SSNIDAAVSTGSIFGIYSPNDECLADVGNESDDPTLGTEEQRCIVNVCQPGSNLLAAGYC 846
Query: 233 LYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQ 292
+YSS+ I ++ G G FTLD G+F+LT +++IP G+IYS N+ Y W + L++
Sbjct: 847 MYSSSVIFVLTIGKGVFVFTLDPMYGEFVLTQENLQIPKSGKIYSFNEGNYKLWDDNLKK 1026
Query: 293 YIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPL 346
YID +++ G K YSARYI SLV D HRTL+YGG+ PRD LRL+YE P+
Sbjct: 1027YIDDLKE-PGANGKPYSARYIGSLVGDFHRTLLYGGIYGYPRDKKSKNGKLRLLYECAPM 1203
>BI267194 similar to GP|17063187|gb| fructose-bisphosphatase-like protein
{Arabidopsis thaliana}, partial (6%)
Length = 427
Score = 112 bits (281), Expect = 2e-25
Identities = 64/114 (56%), Positives = 84/114 (73%), Gaps = 3/114 (2%)
Frame = +1
Query: 3 SAAASPFYQLV-PFHPKFQTFQSKTPLTTPYCTPRLASISG-SAKMRPLRALGGSSSSAG 60
+ +PFYQL+ F PK QTFQSKT T + TP +AS+S +++PL ALG SSSS+
Sbjct: 61 ATTTTPFYQLLLTFRPKLQTFQSKTQ--TLFYTPTMASVSEFRTRVKPLSALGNSSSSSS 234
Query: 61 GDGDDGDGVVTLIEYLGKEG-IDLKDDLVVLLDHIQYACKRIAAIVASPFSCSI 113
+D DG VTL+EY+GK G I + DDLV+L+DHIQYACKRIAA++ASPF+ +I
Sbjct: 235 TVAND-DGSVTLMEYMGKRGGIGVNDDLVILIDHIQYACKRIAALIASPFNYTI 393
>BQ144999 homologue to GP|14318171|gb| fructose-1 6-bisphosphatase {Pisum
sativum}, partial (22%)
Length = 625
Score = 78.6 bits (192), Expect = 4e-15
Identities = 35/83 (42%), Positives = 54/83 (64%)
Frame = +2
Query: 217 LNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIY 276
+N Q GS L+AAGY +YSS+ I ++ G G FTL G+F+LT +++IP G+IY
Sbjct: 353 VNVCQPGSNLLAAGYCMYSSSVIFVLTIGKGVFVFTLXPMYGEFVLTQENLQIPKSGKIY 532
Query: 277 SVNDARYFDWPEGLRQYIDTVRQ 299
S N+ Y W + L++YID +++
Sbjct: 533 SFNEGNYKLWDDNLKKYIDDLKE 601
>BQ138284 similar to PIR|JC7375|JC7 fructose-bisphosphatase (EC 3.1.3.11) -
Aspergillus oryzae, partial (39%)
Length = 646
Score = 70.9 bits (172), Expect = 9e-13
Identities = 43/137 (31%), Positives = 72/137 (52%), Gaps = 5/137 (3%)
Frame = +1
Query: 69 VVTLIEYLGKEGIDLKD---DLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGG 125
++TL +L +E K+ D +L +Q+A K IA + ++ G G
Sbjct: 241 IITLTRFLTEEQSKHKEATGDFTLLCHALQFAFKSIAYYIRRATLVNL-----TGLAGQS 405
Query: 126 GSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIID--DGPYVVVTDPLDG 183
++G D K LDI+ +E+ +S+++ G+V ++ SEE + + D Y V DP+DG
Sbjct: 406 NATGDDQKK-LDIIGDELFISAMRSCGRVRLLVSEEQEELIMFDEFPDARYAVACDPIDG 582
Query: 184 SRNIDASIPTGTIFGIY 200
S N+DA + GTIF +Y
Sbjct: 583 SSNLDAGVSVGTIFAVY 633
>TC82192 similar to GP|15451178|gb|AAK96860.1 sedoheptulose-bisphosphatase
precursor {Arabidopsis thaliana}, partial (40%)
Length = 664
Score = 42.7 bits (99), Expect = 2e-04
Identities = 27/91 (29%), Positives = 45/91 (48%), Gaps = 6/91 (6%)
Frame = +1
Query: 306 KKYSARYICSLVADLHRTLMYG-GVAMN-----PRDHLRLVYEANPLSFLVEQAGGRGSD 359
+KY+ RY +V D+++ ++ G+ N + LRL++E PL L+E AGG SD
Sbjct: 175 EKYTLRYTGGMVPDVNQIIVKEKGIFTNVTSPTAKAKLRLLFEVAPLGLLIENAGGYSSD 354
Query: 360 GKNRILSLQPVKLHQRLPLFLGSLEDMEELE 390
G +L + R + GS ++ E
Sbjct: 355 GHKSVLDKVIENIDDRTQVAYGSKNEVIRFE 447
>TC78279 similar to SP|O20252|S17P_SPIOL Sedoheptulose-1 7-bisphosphatase
chloroplast precursor (EC 3.1.3.37)
(Sedoheptulose-bisphosphatase), partial (49%)
Length = 867
Score = 40.8 bits (94), Expect = 0.001
Identities = 25/76 (32%), Positives = 37/76 (47%), Gaps = 6/76 (7%)
Frame = +2
Query: 131 DAPKPLDIVSNEIILSSLKKSGKVAVMASEE------NDAPTWIIDDGPYVVVTDPLDGS 184
D +D+++N ++ +LK S SEE P DG + V DPLDGS
Sbjct: 536 DEQLAVDMLANNLLFEALKHSHFCKYACSEEIPELQDMGGPV----DGGFSVAFDPLDGS 703
Query: 185 RNIDASIPTGTIFGIY 200
+D + GTIFG++
Sbjct: 704 SIVDTNFTVGTIFGVW 751
>TC81648 similar to PIR|T00840|T00840 probable senescence-related protein
[imported] - Arabidopsis thaliana, partial (36%)
Length = 854
Score = 33.5 bits (75), Expect = 0.15
Identities = 26/80 (32%), Positives = 37/80 (45%)
Frame = -3
Query: 47 RPLRALGGSSSSAGGDGDDGDGVVTLIEYLGKEGIDLKDDLVVLLDHIQYACKRIAAIVA 106
RPL +G S DGDG+ TL E E K + V+L+D ++ +
Sbjct: 369 RPLDFIGDPSI-------DGDGIFTLAETDDGEVTGSKFNAVLLIDEVENGSGDFDKNLF 211
Query: 107 SPFSCSIGKQTGVGAGGGGG 126
F +++GV AGGGGG
Sbjct: 210 GSFR-RCRRRSGVDAGGGGG 154
>BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antigen. [strain
Kaplan PRV] {Pseudorabies virus}, partial (2%)
Length = 1022
Score = 30.0 bits (66), Expect(2) = 0.21
Identities = 13/17 (76%), Positives = 13/17 (76%)
Frame = -3
Query: 120 GAGGGGGSSGRDAPKPL 136
GAGGGG SGR AP PL
Sbjct: 216 GAGGGGDLSGRPAPPPL 166
Score = 21.6 bits (44), Expect(2) = 0.21
Identities = 14/35 (40%), Positives = 17/35 (48%)
Frame = -2
Query: 47 RPLRALGGSSSSAGGDGDDGDGVVTLIEYLGKEGI 81
R L ALGG ++ G G G V L G EG+
Sbjct: 358 RSLPALGGGAA*WGFPGGGGCPRVRLFLGAGMEGV 254
>TC81335
Length = 686
Score = 32.7 bits (73), Expect = 0.26
Identities = 18/46 (39%), Positives = 23/46 (49%)
Frame = -3
Query: 345 PLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELE 390
P S L+ A DG+ IL +QP H +PLFL L D L+
Sbjct: 195 PDSLLITPASAAAPDGRLSILFVQPPVSHDAMPLFLPLLGDFWALQ 58
>TC91220
Length = 603
Score = 31.2 bits (69), Expect = 0.75
Identities = 16/41 (39%), Positives = 24/41 (58%), Gaps = 2/41 (4%)
Frame = -1
Query: 35 PRLASISGS--AKMRPLRALGGSSSSAGGDGDDGDGVVTLI 73
P + ++ G+ A+M PL L +AGG D GDGV T++
Sbjct: 594 PSVGAVDGAFLARMGPLAPLKSQLGAAGGRADVGDGVGTIL 472
>AW697097 similar to PIR|T48399|T483 heat shock transcription factor-like
protein - Arabidopsis thaliana, partial (14%)
Length = 640
Score = 31.2 bits (69), Expect = 0.75
Identities = 25/79 (31%), Positives = 37/79 (46%)
Frame = +2
Query: 109 FSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWI 168
FS S + A GGGGS D P P++ + + L K+ + V N +W
Sbjct: 191 FSSSSLMEYEASAIGGGGSGSSDVPVPMECLQMSPVPPFLSKTFDL-VDDPSLNPIISWS 367
Query: 169 IDDGPYVVVTDPLDGSRNI 187
+ G + VV DPL+ +R I
Sbjct: 368 SNGGSF-VVWDPLEFARII 421
>AW736595 similar to PIR|T48399|T483 heat shock transcription factor-like
protein - Arabidopsis thaliana, partial (16%)
Length = 465
Score = 31.2 bits (69), Expect = 0.75
Identities = 25/79 (31%), Positives = 37/79 (46%)
Frame = +3
Query: 109 FSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWI 168
FS S + A GGGGS D P P++ + + L K+ + V N +W
Sbjct: 54 FSSSSLMEYEASAIGGGGSGSSDVPVPMECLQMSPVPPFLSKTFDL-VDDPSLNPIISWS 230
Query: 169 IDDGPYVVVTDPLDGSRNI 187
+ G + VV DPL+ +R I
Sbjct: 231 SNGGSF-VVWDPLEFARII 284
>AW126412 similar to GP|11994418|db gb|AAF04863.1~gene_id:MBK21.14~similar to
unknown protein {Arabidopsis thaliana}, partial (60%)
Length = 529
Score = 29.6 bits (65), Expect = 2.2
Identities = 14/46 (30%), Positives = 26/46 (56%)
Frame = -1
Query: 184 SRNIDASIPTGTIFGIYNRLEELDNLPTEEKAQLNSLQSGSKLIAA 229
SRN +S + I+GI++R +++P + K +NS S ++ A
Sbjct: 502 SRNFCSSFSSERIYGIFSRNFSSESIPKDCKPSVNSFLENSHIVPA 365
>TC86196 similar to GP|22652125|gb|AAN03626.1 BEL1-related homeotic protein
29 {Solanum tuberosum}, partial (53%)
Length = 2227
Score = 29.6 bits (65), Expect = 2.2
Identities = 30/122 (24%), Positives = 53/122 (42%), Gaps = 8/122 (6%)
Frame = +1
Query: 16 HPKFQTFQSKTPLTTPYCTPRLAS-ISGSAKMRPLRAL---GGSSSSAGG----DGDDGD 67
H ++ + + P T TPR +S S +P R + GGS S A G + +
Sbjct: 556 HVQYNQWNTIDPNTAARDTPRAQQGLSLSLSSQPARFVSMSGGSPSPASGVTTNNNNGNP 735
Query: 68 GVVTLIEYLGKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGS 127
G+ L + +L +++V + ++ K + +IG+ +GVG+GG G
Sbjct: 736 GIQILSSKYLRAAHELLEEVVNVNSGVELGKKSGGGMS----KVNIGESSGVGSGGDGSI 903
Query: 128 SG 129
G
Sbjct: 904 IG 909
>BQ141925 homologue to GP|10727150|gb| Rm62 gene product {alt 1} [Drosophila
melanogaster], partial (3%)
Length = 1059
Score = 29.3 bits (64), Expect = 2.9
Identities = 12/21 (57%), Positives = 13/21 (61%)
Frame = -3
Query: 113 IGKQTGVGAGGGGGSSGRDAP 133
IGK+ G G GGGGG G P
Sbjct: 67 IGKEKGGGGGGGGGGGGGGEP 5
>TC82728 similar to GP|7292941|gb|AAF48332.1| CG11071-PA {Drosophila
melanogaster}, partial (2%)
Length = 734
Score = 29.3 bits (64), Expect = 2.9
Identities = 19/55 (34%), Positives = 27/55 (48%)
Frame = +1
Query: 41 SGSAKMRPLRALGGSSSSAGGDGDDGDGVVTLIEYLGKEGIDLKDDLVVLLDHIQ 95
SGS+ R L S D D GDG V L + LG E + +++D V L ++
Sbjct: 358 SGSSGSGIRRELSFSRWCDDDDNDGGDGRVILDQKLGDEDVIVEEDAVYELPFVE 522
>TC91978 similar to GP|16508407|emb|CAD10175. unnamed protein product
{synthetic construct}, partial (94%)
Length = 820
Score = 28.9 bits (63), Expect = 3.7
Identities = 25/90 (27%), Positives = 33/90 (35%)
Frame = +1
Query: 53 GGSSSSAGGDGDDGDGVVTLIEYLGKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCS 112
GGS+ G D G T+ K + L+ + + D Y C R P
Sbjct: 250 GGST----GYADSVKGRFTISRDNAKNSLYLQMNSLRAEDTAVYYCARGLKSKYRPLFWG 417
Query: 113 IGKQTGVGAGGGGGSSGRDAPKPLDIVSNE 142
G V +GGGG G LDI N+
Sbjct: 418 QGTLVTVSSGGGGSGGGGSGGSALDIQMNQ 507
>TC90584 similar to GP|22136810|gb|AAM91749.1 unknown protein {Arabidopsis
thaliana}, partial (18%)
Length = 686
Score = 28.5 bits (62), Expect = 4.9
Identities = 22/77 (28%), Positives = 31/77 (39%), Gaps = 5/77 (6%)
Frame = +3
Query: 54 GSSSSAGGDGDDGDGVVTLIEYLGKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCSI 113
G+SS GGD ++ VV + D DD + D + CK +A SP+ +
Sbjct: 213 GASSGGGGDDEENGDVVDSSNRVDSHHDDDHDDRDLFGDDNEDYCKTLA---KSPYPIPV 383
Query: 114 -----GKQTGVGAGGGG 125
G GGGG
Sbjct: 384 LPPIRNNTNNPGRGGGG 434
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.137 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,306,123
Number of Sequences: 36976
Number of extensions: 149833
Number of successful extensions: 1129
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 1051
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1103
length of query: 405
length of database: 9,014,727
effective HSP length: 98
effective length of query: 307
effective length of database: 5,391,079
effective search space: 1655061253
effective search space used: 1655061253
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0100a.12


