

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.11
(245 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
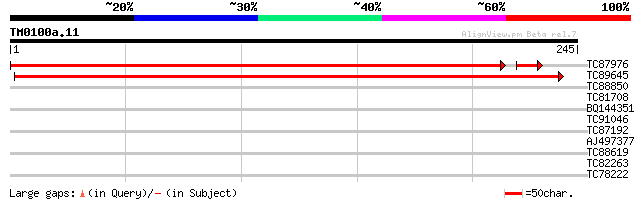
Score E
Sequences producing significant alignments: (bits) Value
TC87976 similar to GP|15144508|gb|AAK84475.1 unknown {Lycopersic... 400 e-113
TC89645 weakly similar to GP|21537016|gb|AAM61357.1 unknown {Ara... 317 3e-87
TC88850 similar to GP|10177090|dbj|BAB10396. soluble starch synt... 29 1.9
TC81708 similar to GP|12083536|gb|AAG48845.1 hypothetical protei... 27 5.5
BQ144351 similar to GP|7242771|emb conserved hypothetical protei... 27 5.5
TC91046 similar to GP|9279622|dbj|BAB01080.1 gene_id:MOJ10.6~unk... 27 5.5
TC87192 weakly similar to PIR|E84526|E84526 probable lysosomal a... 27 7.2
AJ497377 similar to GP|20147355|gb| At1g80210/F18B13_28 {Arabido... 27 7.2
TC88619 homologue to SP|Q43077|AMO_PEA Amine oxidase [copper-con... 27 9.4
TC82263 weakly similar to SP|Q16348|PET2_HUMAN Oligopeptide tran... 27 9.4
TC78222 similar to GP|17064840|gb|AAL32574.1 putative protein ph... 27 9.4
>TC87976 similar to GP|15144508|gb|AAK84475.1 unknown {Lycopersicon
esculentum}, partial (80%)
Length = 948
Score = 400 bits (1029), Expect(2) = e-113
Identities = 191/214 (89%), Positives = 203/214 (94%)
Frame = +2
Query: 1 MVSQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI 60
MVS +ELLKNEIPLEQE VVLAED VNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI
Sbjct: 257 MVSHTVELLKNEIPLEQESVVLAEDIVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI 436
Query: 61 NESARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVT 120
NESARLARLFCEK LP+M FLDSH PNKPE+PYPPHCIAGTDESNLVPALRWLENETNVT
Sbjct: 437 NESARLARLFCEKKLPIMVFLDSHQPNKPEEPYPPHCIAGTDESNLVPALRWLENETNVT 616
Query: 121 LRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGF 180
+RRK+CFDGY+GS EEDGSNVFVDWVKKNKI+T++VVG+CTDICVLDFVCSTMSAKNRGF
Sbjct: 617 IRRKDCFDGYVGSMEEDGSNVFVDWVKKNKIKTMVVVGVCTDICVLDFVCSTMSAKNRGF 796
Query: 181 LKPLENVVVYSHGCATFDVPLEVARNTKGALAHP 214
LKPLENVVVYS+ CATF+VPLEVA N KGALAHP
Sbjct: 797 LKPLENVVVYSNACATFNVPLEVATNIKGALAHP 898
Score = 23.9 bits (50), Expect(2) = e-113
Identities = 9/11 (81%), Positives = 10/11 (90%)
Frame = +1
Query: 220 HMGLYMAKERG 230
H+ LYMAKERG
Sbjct: 916 HVSLYMAKERG 948
>TC89645 weakly similar to GP|21537016|gb|AAM61357.1 unknown {Arabidopsis
thaliana}, partial (83%)
Length = 996
Score = 317 bits (812), Expect = 3e-87
Identities = 143/237 (60%), Positives = 190/237 (79%)
Frame = +2
Query: 3 SQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINE 62
S +EL+K EIP++Q+ ++L+++ GLVLVD++NGFCTVG+GN AP+E + +IS+M+
Sbjct: 80 SPTLELVKEEIPVKQQPLLLSDNFKTGLVLVDLVNGFCTVGSGNFAPKEHDEKISKMVEN 259
Query: 63 SARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLR 122
S +L++ F EKN P+ A+LDSHHP+ PE PYP HC+ G+DES LVP L WLEN+ N TLR
Sbjct: 260 SVKLSKKFAEKNWPIFAYLDSHHPDIPEPPYPSHCLIGSDESKLVPDLLWLENDPNATLR 439
Query: 123 RKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLK 182
+K+C DG+IGS E+DGSNVFVDWVK N+I+ +LV GICTDICVLDF CS +SA+NRGFL
Sbjct: 440 KKDCIDGFIGSYEKDGSNVFVDWVKSNQIKQVLVCGICTDICVLDFTCSVLSARNRGFLS 619
Query: 183 PLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVLF 239
PLENV+V S CAT D+PL VA+ +K ++HPQE +HH+ LY+A RGA+IA+EV F
Sbjct: 620 PLENVIVASQACATCDLPLHVAKASKDLVSHPQELMHHVALYIAVGRGAQIASEVSF 790
>TC88850 similar to GP|10177090|dbj|BAB10396. soluble starch synthase
{Arabidopsis thaliana}, partial (14%)
Length = 752
Score = 28.9 bits (63), Expect = 1.9
Identities = 24/83 (28%), Positives = 30/83 (35%)
Frame = +1
Query: 62 ESARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTL 121
ES L RLF HHP P PH + T+ S +P + N TL
Sbjct: 52 ESLELPRLFL------------HHPRASSAPTRPHPHSTTNSSRFLPQFGFPSKLKNATL 195
Query: 122 RRKECFDGYIGSAEEDGSNVFVD 144
R I + +DG F D
Sbjct: 196 RS-------ISVSSQDGPISFED 243
>TC81708 similar to GP|12083536|gb|AAG48845.1 hypothetical protein {Oryza
sativa}, partial (6%)
Length = 605
Score = 27.3 bits (59), Expect = 5.5
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = -2
Query: 82 DSHHPNKPEDPYPPH 96
DS HP+ P DPY PH
Sbjct: 547 DSLHPSAPLDPYGPH 503
>BQ144351 similar to GP|7242771|emb conserved hypothetical protein SCD8A.23
{Streptomyces coelicolor A3(2)}, partial (2%)
Length = 1019
Score = 27.3 bits (59), Expect = 5.5
Identities = 14/26 (53%), Positives = 16/26 (60%)
Frame = -3
Query: 208 KGALAHPQEFLHHMGLYMAKERGAKI 233
KGA A P +FL GLY K+RG I
Sbjct: 759 KGAWAAPPQFLGGGGLYHPKKRGGGI 682
>TC91046 similar to GP|9279622|dbj|BAB01080.1 gene_id:MOJ10.6~unknown
protein {Arabidopsis thaliana}, partial (6%)
Length = 978
Score = 27.3 bits (59), Expect = 5.5
Identities = 11/40 (27%), Positives = 20/40 (49%)
Frame = +1
Query: 70 FCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPA 109
F + +P ++ ++ P P P+PPH + G + PA
Sbjct: 661 FMDGGIPY-GYVSNNLPPPPPPPFPPHAVTGLSMPSTQPA 777
>TC87192 weakly similar to PIR|E84526|E84526 probable lysosomal acid lipase
[imported] - Arabidopsis thaliana, partial (79%)
Length = 1555
Score = 26.9 bits (58), Expect = 7.2
Identities = 18/55 (32%), Positives = 21/55 (37%), Gaps = 16/55 (29%)
Frame = +1
Query: 83 SHHPNKPEDPYPPHC----------------IAGTDESNLVPALRWLENETNVTL 121
S+H N YP HC I+ T N P LRW +T VTL
Sbjct: 961 SNHQNLTLAAYPDHCLC*WLMGETML*QM*LISSTHSRNCHPHLRWFILKTMVTL 1125
>AJ497377 similar to GP|20147355|gb| At1g80210/F18B13_28 {Arabidopsis
thaliana}, partial (16%)
Length = 667
Score = 26.9 bits (58), Expect = 7.2
Identities = 9/24 (37%), Positives = 13/24 (53%)
Frame = +3
Query: 203 VARNTKGALAHPQEFLHHMGLYMA 226
+ +NT+ HP F+HH Y A
Sbjct: 30 ILQNTRDGKVHPLTFIHHTSTYQA 101
>TC88619 homologue to SP|Q43077|AMO_PEA Amine oxidase [copper-containing]
precursor (EC 1.4.3.6). [Garden pea] {Pisum sativum},
partial (67%)
Length = 1519
Score = 26.6 bits (57), Expect = 9.4
Identities = 15/36 (41%), Positives = 19/36 (52%), Gaps = 4/36 (11%)
Frame = -2
Query: 162 DICVLDFV----CSTMSAKNRGFLKPLENVVVYSHG 193
DIC LDF C + + F P+ N+VV SHG
Sbjct: 681 DICSLDFQYTRQCYSRLDRTTCFELPI*NIVVVSHG 574
>TC82263 weakly similar to SP|Q16348|PET2_HUMAN Oligopeptide transporter
kidney isoform (Peptide transporter 2) (Kidney
H+/peptide cotransporter), partial (3%)
Length = 950
Score = 26.6 bits (57), Expect = 9.4
Identities = 20/76 (26%), Positives = 37/76 (48%)
Frame = +1
Query: 168 FVCSTMSAKNRGFLKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAK 227
FV S + N GF+ + ++V+Y +G FD+P + NT L +G +++
Sbjct: 511 FVLSALD--NMGFVANMVSLVLYFYGVMHFDIP--SSANTLTNFMGSTFLLSLVGGFISD 678
Query: 228 ERGAKIANEVLFGAVE 243
+ +LFG++E
Sbjct: 679 TYLNRFTTCLLFGSLE 726
>TC78222 similar to GP|17064840|gb|AAL32574.1 putative protein phosphatase-2C
(PP2C) {Arabidopsis thaliana}, partial (52%)
Length = 1813
Score = 26.6 bits (57), Expect = 9.4
Identities = 18/58 (31%), Positives = 26/58 (44%), Gaps = 8/58 (13%)
Frame = -1
Query: 81 LDSHHPNKPEDPYPP---HCI--AGTDESNLVPALRWLENETNV---TLRRKECFDGY 130
+ S NKP P+PP H + + T P R + NE N+ T ++ FD Y
Sbjct: 1480 MTSSTKNKPGTPFPPDAKHILNSSNTPPIKCCPRKRKINNEKNIGRRTTKKMSLFDCY 1307
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,211,864
Number of Sequences: 36976
Number of extensions: 120951
Number of successful extensions: 602
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 598
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 601
length of query: 245
length of database: 9,014,727
effective HSP length: 94
effective length of query: 151
effective length of database: 5,538,983
effective search space: 836386433
effective search space used: 836386433
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0100a.11


