

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0096b.3
(223 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
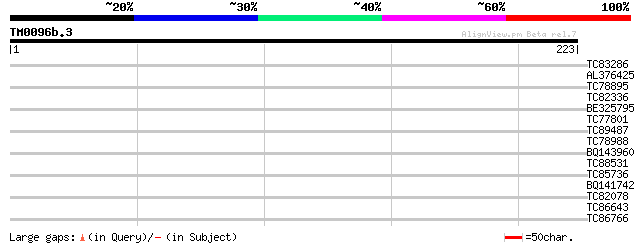
Score E
Sequences producing significant alignments: (bits) Value
TC83286 similar to PIR|B84919|B84919 Not56-like protein [importe... 30 0.96
AL376425 homologue to GP|18377618|gb| unknown protein {Arabidops... 29 1.6
TC78895 similar to GP|6056390|gb|AAF02854.1| Unknown protein {Ar... 29 1.6
TC82336 28 2.1
BE325795 similar to GP|22136848|gb| unknown protein {Arabidopsis... 28 2.8
TC77801 weakly similar to GP|21536807|gb|AAM61139.1 unknown {Ara... 27 4.8
TC89487 similar to GP|13877597|gb|AAK43876.1 RNA lariat debranch... 27 4.8
TC78988 similar to GP|6862912|gb|AAF30301.1| unknown protein {Ar... 27 4.8
BQ143960 weakly similar to SP|P21997|SSGP_ Sulfated surface glyc... 27 4.8
TC88531 homologue to GP|14597044|emb|CAC43563. unnamed protein p... 27 6.2
TC85736 homologue to GP|9230771|gb|AAF85975.1| cytosolic phospho... 27 6.2
BQ141742 weakly similar to PIR|T10798|T107 pherophorin-S - Volvo... 27 6.2
TC82078 similar to GP|12836109|dbj|BAB23506. data source:SPTR s... 27 6.2
TC86643 similar to GP|5922612|dbj|BAA84613.1 EST AU078118(E3904)... 27 8.1
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 27 8.1
>TC83286 similar to PIR|B84919|B84919 Not56-like protein [imported] -
Arabidopsis thaliana, partial (45%)
Length = 845
Score = 29.6 bits (65), Expect = 0.96
Identities = 15/37 (40%), Positives = 21/37 (56%)
Frame = +2
Query: 60 HRMQDVGALVELDPGTPVPEVLALDSSLILPDALKFV 96
HRM V AL L G P P +L ++++L D + FV
Sbjct: 569 HRMCSVVALSSLSMGLPYPTILVVENTLPKIDPIDFV 679
>AL376425 homologue to GP|18377618|gb| unknown protein {Arabidopsis
thaliana}, partial (2%)
Length = 399
Score = 28.9 bits (63), Expect = 1.6
Identities = 24/90 (26%), Positives = 42/90 (46%), Gaps = 2/90 (2%)
Frame = -3
Query: 106 LIIAKQVGVVFDGTDEEAIESYAIHLVANCHGGVYLVFCIGFIIKTFYQPP--PPNDYSC 163
+I+ K V+F +E +E+ +IH N + FC + +T P PP ++
Sbjct: 247 IIVVKYKQVIFKDIVKE-MEAQSIHHQKNLYSHF---FCTFLVFQT*SHPFNFPPFQHTF 80
Query: 164 KEPINPHPIHILEFLGSIIIHLLMCYSLLL 193
++ H +H+ S HLL+ + LLL
Sbjct: 79 PHSLHHHTLHV-----SFHFHLLLLFLLLL 5
>TC78895 similar to GP|6056390|gb|AAF02854.1| Unknown protein {Arabidopsis
thaliana}, partial (28%)
Length = 2089
Score = 28.9 bits (63), Expect = 1.6
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = +3
Query: 140 YLVFCIGFIIKTFYQPPPPNDYSCKEPINPHP 171
+LV + QPP N+ +EP+NPHP
Sbjct: 291 FLVKAVATAATIIEQPPKFNNNHLEEPLNPHP 386
>TC82336
Length = 698
Score = 28.5 bits (62), Expect = 2.1
Identities = 31/103 (30%), Positives = 38/103 (36%), Gaps = 22/103 (21%)
Frame = -3
Query: 122 EAIESYAIHLVANCHGGVYLVF-----CIGFIIKTFYQPPPP---NDYSCKEPI----NP 169
E + SY I L+ +YL CI FI Y PP D C +
Sbjct: 693 EILLSYIIMLILLRRNAIYLRLVKYKKCIYFIQSCHYPYPPS*ASIDQRCGGSLVQSSGT 514
Query: 170 HPIHILEFLGSIIIHLL----------MCYSLLLSNSSHIVMF 202
HPI II+ L+ +SLL SNS HI F
Sbjct: 513 HPICFFSLSKIIILSLIERAPITLSSSKSFSLLSSNSKHIFPF 385
>BE325795 similar to GP|22136848|gb| unknown protein {Arabidopsis thaliana},
partial (6%)
Length = 676
Score = 28.1 bits (61), Expect = 2.8
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = +2
Query: 153 YQPPPPNDYSCKEPINPHPIH 173
+Q PPP + PINP+P H
Sbjct: 614 HQKPPPQTQQNQVPINPNPFH 676
>TC77801 weakly similar to GP|21536807|gb|AAM61139.1 unknown {Arabidopsis
thaliana}, partial (12%)
Length = 1145
Score = 27.3 bits (59), Expect = 4.8
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = -1
Query: 155 PPPPNDYSCKEPINPHPIHILE 176
PPPPN Y ++P P H L+
Sbjct: 623 PPPPNQYHFQQPHQQPPYHQLQ 558
>TC89487 similar to GP|13877597|gb|AAK43876.1 RNA lariat debranching enzyme
- like protein {Arabidopsis thaliana}, partial (31%)
Length = 899
Score = 27.3 bits (59), Expect = 4.8
Identities = 18/44 (40%), Positives = 23/44 (51%)
Frame = +1
Query: 55 SPHIHHRMQDVGALVELDPGTPVPEVLALDSSLILPDALKFVSV 98
S H+H R ALV+ G PV + LALD L D L+ V +
Sbjct: 121 SAHLHCRF---AALVQHGEGGPVTKFLALDKCLPGRDFLQVVEI 243
>TC78988 similar to GP|6862912|gb|AAF30301.1| unknown protein {Arabidopsis
thaliana}, partial (68%)
Length = 2203
Score = 27.3 bits (59), Expect = 4.8
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = -1
Query: 184 HLLMCYSLLLSNSSHIVMFS 203
HLL +S+L +N HI++FS
Sbjct: 1915 HLLFVFSILTNNYKHIILFS 1856
>BQ143960 weakly similar to SP|P21997|SSGP_ Sulfated surface glycoprotein 185
(SSG 185). {Volvox carteri}, partial (13%)
Length = 1023
Score = 27.3 bits (59), Expect = 4.8
Identities = 15/44 (34%), Positives = 20/44 (45%), Gaps = 2/44 (4%)
Frame = +1
Query: 153 YQPPPPNDYSCKEPINPH--PIHILEFLGSIIIHLLMCYSLLLS 194
Y PP P +S P PH P HI SII L+ + ++
Sbjct: 196 YPPPSPPVFSPLSPYTPHIIPQHIKYITTSIIFFLVFTFFFFIN 327
>TC88531 homologue to GP|14597044|emb|CAC43563. unnamed protein product
{Arabidopsis thaliana}, partial (27%)
Length = 898
Score = 26.9 bits (58), Expect = 6.2
Identities = 12/39 (30%), Positives = 22/39 (55%)
Frame = -1
Query: 164 KEPINPHPIHILEFLGSIIIHLLMCYSLLLSNSSHIVMF 202
+EP+ H H+L + L +CYS L++ ++ V+F
Sbjct: 511 QEPLKGHGEHMLVLDTNKPFSLALCYSFALASKANCVLF 395
>TC85736 homologue to GP|9230771|gb|AAF85975.1| cytosolic phosphoglycerate
kinase {Pisum sativum}, complete
Length = 1559
Score = 26.9 bits (58), Expect = 6.2
Identities = 16/49 (32%), Positives = 27/49 (54%)
Frame = +2
Query: 38 LLSGNTNGLSIQTRGMLSPHIHHRMQDVGALVELDPGTPVPEVLALDSS 86
++ G + +++ G L+ + H GA +EL G P+P VLALD +
Sbjct: 1127 IIGGGDSVAAVEKVG-LADKMSHISTGGGASLELLEGKPLPGVLALDDA 1270
>BQ141742 weakly similar to PIR|T10798|T107 pherophorin-S - Volvox carteri,
partial (9%)
Length = 1337
Score = 26.9 bits (58), Expect = 6.2
Identities = 17/45 (37%), Positives = 20/45 (43%)
Frame = +1
Query: 155 PPPPNDYSCKEPINPHPIHILEFLGSIIIHLLMCYSLLLSNSSHI 199
PPPPN KE HP F ++ H L Y LS SS +
Sbjct: 193 PPPPNIRHVKERNKAHPHPYPPFHPTLKTHTLNIYFHPLSYSSFL 327
>TC82078 similar to GP|12836109|dbj|BAB23506. data source:SPTR source
key:Q9NWJ8 evidence:ISS~homolog to CDNA FLJ20798 FIS
CLONE ADSU02031~, partial (3%)
Length = 604
Score = 26.9 bits (58), Expect = 6.2
Identities = 18/74 (24%), Positives = 32/74 (42%)
Frame = +1
Query: 140 YLVFCIGFIIKTFYQPPPPNDYSCKEPINPHPIHILEFLGSIIIHLLMCYSLLLSNSSHI 199
++VFC+ F++ P P+ SC ++ + + F S+ + M L ++S
Sbjct: 283 FVVFCLLFLLFVTTTNPSPSKLSCSS-LSSLELTTVLFFSSVPYIIFMTALSLATSSEEF 459
Query: 200 VMFSKLTSLEKGKS 213
S SL G S
Sbjct: 460 SKLSTFPSLTLGSS 501
>TC86643 similar to GP|5922612|dbj|BAA84613.1 EST AU078118(E3904)
corresponds to a region of the predicted gene.~similar
to Arabidopsis thaliana, partial (39%)
Length = 1705
Score = 26.6 bits (57), Expect = 8.1
Identities = 14/43 (32%), Positives = 23/43 (52%), Gaps = 5/43 (11%)
Frame = +1
Query: 121 EEAIESYAI-----HLVANCHGGVYLVFCIGFIIKTFYQPPPP 158
EEA+ A+ ++++ C G V L + F + T + PPPP
Sbjct: 541 EEAVPCIALSKNDSYVMSACGGKVSLFNMMTFKVMTTFMPPPP 669
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 26.6 bits (57), Expect = 8.1
Identities = 10/19 (52%), Positives = 11/19 (57%)
Frame = +2
Query: 153 YQPPPPNDYSCKEPINPHP 171
Y PPPP+ C EP P P
Sbjct: 260 YSPPPPSPPPCVEPPPPPP 316
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.142 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,834,075
Number of Sequences: 36976
Number of extensions: 128608
Number of successful extensions: 820
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 770
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 820
length of query: 223
length of database: 9,014,727
effective HSP length: 93
effective length of query: 130
effective length of database: 5,575,959
effective search space: 724874670
effective search space used: 724874670
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0096b.3


