

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0095c.6
(384 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
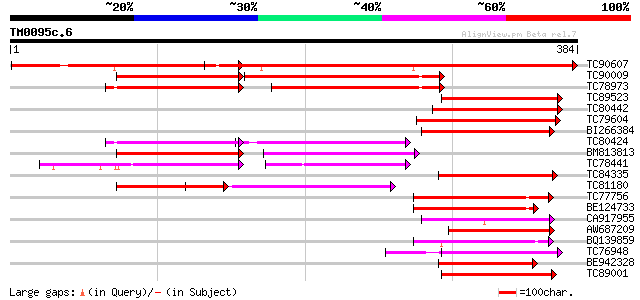
Score E
Sequences producing significant alignments: (bits) Value
TC90607 similar to GP|20466724|gb|AAM20679.1 unknown protein {Ar... 379 e-105
TC90009 homologue to GP|21617909|gb|AAM66959.1 plastid protein {... 104 7e-23
TC78973 similar to PIR|D84745|D84745 plastid protein [imported] ... 94 7e-20
TC89523 weakly similar to GP|17065276|gb|AAL32792.1 Unknown prot... 88 6e-18
TC80442 similar to PIR|F84793|F84793 probable RNA-binding protei... 86 3e-17
TC79604 similar to GP|18175762|gb|AAL59923.1 unknown protein {Ar... 83 2e-16
BI266384 74 9e-14
TC80424 homologue to GP|21593052|gb|AAM65001.1 DAG protein puta... 72 3e-13
BM813813 similar to GP|7549634|gb| DAG protein putative {Arabid... 72 3e-13
TC78441 similar to GP|16226543|gb|AAL16196.1 AT3g15000/K15M2_14 ... 72 5e-13
TC84335 weakly similar to GP|21741678|emb|CAD40580. oj000126_13.... 69 2e-12
TC81180 similar to GP|7549634|gb|AAF63819.1| DAG protein putati... 69 4e-12
TC77756 similar to PIR|T06796|T06796 glycine-rich RNA-binding pr... 68 7e-12
BE124733 similar to PIR|T06796|T06 glycine-rich RNA-binding prot... 65 6e-11
CA917955 65 6e-11
AW687209 weakly similar to GP|9757907|dbj| contains similarity t... 64 1e-10
BQ139859 similar to GP|8777484|dbj contains similarity to RNA-bi... 63 2e-10
TC76948 similar to SP|Q08935|ROC1_NICSY 29 kDa ribonucleoprotein... 61 8e-10
BE942328 weakly similar to GP|21741678|emb oj000126_13.2 {Oryza ... 61 8e-10
TC89001 similar to GP|5726567|gb|AAD48471.1| glycine-rich RNA-bi... 60 1e-09
>TC90607 similar to GP|20466724|gb|AAM20679.1 unknown protein {Arabidopsis
thaliana}, partial (49%)
Length = 1143
Score = 379 bits (973), Expect = e-105
Identities = 194/262 (74%), Positives = 220/262 (83%), Gaps = 10/262 (3%)
Frame = +1
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDYSLSSGQAGLSS-----GLQTRTNMLFPAGNSKHW 187
++Q+ LP + + P+ + +S S SS +S+ + +LFP GNSKHW
Sbjct: 73 TTQIHQLP-IKTKLPNKNNHSSSCSISCSSSFTRISATSNQTSIPQTDMLLFPNGNSKHW 249
Query: 188 LVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDCAQ 247
+VRMDKP V VVTKAQIVD+YAQILTK+MGNEKDAQMCIYHVSWKTNFGFCCELDEDCA
Sbjct: 250 VVRMDKPAVGVVTKAQIVDHYAQILTKIMGNEKDAQMCIYHVSWKTNFGFCCELDEDCAH 429
Query: 248 DLAGVPGVSSVQPDDNFESENKNYE-----DSLSVPNSSEASQEASLRTKKLFVTGLSFY 302
+L+GVPGV SVQPDDNFESENK+YE +SL++PNSSEASQEAS++TKKLFVTGLSFY
Sbjct: 430 ELSGVPGVLSVQPDDNFESENKDYEGRNLENSLNMPNSSEASQEASVKTKKLFVTGLSFY 609
Query: 303 TSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMNGKIINGWM 362
TSEKTLRAAFEGFGELV+VKVIIDKISKRSKGYAFIEYTTEEAA AALKEMNGKIINGWM
Sbjct: 610 TSEKTLRAAFEGFGELVEVKVIIDKISKRSKGYAFIEYTTEEAASAALKEMNGKIINGWM 789
Query: 363 IVVDVAKTNPPRFNRGHARPPA 384
IVVDVAKT PPR+N+GHARP A
Sbjct: 790 IVVDVAKTTPPRYNKGHARPSA 855
Score = 156 bits (394), Expect = 1e-38
Identities = 77/162 (47%), Positives = 109/162 (66%), Gaps = 5/162 (3%)
Frame = +1
Query: 2 FTKCQKISSVNP-NRSQVHQLPTNITLPNKSYHSNLPSSSSSSISCNRFFTIIAAAAATT 60
FTK Q S P N +Q+HQLP LPNK+ HS SS SISC+ FT I+A + T
Sbjct: 31 FTKSQTFPSFRPKNTTQIHQLPIKTKLPNKNNHS-----SSCSISCSSSFTRISATSNQT 195
Query: 61 YPSFTPSTT----HNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDA 116
T +++HW+V MDKP +GV++K+Q++D+Y + L +MG+EK+AQMC+Y
Sbjct: 196 SIPQTDMLLFPNGNSKHWVVRMDKPAVGVVTKAQIVDHYAQILTKIMGNEKDAQMCIYHV 375
Query: 117 SWDTHFGFCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
SW T+FGFCC++DE+ + +L+ +PGVLSV+PD +F S KDY
Sbjct: 376 SWKTNFGFCCELDEDCAHELSGVPGVLSVQPDDNFESENKDY 501
>TC90009 homologue to GP|21617909|gb|AAM66959.1 plastid protein {Arabidopsis
thaliana}, partial (74%)
Length = 1036
Score = 104 bits (259), Expect = 7e-23
Identities = 56/135 (41%), Positives = 82/135 (60%)
Frame = +1
Query: 160 LSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNE 219
L+SG + S T LFP + +HWL+ MDKPG E +K Q++D Y Q L KV+G+E
Sbjct: 235 LNSGNSNFSERPPTDMAPLFPGCDYEHWLIVMDKPGGEGASKQQMIDCYVQTLAKVLGSE 414
Query: 220 KDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSVPN 279
++A+ IY+VS + FGF CE+D++ + L G+PGV V PD + ENK+Y L V
Sbjct: 415 EEAKKKIYNVSCERYFGFGCEIDDETSNKLEGLPGVLFVLPDSYVDPENKDYGAELFV-- 588
Query: 280 SSEASQEASLRTKKL 294
+ E Q + R K++
Sbjct: 589 NGEIVQRSPERQKRV 633
Score = 89.4 bits (220), Expect = 2e-18
Identities = 43/86 (50%), Positives = 60/86 (69%)
Frame = +1
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++MDKP SK Q+ID YV+TL V+GSE+EA+ +Y+ S + +FGF C+ID+E
Sbjct: 313 HWLIVMDKPGGEGASKQQMIDCYVQTLAKVLGSEEEAKKKIYNVSCERYFGFGCEIDDET 492
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S++L LPGVL V PD + KDY
Sbjct: 493 SNKLEGLPGVLFVLPDSYVDPENKDY 570
>TC78973 similar to PIR|D84745|D84745 plastid protein [imported] - Arabidopsis
thaliana, partial (69%)
Length = 1583
Score = 94.4 bits (233), Expect = 7e-20
Identities = 48/117 (41%), Positives = 74/117 (63%)
Frame = +3
Query: 178 LFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGF 237
LFP + HWL+ +DKPG E TK Q++D Y + L +V+G+E++A+ IY+VS + FGF
Sbjct: 933 LFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGF 1112
Query: 238 CCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSVPNSSEASQEASLRTKKL 294
CELDE+ + L G+PGV V PD + E+++Y L V + E Q + R +++
Sbjct: 1113 GCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFV--NGEIVQRSPERQRRV 1277
Score = 86.7 bits (213), Expect = 1e-17
Identities = 42/93 (45%), Positives = 63/93 (67%)
Frame = +3
Query: 66 PSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFC 125
P +N HW++++DKP +K Q+ID YVKTL V+GSE+EA+ +Y+ S + +FGF
Sbjct: 939 PGCDYN-HWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFG 1115
Query: 126 CDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
C++DEE S++L +PGVL V PD + +DY
Sbjct: 1116 CELDEETSNKLEGIPGVLFVLPDSYVDPEHQDY 1214
>TC89523 weakly similar to GP|17065276|gb|AAL32792.1 Unknown protein
{Arabidopsis thaliana}, partial (64%)
Length = 729
Score = 87.8 bits (216), Expect = 6e-18
Identities = 41/82 (50%), Positives = 60/82 (73%)
Frame = +3
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV GLSF+T+EKTL AF +G++V+ KVI D+IS++SKGY F+ + +++ A A+ E
Sbjct: 129 KLFVGGLSFHTTEKTLSDAFSNYGQVVEAKVITDRISEKSKGYGFVTFASQDEAEKAITE 308
Query: 353 MNGKIINGWMIVVDVAKTNPPR 374
MN K +NG ++ VD AK + R
Sbjct: 309 MNEKALNGRVVFVDYAKPDTKR 374
>TC80442 similar to PIR|F84793|F84793 probable RNA-binding protein
[imported] - Arabidopsis thaliana, partial (64%)
Length = 894
Score = 85.5 bits (210), Expect = 3e-17
Identities = 42/88 (47%), Positives = 61/88 (68%)
Frame = +1
Query: 287 ASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAA 346
++L + KLF++GLS T+++ L AF FG+L++ KVI D+ S RSKG+AF+ Y+T E A
Sbjct: 274 STLTSPKLFISGLSRLTTDEKLTEAFSPFGQLLEAKVITDRGSGRSKGFAFVSYSTIEEA 453
Query: 347 GAALKEMNGKIINGWMIVVDVAKTNPPR 374
A + MN K ++GW+I VD AK PR
Sbjct: 454 EKAREGMNAKFLDGWVIFVDPAKPREPR 537
>TC79604 similar to GP|18175762|gb|AAL59923.1 unknown protein {Arabidopsis
thaliana}, partial (56%)
Length = 727
Score = 82.8 bits (203), Expect = 2e-16
Identities = 38/98 (38%), Positives = 64/98 (64%)
Frame = +2
Query: 276 SVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGY 335
S +S ++++ + + LFV+GLS T+ +TLR AF+ FGE+V +V+ D++S SKG+
Sbjct: 191 SFSSSPPSARQMAEPSTNLFVSGLSKRTTTETLREAFQKFGEVVHARVVTDRVSGYSKGF 370
Query: 336 AFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTNPP 373
F++Y T E A ++ M+GK + GW+I + A+ PP
Sbjct: 371 GFVKYATLEDAAKGIEGMDGKFLEGWVIFAEYARPRPP 484
>BI266384
Length = 626
Score = 73.9 bits (180), Expect = 9e-14
Identities = 40/91 (43%), Positives = 60/91 (64%), Gaps = 1/91 (1%)
Frame = +2
Query: 280 SSEASQEASLRTK-KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFI 338
SSE+S E ++ + K+F+ GL T+E L F FGE+ QVK++IDK + +S G+A+I
Sbjct: 170 SSESSVECNMPSSTKIFIKGLPLSTTELHLTKVFSMFGEVTQVKLLIDKKTGQSLGFAYI 349
Query: 339 EYTTEEAAGAALKEMNGKIINGWMIVVDVAK 369
+ E++A +A KEMNGK+ G I V +AK
Sbjct: 350 WFVEEDSAQSAAKEMNGKLFYGRFIYVTIAK 442
>TC80424 homologue to GP|21593052|gb|AAM65001.1 DAG protein putative
{Arabidopsis thaliana}, partial (57%)
Length = 1039
Score = 72.4 bits (176), Expect = 3e-13
Identities = 38/93 (40%), Positives = 55/93 (58%)
Frame = +2
Query: 66 PSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFC 125
P +N HW+++M+ P S+ Q+ID Y++TL TV+GS +EA+ +Y S T+ GF
Sbjct: 167 PGCDYN-HWLIVMEFPKDPAPSRDQMIDTYLQTLATVLGSMEEAKKNMYAFSTTTYTGFQ 343
Query: 126 CDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
C +DE S + LPGVL V PD + DY
Sbjct: 344 CTVDEATSEKFKGLPGVLWVLPDSYIDVKNMDY 442
Score = 67.0 bits (162), Expect = 1e-11
Identities = 37/118 (31%), Positives = 61/118 (51%)
Frame = +2
Query: 154 LKKDYSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILT 213
L +DYS S + R ++ P + HWL+ M+ P ++ Q++D Y Q L
Sbjct: 104 LDEDYSSKR-----SGSNEQRETIMLPGCDYNHWLIVMEFPKDPAPSRDQMIDTYLQTLA 268
Query: 214 KVMGNEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
V+G+ ++A+ +Y S T GF C +DE ++ G+PGV V PD + +N +Y
Sbjct: 269 TVLGSMEEAKKNMYAFSTTTYTGFQCTVDEATSEKFKGLPGVLWVLPDSYIDVKNMDY 442
>BM813813 similar to GP|7549634|gb| DAG protein putative {Arabidopsis
thaliana}, partial (58%)
Length = 605
Score = 72.4 bits (176), Expect = 3e-13
Identities = 35/86 (40%), Positives = 54/86 (62%)
Frame = +1
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++M+ P S+ ++++ YVKTL V+GSE+EA+ +Y S T+ GF + EE+
Sbjct: 76 HWLIIMEFPDNPKPSEDEMVNSYVKTLAQVLGSEEEAKKKIYSVSTSTYIGFGALVSEEL 255
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S ++ LPGVL V PD + KDY
Sbjct: 256 SYKIKELPGVLWVLPDSYLDVPNKDY 333
Score = 60.5 bits (145), Expect = 1e-09
Identities = 32/105 (30%), Positives = 58/105 (54%)
Frame = +1
Query: 173 TRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWK 232
++ +L + +HWL+ M+ P ++ ++V+ Y + L +V+G+E++A+ IY VS
Sbjct: 37 SKETILLDGCDYEHWLIIMEFPDNPKPSEDEMVNSYVKTLAQVLGSEEEAKKKIYSVSTS 216
Query: 233 TNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
T GF + E+ + + +PGV V PD + NK+Y L V
Sbjct: 217 TYIGFGALVSEELSYKIKELPGVLWVLPDSYLDVPNKDYGGDLFV 351
>TC78441 similar to GP|16226543|gb|AAL16196.1 AT3g15000/K15M2_14
{Arabidopsis thaliana}, partial (43%)
Length = 1476
Score = 71.6 bits (174), Expect = 5e-13
Identities = 55/173 (31%), Positives = 84/173 (47%), Gaps = 35/173 (20%)
Frame = +3
Query: 21 LPTNITLP---NKSYHSNLPSSSSSSISCNRFFTIIAAAAATT----YPSFTPSTTH--- 70
+P +TL ++S+ + + SSS++S R ++A A T+ PSF +T
Sbjct: 51 IPKTLTLTTFLSRSFSTTTTNPSSSALSFLRRLRPLSATAITSRHILLPSFRAFSTRPTT 230
Query: 71 -----------NR--------------HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGS 105
NR HW+V+M+KP G ++ ++ID Y+KTL V+GS
Sbjct: 231 SSLNDPNPNWSNRPPKETILLDGCDFEHWLVVMEKPE-GDPTRDEIIDSYIKTLAEVIGS 407
Query: 106 EKEAQMCLYDASWDTHFGFCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
E+EA+ +Y S +F F EE+S +L LP V V PD N +KDY
Sbjct: 408 EEEARKSIYSVSTRHYFAFGALCSEELSYKLKELPKVRWVLPDSYLNVREKDY 566
Score = 56.6 bits (135), Expect = 2e-08
Identities = 32/98 (32%), Positives = 53/98 (53%)
Frame = +3
Query: 174 RTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKT 233
+ +L + +HWLV M+KP + T+ +I+D Y + L +V+G+E++A+ IY VS +
Sbjct: 276 KETILLDGCDFEHWLVVMEKPEGDP-TRDEIIDSYIKTLAEVIGSEEEARKSIYSVSTRH 452
Query: 234 NFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
F F E+ + L +P V V PD K+Y
Sbjct: 453 YFAFGALCSEELSYKLKELPKVRWVLPDSYLNVREKDY 566
>TC84335 weakly similar to GP|21741678|emb|CAD40580. oj000126_13.2 {Oryza
sativa}, partial (39%)
Length = 678
Score = 69.3 bits (168), Expect = 2e-12
Identities = 32/81 (39%), Positives = 52/81 (63%)
Frame = +2
Query: 291 TKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAAL 350
T K+FV GL+F T+E+ L AF +G +V+ ++++K KR KG+ ++ + EE A A
Sbjct: 197 TSKIFVKGLAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQ 376
Query: 351 KEMNGKIINGWMIVVDVAKTN 371
MNGKI++G ++ VD+ N
Sbjct: 377 IGMNGKILHGRVLYVDMDPPN 439
>TC81180 similar to GP|7549634|gb|AAF63819.1| DAG protein putative
{Arabidopsis thaliana}, partial (50%)
Length = 687
Score = 68.6 bits (166), Expect = 4e-12
Identities = 32/76 (42%), Positives = 50/76 (65%)
Frame = +3
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++M+ P S+ ++++ YVKTL V+GSE+EA+ +Y S T+ GF + EE+
Sbjct: 285 HWLIIMEFPDNPKPSEDEMVNSYVKTLAQVLGSEEEAKKKIYSVSTSTYIGFGALVSEEL 464
Query: 133 SSQLASLPGVLSVRPD 148
S ++ LPGVL V PD
Sbjct: 465 SYKIKELPGVLWVLPD 512
Score = 55.5 bits (132), Expect = 3e-08
Identities = 40/142 (28%), Positives = 66/142 (46%)
Frame = +3
Query: 120 THFGFCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDYSLSSGQAGLSSGLQTRTNMLF 179
T F F S Q +P VR F S YS + + S + +L
Sbjct: 108 TRFAFALS---SASRQTLPIPHSFPVR----FKSSGSGYSPLNDPSPNWSNRPPKETILL 266
Query: 180 PAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCC 239
+ +HWL+ M+ P ++ ++V+ Y + L +V+G+E++A+ IY VS T GF
Sbjct: 267 DGCDYEHWLIIMEFPDNPKPSEDEMVNSYVKTLAQVLGSEEEAKKKIYSVSTSTYIGFGA 446
Query: 240 ELDEDCAQDLAGVPGVSSVQPD 261
+ E+ + + +PGV V PD
Sbjct: 447 LVSEELSYKIKELPGVLWVLPD 512
>TC77756 similar to PIR|T06796|T06796 glycine-rich RNA-binding protein -
garden pea, partial (89%)
Length = 813
Score = 67.8 bits (164), Expect = 7e-12
Identities = 36/95 (37%), Positives = 61/95 (63%)
Frame = +3
Query: 274 SLSVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSK 333
S P SS + + + KLF+ GLS+ +++LR AF +G++V+ +VI D+ + RS+
Sbjct: 132 STQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSR 311
Query: 334 GYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVA 368
G+ F+ +T+EE+A +AL M+G+ +NG I V A
Sbjct: 312 GFGFVNFTSEESATSAL-SMDGQDLNGRNIRVSYA 413
>BE124733 similar to PIR|T06796|T06 glycine-rich RNA-binding protein - garden
pea, partial (86%)
Length = 580
Score = 64.7 bits (156), Expect = 6e-11
Identities = 31/85 (36%), Positives = 57/85 (66%)
Frame = +3
Query: 274 SLSVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSK 333
S P SS + + + KLF+ GLS+ +++LR AF +G++V+ +VI D+ + RS+
Sbjct: 90 STQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSR 269
Query: 334 GYAFIEYTTEEAAGAALKEMNGKII 358
G+ F+ +T+EE+A +AL M+G+++
Sbjct: 270 GFGFVNFTSEESATSAL-SMDGQVV 341
>CA917955
Length = 870
Score = 64.7 bits (156), Expect = 6e-11
Identities = 40/104 (38%), Positives = 60/104 (57%), Gaps = 14/104 (13%)
Frame = +2
Query: 280 SSEASQEASLRTK-KLFVTGLSFYTSEKTLRAAFEGFGELVQ-------------VKVII 325
SSE+S E ++ + K+F+ GL T+E L F FGE+ Q VK++I
Sbjct: 113 SSESSVECNMPSSTKIFIKGLPLSTTELHLTKVFSMFGEVTQGN*L*IYFGCFQRVKLLI 292
Query: 326 DKISKRSKGYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAK 369
DK + +S G+A+I + E++A +A KEMNGK+ G I V +AK
Sbjct: 293 DKKTGQSLGFAYIWFVEEDSAQSAAKEMNGKLFYGRFIYVTIAK 424
>AW687209 weakly similar to GP|9757907|dbj| contains similarity to RNA
binding protein~gene_id:MMN10.9 {Arabidopsis thaliana},
partial (64%)
Length = 531
Score = 63.9 bits (154), Expect = 1e-10
Identities = 30/72 (41%), Positives = 48/72 (66%)
Frame = +2
Query: 298 GLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMNGKI 357
GLSFYT+++ L + F FG L + +I D+ ++R KG+ F+ Y +E A A K +NG+I
Sbjct: 206 GLSFYTTQQQLESLFSPFGVLTEATLITDQNTQRPKGFGFVSYKSEIEAEKARKALNGRI 385
Query: 358 INGWMIVVDVAK 369
++G +I V+ AK
Sbjct: 386 VDGRLIFVEHAK 421
>BQ139859 similar to GP|8777484|dbj contains similarity to RNA-binding
protein~gene_id:K15M2.15 {Arabidopsis thaliana}, partial
(29%)
Length = 562
Score = 62.8 bits (151), Expect = 2e-10
Identities = 41/98 (41%), Positives = 57/98 (57%), Gaps = 3/98 (3%)
Frame = +3
Query: 274 SLSVPNSSEASQEASLRT---KKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISK 330
SL P+ +A + + R +KLFV GLS T+ +TLR F +G+L + VI DK +
Sbjct: 240 SLQHPDILDAVRSVADRDTTRRKLFVRGLSGETTTETLRTVFSSYGDLDEAIVIFDKATG 419
Query: 331 RSKGYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVA 368
RSKGY F+ Y + A ALKE + K I+G M V +A
Sbjct: 420 RSKGYGFVVYRHVDGAVLALKEPSKK-IDGRMTVTQLA 530
>TC76948 similar to SP|Q08935|ROC1_NICSY 29 kDa ribonucleoprotein A
chloroplast precursor (CP29A). [Wood tobacco] {Nicotiana
sylvestris}, partial (61%)
Length = 1374
Score = 60.8 bits (146), Expect = 8e-10
Identities = 34/120 (28%), Positives = 58/120 (48%)
Frame = +1
Query: 255 VSSVQPDDNFESENKNYEDSLSVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEG 314
+S V F+ E ++D P+ S ++LFV L F L FE
Sbjct: 253 ISRVAVSSEFDQEEDTFDDGDDTPSYSP--------NQRLFVGNLPFSVDSAQLAEIFEN 408
Query: 315 FGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTNPPR 374
G++ V+VI DK + RS+G+ F+ ++ AA +++NG +++G + V+ PPR
Sbjct: 409 AGDVEMVEVIYDKSTGRSRGFGFVTMSSAAEVEAAAQQLNGYVVDGRELRVNAGPPPPPR 588
Score = 47.8 bits (112), Expect = 7e-06
Identities = 23/82 (28%), Positives = 47/82 (57%)
Frame = +1
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
++ V L++ L + F G++++ KVI D+ S RS+G+ F+ +++ + +A++
Sbjct: 679 RVHVGNLAWGVDNLALESLFGEQGQVLEAKVIYDRESGRSRGFGFVTFSSADEVDSAIRT 858
Query: 353 MNGKIINGWMIVVDVAKTNPPR 374
++G +NG I V A + P R
Sbjct: 859 LDGADLNGRAIRVSPADSRPKR 924
>BE942328 weakly similar to GP|21741678|emb oj000126_13.2 {Oryza sativa},
partial (34%)
Length = 548
Score = 60.8 bits (146), Expect = 8e-10
Identities = 27/67 (40%), Positives = 43/67 (63%)
Frame = +3
Query: 291 TKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAAL 350
T K+FV GL+F T+E+ L AF +G +V+ ++++K KR KG+ ++ + EE A A
Sbjct: 111 TSKIFVKGLAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQ 290
Query: 351 KEMNGKI 357
MNGK+
Sbjct: 291 IGMNGKV 311
>TC89001 similar to GP|5726567|gb|AAD48471.1| glycine-rich RNA-binding
protein {Glycine max}, partial (90%)
Length = 488
Score = 60.5 bits (145), Expect = 1e-09
Identities = 30/78 (38%), Positives = 50/78 (63%)
Frame = +3
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
+ FV GL++ T + L AF FGE+ KVI D+ + RS+G+ F+ + E++ A++E
Sbjct: 54 RCFVGGLAWATDSQALEQAFSKFGEITDSKVINDRETGRSRGFGFVTFAEEKSMRDAIEE 233
Query: 353 MNGKIINGWMIVVDVAKT 370
MNG+ I+G I V+ A++
Sbjct: 234 MNGQDIDGRNITVNEAQS 287
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.130 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,673,594
Number of Sequences: 36976
Number of extensions: 188330
Number of successful extensions: 1455
Number of sequences better than 10.0: 152
Number of HSP's better than 10.0 without gapping: 1383
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1442
length of query: 384
length of database: 9,014,727
effective HSP length: 98
effective length of query: 286
effective length of database: 5,391,079
effective search space: 1541848594
effective search space used: 1541848594
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0095c.6


