

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094b.3
(337 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
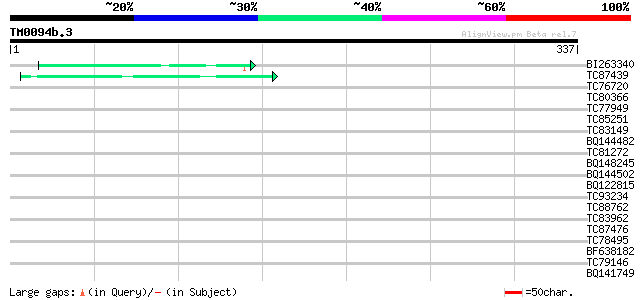
BI263340 weakly similar to PIR|T49223|T492 uclacyanin 3 [importe... 42 4e-04
TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in... 41 7e-04
TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {... 40 0.001
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 40 0.001
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 40 0.002
TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean... 38 0.006
TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk p... 38 0.006
BQ144482 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 36 0.024
TC81272 similar to PIR|T05225|T05225 extensin homolog F17I5.160 ... 36 0.024
BQ148245 GP|21554860|gb unknown {Arabidopsis thaliana}, partial ... 35 0.054
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 34 0.070
BQ122815 weakly similar to OMNI|MT3615.1 PE_PGRS family protein ... 34 0.070
TC93234 weakly similar to GP|21751020|dbj|BAC03887. unnamed prot... 34 0.070
TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Ar... 34 0.070
TC83962 similar to PIR|B86289|B86289 hypothetical protein AAF719... 34 0.070
TC87476 homologue to PIR|T11622|T11622 extensin class 1 precurso... 34 0.091
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 34 0.091
BF638182 weakly similar to PIR|T45594|T45 glucosidase-like prote... 34 0.091
TC79146 similar to GP|9757884|dbj|BAB08471.1 RuvB DNA helicase-l... 33 0.12
BQ141749 homologue to GP|20160651|dbj hypothetical protein {Oryz... 33 0.16
>BI263340 weakly similar to PIR|T49223|T492 uclacyanin 3 [imported] -
Arabidopsis thaliana, partial (14%)
Length = 681
Score = 41.6 bits (96), Expect = 4e-04
Identities = 37/133 (27%), Positives = 53/133 (39%), Gaps = 4/133 (3%)
Frame = +3
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P + SP P ++ GGG+ P P T P S + +QTP
Sbjct: 159 PHRSTSPSHGSSSPPSHGSSSPPSHGGGYYTPTPPSTGCGYSPPHDPSTSTPSHNQTPST 338
Query: 78 P-RPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSP- 135
P P + SP P P+ VTL P +P++P G + PT P SP
Sbjct: 339 PSNPPSSGGYYNSP----PSTPTDPPVTLTPPSPSTPIDPG-----TPTVPTPPFLPSPS 491
Query: 136 PYP--FSFLLSHP 146
P+P ++ +HP
Sbjct: 492 PFPGTCNYWRTHP 530
>TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in cucumber
hypocotyls {Cucumis sativus}, partial (57%)
Length = 897
Score = 40.8 bits (94), Expect = 7e-04
Identities = 39/153 (25%), Positives = 58/153 (37%)
Frame = +3
Query: 7 VAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHP 66
VA VGG+ SP +A + S P ++P+ P PV + + P S
Sbjct: 84 VAGVGGQ---SPSAAPTTSPSTPAATTPVSSPVAAPPTTPTTPAPVASPKSSPPATSPKA 254
Query: 67 SEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQ 126
+ P A P + +P S PP P PVK P+ +SP A
Sbjct: 255 A------APTATPPAASSPPAVTPVSTPPPAPVPVKSPPTPAPVSSP---------PAVT 389
Query: 127 PTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
P +++P P +K GAP+ +P
Sbjct: 390 PVAAPTTTPAVPAPAPSKGKKNKKKHGAPAPSP 488
Score = 37.0 bits (84), Expect = 0.011
Identities = 31/110 (28%), Positives = 47/110 (42%)
Frame = +3
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P ++ P + V P P P+ P PV + A P+A+ + TP
Sbjct: 276 PAASSPPAVTPVSTPPPAPVPVKSPP----TPAPVSSPPAVTPVAA-------PTTTPAV 422
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQP 127
P P ++G+++ P PSP LGP AP + G G E+ S+ P
Sbjct: 423 PAPAP-SKGKKNKKKHGAPAPSP--ALLGPPAPPA-GAPGPSEDASSPGP 560
>TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {Nicotiana
sylvestris}, partial (51%)
Length = 1290
Score = 40.4 bits (93), Expect = 0.001
Identities = 27/98 (27%), Positives = 42/98 (42%), Gaps = 2/98 (2%)
Frame = +1
Query: 34 PQTTPLTGTGGGFVAPDPVVTDSACD--PLASDHPSEVNASQTPDAPRPVTLAEGQRSPS 91
P TP T + +P P+ T S + P + P+ S +P AP P SPS
Sbjct: 562 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 741
Query: 92 SIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTG 129
P PSP + +A + G+G +G++ +G
Sbjct: 742 PSSSPAPSPDEAA-DNNAISHTGIGEDGKSSGGGMSSG 852
Score = 30.8 bits (68), Expect = 0.77
Identities = 25/75 (33%), Positives = 31/75 (41%), Gaps = 3/75 (4%)
Frame = +1
Query: 69 VNASQTPDAPRPVTLAEGQ-RSPSSIRPPIPSPVKVTLGPSAPN-SPGVGGEGENLSAAQ 126
VN + + D+P P SPS P PSP P AP +P ++ A
Sbjct: 535 VNIASSADSPAPTPATNSSLNSPSPTPIPTPSPAN---SPPAPTPTPTPSPHSDSPPAPS 705
Query: 127 PTGPSSSSP-PYPFS 140
P SSSP P P S
Sbjct: 706 PDNSPSSSPSPSPSS 750
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 40.0 bits (92), Expect = 0.001
Identities = 36/116 (31%), Positives = 43/116 (37%)
Frame = +3
Query: 23 SPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVT 82
SP + P P T A P VT SA P PS S TP AP P T
Sbjct: 6 SPPPATPSSPPPSTP----------ANPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPST 155
Query: 83 LAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+ SP + P I P PS P+ + P+ PSS +PP P
Sbjct: 156 PSNSPPSPPTT-PAISPPSGGGTTPSPPSRTPPSSDDS------PSPPSSKTPPPP 302
Score = 31.2 bits (69), Expect = 0.59
Identities = 27/98 (27%), Positives = 35/98 (35%), Gaps = 15/98 (15%)
Frame = +3
Query: 18 PVSACSPKRSKVGEPDPQT--TPLTGTGGGFVAPDPVVTDSACDPLASDH---------- 65
P + SP S P P T P T +P P T SA P +
Sbjct: 15 PATPSSPPPSTPANPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPPTTPA 194
Query: 66 ---PSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSP 100
PS + +P + P + + PSS PP PSP
Sbjct: 195 ISPPSGGGTTPSPPSRTPPSSDDSPSPPSSKTPPPPSP 308
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 39.7 bits (91), Expect = 0.002
Identities = 37/120 (30%), Positives = 47/120 (38%), Gaps = 1/120 (0%)
Frame = +3
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVV-TDSACDPLASDHPSEVNASQT 74
SSPV P S P P + G +P PVV T A P+AS S +
Sbjct: 615 SSPVPTSGPTASS---PSPVVSTPPAGGPMASSPSPVVSTPPAGGPMAS--------SPS 761
Query: 75 PDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSS 134
P P G S ++ PP P P ++P G G G +AA P P S+S
Sbjct: 762 PAGGPPALSPAGGPSTAAGGPPAPGPGGAA---TSPGPGGAGASGPGGAAAAPGSPGSNS 932
Score = 30.0 bits (66), Expect = 1.3
Identities = 28/87 (32%), Positives = 36/87 (41%), Gaps = 2/87 (2%)
Frame = +3
Query: 52 VVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIR-PPIPSPVKVTLGPSAP 110
+V S P +S PS S +P T + PS + PP SPV T GP+A
Sbjct: 474 LVVISPRTPKSSPSPSAGGLSPPSPSPTTTTPSPSGSPPSPVAIPPASSPVP-TSGPTAS 650
Query: 111 N-SPGVGGEGENLSAAQPTGPSSSSPP 136
+ SP V A P S+PP
Sbjct: 651 SPSPVVSTPPAGGPMASSPSPVVSTPP 731
>TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean, partial
(93%)
Length = 1554
Score = 37.7 bits (86), Expect = 0.006
Identities = 34/123 (27%), Positives = 44/123 (35%), Gaps = 3/123 (2%)
Frame = -2
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAP---DPVVTDSACDPLASDHPSEVNAS 72
S P +PK + V P P+ P + P PV T + PL
Sbjct: 749 SQPPVTVAPKSAPVTSPAPKIAPASSPKVPPPQPPKSSPVSTPTLPPPLPPPPKISPTPV 570
Query: 73 QTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSS 132
QTP AP PV + P P+P K P+ SP + + A P P S
Sbjct: 569 QTPPAPAPV---------KATPVPAPAPAKQAPTPAPATSPPIPAPTPAIEAPVP-APES 420
Query: 133 SSP 135
S P
Sbjct: 419 SKP 411
Score = 30.4 bits (67), Expect = 1.0
Identities = 29/100 (29%), Positives = 40/100 (40%), Gaps = 1/100 (1%)
Frame = -2
Query: 2 SVKPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL 61
S P++A + P PK S V P T P ++P PV T A P+
Sbjct: 704 SPAPKIAPASSPKVPPPQP---PKSSPVSTP---TLPPPLPPPPKISPTPVQTPPAPAPV 543
Query: 62 -ASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSP 100
A+ P+ A Q P P P T +I P+P+P
Sbjct: 542 KATPVPAPAPAKQAP-TPAPATSPPIPAPTPAIEAPVPAP 426
Score = 30.0 bits (66), Expect = 1.3
Identities = 25/93 (26%), Positives = 36/93 (37%), Gaps = 5/93 (5%)
Frame = -2
Query: 51 PVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP-SA 109
PV P+ S P AS +P P P S ++ PP+P P K++ P
Sbjct: 740 PVTVAPKSAPVTSPAPKIAPAS-SPKVPPPQPPKSSPVSTPTLPPPLPPPPKISPTPVQT 564
Query: 110 PNSP----GVGGEGENLSAAQPTGPSSSSPPYP 138
P +P + PT ++SPP P
Sbjct: 563 PPAPAPVKATPVPAPAPAKQAPTPAPATSPPIP 465
>TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk protein
{Argiope trifasciata}, partial (3%)
Length = 1177
Score = 37.7 bits (86), Expect = 0.006
Identities = 26/92 (28%), Positives = 37/92 (39%), Gaps = 2/92 (2%)
Frame = +3
Query: 23 SPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVT 82
SP R + +P P + T P P + P PS + P P T
Sbjct: 216 SPPRMRRSKPSPSSPAATPLWPQRTTPPPSPSTPRPSP*TPATPSTSPTAPLPTPPPRTT 395
Query: 83 L--AEGQRSPSSIRPPIPSPVKVTLGPSAPNS 112
L A R PS PP P P V++ P++P++
Sbjct: 396 LPPALTPRPPSPSTPPTPKPGPVSVSPASPSA 491
Score = 33.1 bits (74), Expect = 0.16
Identities = 25/89 (28%), Positives = 36/89 (40%)
Frame = +3
Query: 48 APDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP 107
+P P ++ S PS S+TP P P L P +PP SP ++
Sbjct: 81 SPTPSAPRTSSRSTPSTSPSRRPRSKTPLPPPPPPL------PQHPKPPQKSPPRMRRSK 242
Query: 108 SAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
+P+SP AA P P ++PP
Sbjct: 243 PSPSSP----------AATPLWPQRTTPP 299
>BQ144482 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (16%)
Length = 993
Score = 35.8 bits (81), Expect = 0.024
Identities = 22/75 (29%), Positives = 30/75 (39%)
Frame = +3
Query: 64 DHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLS 123
D P + P P P+ P +RPP PSP L P +P+ P +
Sbjct: 150 DRPPPPKVAPRPPFPPPL--------PFYLRPPSPSPSPPPLSPPSPSPPPPPPQPLLGP 305
Query: 124 AAQPTGPSSSSPPYP 138
+P GP +PP P
Sbjct: 306 GLRPPGPRPPAPPPP 350
Score = 28.1 bits (61), Expect = 5.0
Identities = 27/117 (23%), Positives = 36/117 (30%), Gaps = 12/117 (10%)
Frame = +2
Query: 12 GKRQSSPVSACSPKRSKVGE-PDPQTTPLTGTGGGFVAPDPVVTDSACD----------- 59
G R+ P + G P P P + F P P+++
Sbjct: 83 GGRRGGRAGRARPGAGRGGAAPRPAPAPQSRPAPTFSPPPPLLSPPPLPLSLPSPSFAPL 262
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVG 116
PL P + P PRP R+P P P PV P+ P P G
Sbjct: 263 PLPPPPPPSASPWPRPSPPRP----PAPRAPPPRPGPPPGPVPPECWPAPPGGPVTG 421
>TC81272 similar to PIR|T05225|T05225 extensin homolog F17I5.160 -
Arabidopsis thaliana, partial (3%)
Length = 757
Score = 35.8 bits (81), Expect = 0.024
Identities = 28/108 (25%), Positives = 36/108 (32%), Gaps = 1/108 (0%)
Frame = +3
Query: 32 PDPQTTPLTGTGGGFVAPDPVVTDSACD-PLASDHPSEVNASQTPDAPRPVTLAEGQRSP 90
P P TP+ GG P P + + P AS P + P P Q P
Sbjct: 102 PVPLPTPIPTGPGGVPQPGPGASGAPQPGPGASGAPQPGPGASGAPQPGPGASGAPQPGP 281
Query: 91 SSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+ P P P P + G G S A P +S P+P
Sbjct: 282 GASGAPQPGPDASGAPQPGPGASGAPQPGPGASGAPQPAPEASGAPHP 425
Score = 29.6 bits (65), Expect = 1.7
Identities = 38/146 (26%), Positives = 51/146 (34%), Gaps = 1/146 (0%)
Frame = +1
Query: 2 SVKPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL 61
S + A + SS +P S + P P V P P + L
Sbjct: 7 STRSTSAPMASVTSSSLARRTAPSPSLPSQSAPSPCPPRSPPAPVVFPSPAPVPAVLLSL 186
Query: 62 ASDHPSEVN-ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGE 120
A P ++ A P P PV SP + +PSP + P AP SP +
Sbjct: 187 ALVLPVLLSPALVLPVLPSPVLALPVLPSPVPVPLALPSPALM---PVAPLSPALVLPAP 357
Query: 121 NLSAAQPTGPSSSSPPYPFSFLLSHP 146
LS A S PP P + L+ P
Sbjct: 358 -LSPALVLPVLPSPPPRPAALLIPDP 432
>BQ148245 GP|21554860|gb unknown {Arabidopsis thaliana}, partial (8%)
Length = 432
Score = 34.7 bits (78), Expect = 0.054
Identities = 27/100 (27%), Positives = 41/100 (41%), Gaps = 1/100 (1%)
Frame = +3
Query: 49 PDPVVTD-SACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP 107
P P+++ S DP+ +S T P P +++ Q I P +PSP G
Sbjct: 96 PSPILSSYSQSDPIPELPTLPTTSSATKTDPSPSSISPFQNLSPEIAPLLPSP-----GG 260
Query: 108 SAPNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPY 147
+ P G + PT PS+ SPP P + P+
Sbjct: 261 ALPTPTG---------SDIPTIPSNPSPPNPDDVIAPGPF 353
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 34.3 bits (77), Expect = 0.070
Identities = 39/137 (28%), Positives = 46/137 (33%), Gaps = 2/137 (1%)
Frame = -1
Query: 7 VAEVGGKRQSSPVSAC--SPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASD 64
VA V G R + A P+ + P P T P G G P P T + P A
Sbjct: 392 VAVVPGPRSGNSPRAAHRGPEPRPLPRPPPPTRPPAAPGPG-PPPAPGPTPLSKPPAAEG 216
Query: 65 HPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSA 124
P A P A R PS+ P P P P AP P +L
Sbjct: 215 ---------IPPATAPPPRARRLRPPSA---PAPPPTPPAAAPPAPTPPHAPA*APHL-- 78
Query: 125 AQPTGPSSSSPPYPFSF 141
P P + PP P F
Sbjct: 77 --PPPPPAPRPPRPPPF 33
Score = 32.3 bits (72), Expect = 0.27
Identities = 22/77 (28%), Positives = 30/77 (38%), Gaps = 4/77 (5%)
Frame = -2
Query: 66 PSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAA 125
P ++ + PRP A + PS PP P T P SP G+G+
Sbjct: 523 PEPLHNPRPRPPPRPPPAAPARAPPSG--PPAPPTPPATRAPPVRASPWCPGQGQETPQG 350
Query: 126 QPTGPSSSSP----PYP 138
+ TG S P P+P
Sbjct: 349 RHTGVQSPVPCPALPHP 299
Score = 30.8 bits (68), Expect = 0.77
Identities = 28/105 (26%), Positives = 36/105 (33%)
Frame = -1
Query: 32 PDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPS 91
PDP P G V P P S P A+ E PRP +P
Sbjct: 431 PDPAGHPRAPRQGVAVVPGP---RSGNSPRAAHRGPEPRP-----LPRPPPPTRPPAAPG 276
Query: 92 SIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
PP P P ++ P+A P +P PS+ +PP
Sbjct: 275 PGPPPAPGPTPLSKPPAAEGIPPATAPPPRARRLRP--PSAPAPP 147
>BQ122815 weakly similar to OMNI|MT3615.1 PE_PGRS family protein
{Mycobacterium tuberculosis CDC1551}, partial (2%)
Length = 590
Score = 34.3 bits (77), Expect = 0.070
Identities = 23/85 (27%), Positives = 32/85 (37%)
Frame = +2
Query: 47 VAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLG 106
V P+ V+ +P +P+ PDAP+P GQ P I +P +
Sbjct: 5 VPPEIVIDPPQFNPFHDPNPNP-----KPDAPKPPLTPPGQDMPFPIPDVVPPRPPPDVV 169
Query: 107 PSAPNSPGVGGEGENLSAAQPTGPS 131
P P P + PTGPS
Sbjct: 170 PPCPPGPDIVPPPSTPPIPPPTGPS 244
>TC93234 weakly similar to GP|21751020|dbj|BAC03887. unnamed protein product
{Homo sapiens}, partial (11%)
Length = 591
Score = 34.3 bits (77), Expect = 0.070
Identities = 40/141 (28%), Positives = 54/141 (37%), Gaps = 1/141 (0%)
Frame = +2
Query: 20 SACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ-TPDAP 78
S+ +RS+ T TG AP V+ A P PS +S + P
Sbjct: 185 SSIPSQRSRTSRSPSSRPRHTSTG---TAPPFAVSSPASSPTT---PSSTRSSPGSTSTP 346
Query: 79 RPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
R + + R P S PP P + T G S+P+S PSSSSPP P
Sbjct: 347 RTPSTSFSARPPPSTPPPTPFRSRPTKG-SSPSST----------------PSSSSPPAP 475
Query: 139 FSFLLSHPYLEKLKGAPSQNP 159
S +S AP++ P
Sbjct: 476 TSPAIS----PSRPSAPTRRP 526
Score = 32.7 bits (73), Expect = 0.20
Identities = 28/98 (28%), Positives = 40/98 (40%)
Frame = +2
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
SSP S+ + S P +TP T P + SA P ++ P TP
Sbjct: 278 SSPASSPTTPSSTRSSPGSTSTPRT----------PSTSFSARPPPSTPPP-------TP 406
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSP 113
RP + +PSS P P+P + PS P++P
Sbjct: 407 FRSRPTKGSSPSSTPSSSSP--PAPTSPAISPSRPSAP 514
>TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 34.3 bits (77), Expect = 0.070
Identities = 32/121 (26%), Positives = 51/121 (41%)
Frame = +3
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P A +P + + Q T A PV T ++ P ++ S V+ + +P A
Sbjct: 21 PGVAQTPSGNGIAPSPYQFNNPTSPSSSPSASSPVSTTTS-PPASTPASSPVSTTTSPPA 197
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPY 137
P + P++ PP P+P P + NSP G +AA P+ PSS++
Sbjct: 198 STPAS----SPVPTTTSPPAPTPAS---SPVSTNSPTASPAGSLPAAATPS-PSSTTAGS 353
Query: 138 P 138
P
Sbjct: 354 P 356
Score = 32.3 bits (72), Expect = 0.27
Identities = 29/97 (29%), Positives = 38/97 (38%), Gaps = 4/97 (4%)
Frame = +3
Query: 16 SSPVSACS--PKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
SSPVS + P + P P TT P A P++++ P+ A
Sbjct: 162 SSPVSTTTSPPASTPASSPVPTTT------------SPPAPTPASSPVSTNSPTASPAGS 305
Query: 74 TPDA--PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS 108
P A P P + G SPSS P S + T G S
Sbjct: 306 LPAAATPSPSSTTAGSPSPSSTPPGTRSSNETTPGRS 416
>TC83962 similar to PIR|B86289|B86289 hypothetical protein AAF71991.1
[imported] - Arabidopsis thaliana, partial (11%)
Length = 679
Score = 34.3 bits (77), Expect = 0.070
Identities = 31/102 (30%), Positives = 44/102 (42%), Gaps = 4/102 (3%)
Frame = +2
Query: 62 ASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGEN 121
A+ P+ ++ S TP P + S SS PP PS T P P S N
Sbjct: 47 ANKSPA*ISNSSTPS---PSSSTSTSTSSSSSPPPTPSTSSTTPSPPPPTSLSSPTPTSN 217
Query: 122 -LSAAQPTGPS---SSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
LS+A PT P+ ++ P P + S+ +L + PS P
Sbjct: 218 HLSSA*PTTPTNTLTAVPSTPLNSPCSNLHLPQQTSLPSPLP 343
>TC87476 homologue to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 939
Score = 33.9 bits (76), Expect = 0.091
Identities = 34/112 (30%), Positives = 38/112 (33%), Gaps = 1/112 (0%)
Frame = +3
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVT-LAEGQRSPSSIRPPIPSPVKVTLGP 107
P P V S P S P V S P +P P P S PP P K P
Sbjct: 387 PPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSASPPPPYYYKSPPPP 566
Query: 108 SAPNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
S P G + P PS S PP PY+ K PS +P
Sbjct: 567 SPSPPPPYGYKS-------PPPPSPSPPP---------PYIYKSPPPPSPSP 674
Score = 32.7 bits (73), Expect = 0.20
Identities = 39/134 (29%), Positives = 46/134 (34%)
Frame = +3
Query: 15 QSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQT 74
+S P + SP V + P +P P P V S P AS P S
Sbjct: 405 KSPPPPSPSPPPPYVYKSPPPPSPSP--------PPPYVYKSPPPPSASPPPPYYYKSPP 560
Query: 75 PDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSS 134
P +P P G +SP PP PSP + S P PS S
Sbjct: 561 PPSPSPPP-PYGYKSPP---PPSPSPPPPYIYKS------------------PPPPSPSP 674
Query: 135 PPYPFSFLLSHPYL 148
PPY HPYL
Sbjct: 675 PPY-------HPYL 695
Score = 32.0 bits (71), Expect = 0.35
Identities = 33/111 (29%), Positives = 38/111 (33%)
Frame = +2
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS 108
P P V S P S P V S P +P P + P PP PSP + S
Sbjct: 50 PPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPP----PPTPSPPPPYIYKS 217
Query: 109 APNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
P P P PS S PP PY+ K PS +P
Sbjct: 218 PP--PPSPSPPPPYVYKSPPPPSPSPPP---------PYVYKSPPPPSPSP 337
Score = 31.6 bits (70), Expect = 0.45
Identities = 32/111 (28%), Positives = 38/111 (33%)
Frame = +2
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS 108
P P + S P S P V S P +P P + P PP PSP + S
Sbjct: 2 PPPYIYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPP----PPSPSPPPPYVYKS 169
Query: 109 APNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
P P P PS S PP PY+ K PS +P
Sbjct: 170 PP--PPTPSPPPPYIYKSPPPPSPSPPP---------PYVYKSPPPPSPSP 289
Score = 27.3 bits (59), Expect = 8.5
Identities = 32/127 (25%), Positives = 42/127 (32%)
Frame = +2
Query: 15 QSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQT 74
+S P + SP V + P +P P P V S P S P + S
Sbjct: 68 KSPPPPSPSPPPPYVYKSPPPPSPSP--------PPPYVYKSPPPPTPSPPPPYIYKSPP 223
Query: 75 PDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSS 134
P +P P + P PP PSP + S P P S S
Sbjct: 224 PPSPSPPPPYVYKSPP----PPSPSPPPPYVYKSPP-------------------PPSPS 334
Query: 135 PPYPFSF 141
PP P+ +
Sbjct: 335 PPPPYIY 355
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 33.9 bits (76), Expect = 0.091
Identities = 43/175 (24%), Positives = 60/175 (33%), Gaps = 12/175 (6%)
Frame = +3
Query: 8 AEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTD----SACDPLAS 63
+ VG ++ S SP P TTP +P PV + + P A
Sbjct: 219 SSVGAQQAPSTSPNSSPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQ 398
Query: 64 DHPSEVNASQTP----DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEG 119
P V +S P P P+T Q +P PP SP + P+ P P
Sbjct: 399 STPPPVQSSPPPVQQSPPPTPLTPPPVQSTPPPASPPPASPPPFSPPPATP-PPATPPPA 575
Query: 120 ENLSAAQPT----GPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEAVLNL 170
A PT P+++ P P P L AP +P D + +L
Sbjct: 576 TPPPALTPTPLSSPPATTPAPAPAKLKSKAPAL-----APVLSPSDAPAPGLSSL 725
>BF638182 weakly similar to PIR|T45594|T45 glucosidase-like protein -
Arabidopsis thaliana, partial (46%)
Length = 674
Score = 33.9 bits (76), Expect = 0.091
Identities = 28/93 (30%), Positives = 35/93 (37%), Gaps = 5/93 (5%)
Frame = +3
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPI---PSPVKVTL 105
P + +A P S HP S P P + Q PSS+ PI PSP
Sbjct: 66 PTTSASTTAPSPTTSHHPQPSPLSSKPKPQSPTLKSSTQTLPSSVPSPIQTSPSP*LSPT 245
Query: 106 GPSAP--NSPGVGGEGENLSAAQPTGPSSSSPP 136
S P NSP + S+ PSS+ P
Sbjct: 246 PTSLPSQNSPPLKNGSPPTSSLSTPKPSSTVSP 344
>TC79146 similar to GP|9757884|dbj|BAB08471.1 RuvB DNA helicase-like protein
{Arabidopsis thaliana}, partial (47%)
Length = 792
Score = 33.5 bits (75), Expect = 0.12
Identities = 26/75 (34%), Positives = 37/75 (48%), Gaps = 5/75 (6%)
Frame = -1
Query: 88 RSPSSIRPPI--PSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSH 145
+SP S PP PSP T ++P+SP + ++ S +P P SPP+P F
Sbjct: 657 QSPQSSSPPNHKPSPSPSTSTSTSPSSPQLQPPPDDQS--EPQQPHLQSPPFPP*FSHQS 484
Query: 146 PY---LEKLKGAPSQ 157
PY KL+ + SQ
Sbjct: 483 PYEKPASKLRFSTSQ 439
>BQ141749 homologue to GP|20160651|dbj hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1321
Score = 33.1 bits (74), Expect = 0.16
Identities = 24/83 (28%), Positives = 39/83 (46%), Gaps = 5/83 (6%)
Frame = -1
Query: 61 LASDHPSEVNASQTPDAPRPV--TLAEG-QRSPSSIRPPIPSPVKVTLGPSAPNSPGVGG 117
+ D P E A+ +P +P P ++ +G + S+RP I P + L NSPG+ G
Sbjct: 565 IEGDAPHEPRATPSPPSPTPFLNSILQGTSHNTESLRPRIVPPPYLPLQSLPGNSPGIPG 386
Query: 118 --EGENLSAAQPTGPSSSSPPYP 138
EN++ + S+P P
Sbjct: 385 IRGPENINESTKKYKEKSTPNTP 317
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,855,809
Number of Sequences: 36976
Number of extensions: 133013
Number of successful extensions: 1106
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 982
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1058
length of query: 337
length of database: 9,014,727
effective HSP length: 97
effective length of query: 240
effective length of database: 5,428,055
effective search space: 1302733200
effective search space used: 1302733200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0094b.3


