

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094a.6
(990 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
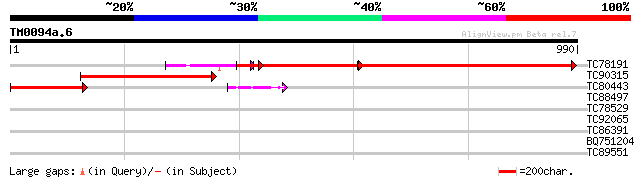
TC78191 homologue to GP|13111324|dbj|BAB32793. 110 kDa protein H... 640 0.0
TC90315 homologue to GP|21929218|dbj|BAC06183. 110kDa protein HM... 412 e-115
TC80443 homologue to GP|21929218|dbj|BAC06183. 110kDa protein HM... 208 9e-54
TC88497 similar to GP|11994617|dbj|BAB02754. contains similarity... 37 0.049
TC78529 similar to GP|14030876|gb|AAK52082.1 nuclease {Nicotiana... 34 0.32
TC92065 similar to GP|21539445|gb|AAM53275.1 putative copper ami... 31 2.7
TC86391 similar to GP|4097571|gb|AAD09514.1| GMFP5 {Glycine max}... 30 3.5
BQ751204 similar to PIR|T36948|T369 probable regulator - Strepto... 30 4.6
TC89551 similar to GP|17887381|gb|AAL40864.1 receptor protein ki... 30 4.6
>TC78191 homologue to GP|13111324|dbj|BAB32793. 110 kDa protein HMP {Pisum
sativum}, partial (84%)
Length = 2058
Score = 640 bits (1650), Expect = 0.0
Identities = 320/382 (83%), Positives = 346/382 (89%)
Frame = +1
Query: 608 KETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGTFLGSLWESR 667
K+ + FA SGVRCPGR EPYS+EAIALMRR+IM RDVEIEVETVDR GTFLGSLWESR
Sbjct: 637 KKLAVLLFAFSGVRCPGREEPYSDEAIALMRRRIMHRDVEIEVETVDRTGTFLGSLWESR 816
Query: 668 TNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVEGEEVSNGANV 727
N A+ LLEAGLAKLQTSFGSDRIP+ H+L++AEQSAK +KLKIWEN+VEGE V +GANV
Sbjct: 817 ANGAVPLLEAGLAKLQTSFGSDRIPDLHVLEQAEQSAKSKKLKIWENYVEGEVVPSGANV 996
Query: 728 ESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGAFSPKKGDIV 787
ESKQQEVLKV VTEVLGGGKFYVQTVGDQKIASIQ QLA+LNLK+APV+GAF+PKKGDIV
Sbjct: 997 ESKQQEVLKVTVTEVLGGGKFYVQTVGDQKIASIQNQLASLNLKDAPVIGAFNPKKGDIV 1176
Query: 788 LCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPG 847
LCYFHAD SWYRAMVVNTPRGPVES KD FEVFYIDYGNQE V YSQLRPLD SVSAAPG
Sbjct: 1177 LCYFHADSSWYRAMVVNTPRGPVESSKDAFEVFYIDYGNQEVVPYSQLRPLDPSVSAAPG 1356
Query: 848 LAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTI 907
LAQLCSLAYIK P+LEEDFGQEAAEYLSELTLSSGKEFRA VEE+DT+GGKVKGQGTG I
Sbjct: 1357 LAQLCSLAYIKLPNLEEDFGQEAAEYLSELTLSSGKEFRAMVEEKDTTGGKVKGQGTGPI 1536
Query: 908 LAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQ 967
+AVTLVAVD+EISVNAAMLQEGLARMEKRNRWDR RK LD+L+ FQ +AR RRGMWQ
Sbjct: 1537 IAVTLVAVDSEISVNAAMLQEGLARMEKRNRWDRTARKQALDNLEMFQGEARTARRGMWQ 1716
Query: 968 YGDVESDDEDTAPPARKAGAGR 989
YGD++SDDEDTAPP RKAG GR
Sbjct: 1717 YGDIQSDDEDTAPPQRKAGGGR 1782
Score = 283 bits (724), Expect = 2e-76
Identities = 144/192 (75%), Positives = 161/192 (83%)
Frame = +3
Query: 426 RDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGSVFL 485
+++ P P R K RL+GRQV+V+MEYSRK+ P DGSAVP A DSRVMDFGSVF+
Sbjct: 99 KNQLPMPVKR--KSS*EHRLIGRQVNVQMEYSRKVGPVDGSAVPPGAVDSRVMDFGSVFV 272
Query: 486 LSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYYDALLTAES 545
LS+ KAD DD PS A SQ TG+NV EL++GRGFGTVIRHRDFEERSN+YDALL AE+
Sbjct: 273 LSSGKADGDDAPSPAVPA-SQQTGLNVAELIIGRGFGTVIRHRDFEERSNFYDALLAAEA 449
Query: 546 RALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSGHRFKLL 605
RA+SGRKGIHSAKDPPVMHITDL T SAKKAKDFLPFLHRSRR+PAVVEYV SGHRFKLL
Sbjct: 450 RAISGRKGIHSAKDPPVMHITDLITASAKKAKDFLPFLHRSRRVPAVVEYVFSGHRFKLL 629
Query: 606 IPKETCSIAFAL 617
IPKETCSIAF +
Sbjct: 630 IPKETCSIAFCI 665
Score = 98.2 bits (243), Expect = 1e-20
Identities = 46/48 (95%), Positives = 47/48 (97%)
Frame = +2
Query: 396 ADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRT 443
ADDSIPYGSP AERRVNLSSIRCPK+GNPRRDEKPAPYAREAKEFLRT
Sbjct: 2 ADDSIPYGSPQAERRVNLSSIRCPKMGNPRRDEKPAPYAREAKEFLRT 145
Score = 52.4 bits (124), Expect = 9e-07
Identities = 43/164 (26%), Positives = 77/164 (46%), Gaps = 7/164 (4%)
Frame = +1
Query: 272 DPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPDGESAKDLALELVENGYAKY 331
+P+ +A R+++RDV I +E VD+ +GS++ ES + A+ L+E G AK
Sbjct: 694 EPYSDEAIALMRRRIMHRDVEIEVETVDRTGTFLGSLW----ESRANGAVPLLEAGLAK- 858
Query: 332 VEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNY-----VPPASNSKAIHNQNFTGKVVE 386
++ S L+ AE AK +L++W NY VP +N ++ + V E
Sbjct: 859 LQTSFGSDRIPDLHVLEQAEQSAKSKKLKIWENYVEGEVVPSGANVESKQQEVLKVTVTE 1038
Query: 387 VVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVG--NPRRDE 428
V+ G V + + + +L+ P +G NP++ +
Sbjct: 1039VLGGGKFYVQTVGDQKIASIQNQLASLNLKDAPVIGAFNPKKGD 1170
>TC90315 homologue to GP|21929218|dbj|BAC06183. 110kDa protein HMP {Pisum
sativum}, partial (62%)
Length = 719
Score = 412 bits (1058), Expect = e-115
Identities = 208/239 (87%), Positives = 225/239 (94%), Gaps = 1/239 (0%)
Frame = +3
Query: 124 KGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAAN 183
KGE SP+LAELLRLEEQAKQEGLGRWSKVPGAAEAS+RNLPPSA+GDASNFDAMGLLA N
Sbjct: 3 KGEASPFLAELLRLEEQAKQEGLGRWSKVPGAAEASVRNLPPSALGDASNFDAMGLLAKN 182
Query: 184 KGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADEN 243
KG PMEA+VEQVRDGSTLR+YLLPEFQFVQVFVAGIQ+PQMGRRAAPE+VV E+T D
Sbjct: 183 KGVPMEALVEQVRDGSTLRIYLLPEFQFVQVFVAGIQAPQMGRRAAPESVVVPEVTVDTT 362
Query: 244 DGDVPGEPRPPLTSAQRLAVS-SSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFS 302
+GDVP EPR PLTSAQRLAVS S+ ET+ADPFG DAKFFTEMRVLNRDVRIVLEGVDKFS
Sbjct: 363 NGDVPAEPRAPLTSAQRLAVSASAAETSADPFGADAKFFTEMRVLNRDVRIVLEGVDKFS 542
Query: 303 NLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRM 361
NLIGSVYYPDGESAKDLALELVENG+AKYVEWSANMME+EAK++LK AELEAKK+RLR+
Sbjct: 543 NLIGSVYYPDGESAKDLALELVENGFAKYVEWSANMMEDEAKKKLKAAELEAKKTRLRI 719
>TC80443 homologue to GP|21929218|dbj|BAC06183. 110kDa protein HMP {Pisum
sativum}, partial (34%)
Length = 617
Score = 208 bits (529), Expect = 9e-54
Identities = 99/136 (72%), Positives = 120/136 (87%), Gaps = 1/136 (0%)
Frame = +1
Query: 1 MASAATGATGWYRGRVKAVPSGDCLVIVAVASS-KPGPLPEKSITLASLITPRLARRGGV 59
MA+ A G + WY+ +VKAVPSGDC+V+V+VA++ K G LPEKSITL+SLI PRLARRGGV
Sbjct: 205 MAATAAGNSAWYKAKVKAVPSGDCIVVVSVAANAKLGVLPEKSITLSSLIAPRLARRGGV 384
Query: 60 DEPFAWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGEKNVGVLVVSQGWAKVRE 119
DEPFAWESRE+LRKL IGKE+TFR+DY V SINR+FGTVFLG+KNV +LVVSQGWAKVRE
Sbjct: 385 DEPFAWESREFLRKLLIGKEITFRIDYTVPSINREFGTVFLGDKNVALLVVSQGWAKVRE 564
Query: 120 QGQQKGEVSPYLAELL 135
QGQQKG+ SP+ +++
Sbjct: 565 QGQQKGKASPFFGKIV 612
Score = 48.5 bits (114), Expect = 1e-05
Identities = 36/109 (33%), Positives = 58/109 (53%), Gaps = 3/109 (2%)
Frame = +1
Query: 380 FTGKVVEVVSGDCIIVADDSIPYGSPLA---ERRVNLSSIRCPKVGNPRRDEKPAPYARE 436
+ KV V SGDCI+V S+ + L E+ + LSS+ P++ RR P+A E
Sbjct: 238 YKAKVKAVPSGDCIVVV--SVAANAKLGVLPEKSITLSSLIAPRLA--RRGGVDEPFAWE 405
Query: 437 AKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGSVFL 485
++EFLR L+G++++ ++Y+ VPS +FG+VFL
Sbjct: 406 SREFLRKLLIGKEITFRIDYT----------VPSIN-----REFGTVFL 507
>TC88497 similar to GP|11994617|dbj|BAB02754. contains similarity to kinesin
protein~gene_id:MGL6.9 {Arabidopsis thaliana}, partial
(26%)
Length = 1310
Score = 36.6 bits (83), Expect = 0.049
Identities = 26/82 (31%), Positives = 34/82 (40%), Gaps = 14/82 (17%)
Frame = +2
Query: 420 KVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEY-----------SRKIVPTDGSAV 468
K GNPR+D+ P P + KE L T L E Y RK++ + S
Sbjct: 284 KSGNPRKDQAPNPVPQSNKEVLSTSSLPDSACAEDVYYQRQEVKTGDMGRKVIENENSLY 463
Query: 469 PSPAA---DSRVMDFGSVFLLS 487
S AA D + F S FL +
Sbjct: 464 SSAAAADVDKQPSSFSSTFLFN 529
>TC78529 similar to GP|14030876|gb|AAK52082.1 nuclease {Nicotiana tabacum},
partial (56%)
Length = 618
Score = 33.9 bits (76), Expect = 0.32
Identities = 23/92 (25%), Positives = 46/92 (50%)
Frame = +3
Query: 271 ADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPDGESAKDLALELVENGYAK 330
A P+G +AK V + +R+++ G D++ L+G +Y + ++L L+ K
Sbjct: 372 AMPYGKEAKTELTKIVQGKSLRVLIYGEDQYQRLVGDIYC-NNIFVQELMLK-------K 527
Query: 331 YVEWSANMMEEEAKRRLKTAELEAKKSRLRMW 362
+ W + + + L+T E EA+ R+ +W
Sbjct: 528 GLAW--HYAAYDKRPELETWEKEARAKRVGLW 617
>TC92065 similar to GP|21539445|gb|AAM53275.1 putative copper amine oxidase
{Arabidopsis thaliana}, partial (2%)
Length = 1396
Score = 30.8 bits (68), Expect = 2.7
Identities = 19/62 (30%), Positives = 33/62 (52%)
Frame = -1
Query: 402 YGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIV 461
Y L+ERR +S RC + PRR + +R ++ + R+RL R + + SR+ +
Sbjct: 1000 YPMALSERRCRVS*PRCSRSCRPRR-QSALSTSRPSRTYTRSRLR*RASTCSLTVSRQHL 824
Query: 462 PT 463
P+
Sbjct: 823 PS 818
>TC86391 similar to GP|4097571|gb|AAD09514.1| GMFP5 {Glycine max}, partial
(54%)
Length = 1353
Score = 30.4 bits (67), Expect = 3.5
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = +3
Query: 932 RMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGD 970
R E++ RW RKER+ G + + R+E+ G W+ G+
Sbjct: 858 REERKGRWRRKEREEG-------R*EERREKSGRWRCGE 953
>BQ751204 similar to PIR|T36948|T369 probable regulator - Streptomyces
coelicolor, partial (9%)
Length = 715
Score = 30.0 bits (66), Expect = 4.6
Identities = 16/34 (47%), Positives = 19/34 (55%), Gaps = 2/34 (5%)
Frame = +2
Query: 248 PGEPRPPLTSAQR--LAVSSSTETAADPFGPDAK 279
PG PRPP TS R L+V S T T+ P A+
Sbjct: 233 PGSPRPPTTSGMRPSLSVGSPTTTSPTLLRPTAR 334
>TC89551 similar to GP|17887381|gb|AAL40864.1 receptor protein kinase-like
protein {Capsicum annuum}, partial (63%)
Length = 1291
Score = 30.0 bits (66), Expect = 4.6
Identities = 17/47 (36%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Frame = +2
Query: 527 HRDFEERSNYYDALLT-AESRALSGRKGIHSAKDPPVMHITDLTTTS 572
H + +SNYYDA+L E +S KG + +PP + DL S
Sbjct: 149 HPNTASKSNYYDAILNGVEIFKISDSKGNLAGTNPPPPQVQDLIDPS 289
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.133 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,564,425
Number of Sequences: 36976
Number of extensions: 295652
Number of successful extensions: 1087
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1074
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1083
length of query: 990
length of database: 9,014,727
effective HSP length: 106
effective length of query: 884
effective length of database: 5,095,271
effective search space: 4504219564
effective search space used: 4504219564
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0094a.6


