

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.9
(742 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
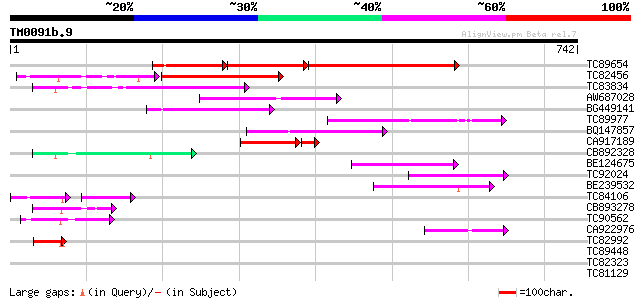
Score E
Sequences producing significant alignments: (bits) Value
TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein... 159 5e-88
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 167 2e-50
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 149 4e-36
AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~... 143 3e-34
BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidop... 125 5e-29
TC89977 similar to GP|5764395|gb|AAD51282.1| far-red impaired re... 112 7e-25
BQ147857 weakly similar to GP|15983442|gb At1g10240/F14N23_12 {A... 102 5e-22
CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.3... 79 4e-17
CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sin... 72 6e-13
BE124675 weakly similar to GP|13365573|db hypothetical protein~s... 67 2e-11
TC92024 similar to PIR|T05645|T05645 hypothetical protein F20D10... 66 6e-11
BE239532 64 2e-10
TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity ... 40 2e-07
CB893278 weakly similar to PIR|T05645|T056 hypothetical protein ... 52 6e-07
TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 49 9e-06
CA922976 weakly similar to GP|9502168|gb|A contains simlarity to... 44 2e-04
TC82992 homologue to PIR|T05645|T05645 hypothetical protein F20D... 44 3e-04
TC89448 similar to PIR|C84864|C84864 hypothetical protein At2g43... 40 0.004
TC82323 similar to GP|7673677|gb|AAF66982.1| transposase {Zea ma... 39 0.006
TC81129 similar to GP|5764395|gb|AAD51282.1| far-red impaired re... 39 0.009
>TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein F20D10.290
- Arabidopsis thaliana, partial (65%)
Length = 1358
Score = 159 bits (403), Expect(3) = 5e-88
Identities = 77/197 (39%), Positives = 121/197 (61%)
Frame = +1
Query: 392 NEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQEFVLKFEKA 451
NEW Q +Y R+ W+P+Y R +FF + +E +N FFD V+ASTTLQ V ++EKA
Sbjct: 697 NEWPQSMYNARQHWVPVYLRGSFFGEIPLNDGNECLNFFFDGHVNASTTLQLLVRQYEKA 876
Query: 452 VESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKIR 511
V + E E K ++E+ + + +L T S +E+ AAS+YTR +F KF+ EL KI
Sbjct: 877 VSTWHERELKADFETSNSNPVLRTPSPMEKQAASLYTRKMFMKFQEELVGTMANPATKIE 1056
Query: 512 RDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQI 571
G Y+V+ + + + TV + A C +++E+ GI+CRHIL +F+AK V+ +
Sbjct: 1057DSGTITTYRVAKFGENQTSHTVAFNSSEMKASCSCQMFEYSGIVCRHILAVFRAKNVLTL 1236
Query: 572 PSHFIMERWTKDANRGL 588
PSH++++RWT +A G+
Sbjct: 1237PSHYVLKRWTMNAKTGV 1287
Score = 105 bits (263), Expect(3) = 5e-88
Identities = 48/98 (48%), Positives = 64/98 (64%)
Frame = +2
Query: 187 YIRNVRRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQL 246
++ + R++ L G Q V DY KR + EN FFYA+Q D D+ N+FW D R +Y
Sbjct: 83 HMSSTRQRTLGGGGHQ-VLDYLKRMRAENHAFFYAVQSDVDNAGGNIFWADETCRTNYSY 259
Query: 247 FGDVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCA 284
FGD + FDTTYKTN+Y +PFA F G N+H Q +L+GCA
Sbjct: 260 FGDTVIFDTTYKTNQYRVPFASFTGFNHHGQPVLFGCA 373
Score = 99.8 bits (247), Expect(3) = 5e-88
Identities = 43/107 (40%), Positives = 67/107 (62%)
Frame = +3
Query: 285 LLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKVFPESRHRLCLWHIIKK 344
L+ +ESE S++WLFKT L+A+ G+ P+SI TD D ++ A+A+V P +RHR C W I ++
Sbjct: 375 LILNESEPSYIWLFKTWLRAVSGRPPVSITTDLDPVIQVAVAQVLPPTRHRFCKWSIFRE 554
Query: 345 FPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRIMVEYNLKE 391
KLAH+Y F + K+C+ S +I FE W ++ Y + +
Sbjct: 555 NRSKLAHLYQSNPTFDNEFKKCVHESVTIDEFESCWRSLLERYYIMD 695
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 167 bits (424), Expect(2) = 2e-50
Identities = 77/160 (48%), Positives = 103/160 (64%)
Frame = +2
Query: 199 GDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYK 258
GD A+ + R Q +NS F+ A+ D+ + NLFW DAR R +Y+ FG+VIT DTTY
Sbjct: 713 GDVDAIHSFFHRMQKQNSQFYCAMDMDDKRNIQNLFWADARCRAAYEYFGEVITLDTTYL 892
Query: 259 TNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQD 318
T+KY +P PFVG+N+H Q+IL GCA+L + + WLF L+ M G P IIT++D
Sbjct: 893 TSKYDLPLVPFVGVNHHGQTILLGCAILSNLDAKTLTWLFTRWLECMHGHAPNGIITEED 1072
Query: 319 LAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSI 358
AMK A+ FP++RHR CLWHI+KK PE L +SI
Sbjct: 1073KAMKNAIEVAFPKARHRWCLWHIMKKVPEMLGKYSRNESI 1192
Score = 50.4 bits (119), Expect(2) = 2e-50
Identities = 51/197 (25%), Positives = 87/197 (43%), Gaps = 9/197 (4%)
Frame = +1
Query: 9 SSKDFTTSILDTSDIGGNSKLEIGMEFESIEKVREFYNSFAKKNGFGVRVRSSK----PK 64
S F+ +I+ + D + GM F S E+V +Y ++A + GFGV SSK K
Sbjct: 160 SDDKFSAAIVVSED----EEPRTGMTFRSEEEVIRYYMNYANRVGFGVTKISSKNADDGK 327
Query: 65 RAVLVCCNEGQHKVKISRTEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSF 124
+ + CN + V S+ Q T++T+ R+ A + + +T S+ ++
Sbjct: 328 KYFTLACNCARRYVSTSKNPSKQYLTSKTQ-----CRARLNACVALDGSSTISRIVL--- 483
Query: 125 NNDHNHVMVSPKSVSYMRCHKKMSVAAKSLVEKFEEEGIPTGK----VAAIFND-GDSTF 179
+HNH + K Y R + + +E ++ GI + V N G+ T
Sbjct: 484 --EHNHELGQTKG-RYFRLDESSGTHVERNLELNDQGGINVDRNLQSVGLEMNGCGNLTS 654
Query: 180 TNRDCWNYIRNVRRKNL 196
DC N+++ VRR L
Sbjct: 655 GENDCRNFVQKVRRLRL 705
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 149 bits (376), Expect = 4e-36
Identities = 95/293 (32%), Positives = 149/293 (50%), Gaps = 10/293 (3%)
Frame = +3
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRV------RSSKPKRAVLVCCNEGQHKVKISRTE 84
+ MEFES + +Y +AK+ GF VRV R+SK K ++CC+ K RT+
Sbjct: 219 LAMEFESYDDAYSYYICYAKEVGFCVRVKNSWFKRNSKEKYGAVLCCSSQGFK----RTK 386
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCH 144
++ + +T R+GC A +I + +W I +HNHV+ + + H
Sbjct: 387 DVNNLRKET-------RTGCPA-MIRMKLVESQRWRICEVTLEHNHVLGA-------KIH 521
Query: 145 KKMSVAAKSLVEKFEEEG--IPTGKVAAIFNDGDSTFTN--RDCWNYIRNVRRKNLDVGD 200
K + K+ + + EG I I +G+ + RD + + + NL GD
Sbjct: 522 KSIK---KNSLPSSDAEGKTIKVYHALVIDTEGNDNLNSNARDDRAFSKYSNKLNLRKGD 692
Query: 201 AQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTN 260
QA++++ R Q+ N NFFY + +++ + N WVDA+SR + F DVI FD TY N
Sbjct: 693 TQAIYNFLCRMQLTNPNFFYLMDFNDEGHLRNALWVDAKSRAACGYFSDVIYFDNTYLVN 872
Query: 261 KYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISI 313
KY +P VG+N+H QS+L GC LL E S+ WLF+T ++ + P +I
Sbjct: 873 KYEIPLVALVGINHHGQSVLLGCGLLAGEIIESYKWLFRTWIKCIPRCSPQTI 1031
>AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~similar to
Arabidopsis thaliana F20D10.300, partial (11%)
Length = 662
Score = 143 bits (360), Expect = 3e-34
Identities = 67/186 (36%), Positives = 109/186 (58%)
Frame = +1
Query: 249 DVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGK 308
DV+ FDTTYK NKY+ P F G N+HSQ+I++G ALL+DE+ S+ W+ L+ M K
Sbjct: 1 DVLAFDTTYKKNKYNYPLCIFSGCNHHSQTIIFGVALLEDETIESYKWVLNRFLECMENK 180
Query: 309 KPISIITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIR 368
P +++TD D +M+ A+ +VFP++ HRLC WH+ K E + K++ F ++ +
Sbjct: 181 FPKAVVTDGDGSMREAIKQVFPDASHRLCAWHLHKNAQENI-----KKTPFLEGFRKAMY 345
Query: 369 GSHSIQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESIN 428
+ + + FE+ W ++ + L+ N W+ Y + W Y R FF + TT + E+IN
Sbjct: 346 SNFTPEQFEDFWSELIQKNELEGNAWVIKTYANKSLWATAYLRDKFFGRIRTTSQCEAIN 525
Query: 429 AFFDSF 434
A ++
Sbjct: 526 AIVKTY 543
>BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (33%)
Length = 690
Score = 125 bits (315), Expect = 5e-29
Identities = 58/168 (34%), Positives = 97/168 (57%)
Frame = +3
Query: 179 FTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDA 238
FT +D N +++ R+ + + + + C+ + ++ NF + D ++R+ N+ W A
Sbjct: 171 FTEKDVRNLLQSFRKLDPEE-ETLDLLRMCRNIKDKDPNFKFEYTLDANNRLENIAWSYA 347
Query: 239 RSRLSYQLFGDVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLF 298
S Y +FGD + FDTT++ + MP +VG+NN+ +GC LL+DE+ SF W
Sbjct: 348 SSIQLYDIFGDAVVFDTTHRLTAFDMPLGLWVGINNYGMPCFFGCVLLRDETVRSFSWAI 527
Query: 299 KTLLQAMGGKKPISIITDQDLAMKAALAKVFPESRHRLCLWHIIKKFP 346
K L M GK P +I+TDQ++ +K AL+ P ++H C+W I+ KFP
Sbjct: 528 KAFLGFMNGKAPQTILTDQNICLKEALSAEMPMTKHAFCIWMIVAKFP 671
>TC89977 similar to GP|5764395|gb|AAD51282.1| far-red impaired response
protein {Arabidopsis thaliana}, partial (35%)
Length = 1523
Score = 112 bits (279), Expect = 7e-25
Identities = 73/235 (31%), Positives = 119/235 (50%)
Frame = +2
Query: 416 AGMNTTQRSESINAFFDSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILST 475
AG +T QRSES+N+FFD ++ TL+EFV ++ + +R + E ++++ HK L +
Sbjct: 2 AGTSTAQRSESMNSFFDKYIHKKITLKEFVKQYRLILLNRYDEEEIADFDTLHKQPALKS 181
Query: 476 GSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDI 535
S E+ ++IYT IF KF+ E+ + DG Y V + Y+ + F V
Sbjct: 182 PSPWEKQMSTIYTHAIFKKFQVEVLGVAGCQSRIEVGDGTAVRYIVQD-YEKDEEFLVTW 358
Query: 536 DLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDN 595
S C L+E+ G LCRH L + Q G +PSH+IM+RWTKDA + + D
Sbjct: 359 KELSSEVSCFCRLFEYKGFLCRHALSVLQRCGCSSVPSHYIMKRWTKDAK--IREPAADR 532
Query: 596 DLGEKSDTLKILRRVHVQREASFLADLAEESEEAYNFIISEMKQTHKSAAAMKTS 650
++ T + R + + + L++ SEE+YN + + K+ + S
Sbjct: 533 T--QRIQT-RPQRYNDLCKRSIALSEEGSSSEESYNIALRALIDALKNCVLVNNS 688
>BQ147857 weakly similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (26%)
Length = 560
Score = 102 bits (254), Expect = 5e-22
Identities = 57/186 (30%), Positives = 96/186 (50%), Gaps = 1/186 (0%)
Frame = +1
Query: 310 PISIITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQ-SIFKRDMKRCIR 368
P +++TD + +K A+A PES+H C+WHI+ KF + + Q +K D R +
Sbjct: 4 PQTLLTDHNTWLKEAIAVEMPESKHAFCIWHILSKFSDWXYLLLGSQYDEWKADFHR-LY 180
Query: 369 GSHSIQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESIN 428
+ FE+ W +++ +Y L N+ + LY +R W + R FFAG+ + ++ESIN
Sbjct: 181 NLEMXEDFEKSWRQMVDKYGLHANKHIISLYSLRTFWALPFLRRYFFAGLTSXCQTESIN 360
Query: 429 AFFDSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYT 488
F F+ A + F+ + V+ A K+ + + + L TGS +E HAA+I T
Sbjct: 361 VFIQRFLSAQSQPXRFLQQVADIVDFNDRAGAKQXMQRKMQKXCLKTGSPIESHAATILT 540
Query: 489 RNIFAK 494
+K
Sbjct: 541 PYALSK 558
>CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (12%)
Length = 312
Score = 79.3 bits (194), Expect(2) = 4e-17
Identities = 34/79 (43%), Positives = 52/79 (65%)
Frame = +1
Query: 302 LQAMGGKKPISIITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKR 361
L AM G+ P+SI TD D +++A+ +VFP++RHR C WHI K+ EKL+HI+ + F+
Sbjct: 1 LTAMSGRPPLSITTDHDSVIQSAIMQVFPDTRHRFCKWHIFKQCQEKLSHIFLQFPNFEA 180
Query: 362 DMKRCIRGSHSIQSFEEEW 380
+ +C+ + SI FE W
Sbjct: 181 EFHKCVNLTDSIDEFESCW 237
Score = 27.3 bits (59), Expect(2) = 4e-17
Identities = 8/23 (34%), Positives = 15/23 (64%)
Frame = +3
Query: 383 IMVEYNLKENEWLQGLYKIRESW 405
++ Y+L++NEWLQ ++ W
Sbjct: 243 LLDRYDLRDNEWLQAIHSACRQW 311
>CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sinorhizobium
meliloti}, partial (1%)
Length = 828
Score = 72.4 bits (176), Expect = 6e-13
Identities = 52/227 (22%), Positives = 92/227 (39%), Gaps = 13/227 (5%)
Frame = +1
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRV--------RSSKPKRAVLVCCNEGQHKVKISR 82
+G S ++ + Y A K GF VR K + C +G
Sbjct: 160 LGQVVNSKDEAYDLYQEHAFKTGFSVRKGKELYYDNEKKKTRLKDFYCSKQGFKN----- 324
Query: 83 TEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMR 142
+ + K + R+ C A ++ T + W + DHNH V + +R
Sbjct: 325 ----NEPDGEVAYKRADSRTNCLA-MVRFNVTKEGVWKVTKLILDHNHEFVPLQQRYLLR 489
Query: 143 CHKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNR-----DCWNYIRNVRRKNLD 197
+ MS + ++ + I V R D NY+ + K ++
Sbjct: 490 SMRNMSNLKEDRIKSLVNDAIRVTNVGGYLGKEVGGVDKRGVMLKDMHNYVSTGKLKFIE 669
Query: 198 VGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSY 244
GDAQ++ ++ + KQ ++S F+Y++Q D +SR+ N+FW D +S + Y
Sbjct: 670 AGDAQSLLNHLQSKQAQDSMFYYSVQLDHESRLNNVFWRDGKSVVDY 810
>BE124675 weakly similar to GP|13365573|db hypothetical protein~similar to
Arabidopsis thaliana F20D10.300, partial (4%)
Length = 448
Score = 67.0 bits (162), Expect = 2e-11
Identities = 41/140 (29%), Positives = 65/140 (46%)
Frame = +2
Query: 448 FEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTR 507
FE+ VE + E + + E + L + +HA+ YT F F+ EK
Sbjct: 2 FERIVEEQRYKEIEASDEMKGCLPKLMGNVVVLKHASVAYTPRAFEVFQQRYEKSLNVIV 181
Query: 508 HKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKG 567
++ +RDG + Y+V N Y +TV C +E +G LC H L + +
Sbjct: 182 NQHKRDGYLFEYKV-NTYGHARQYTVTFSSSDNTVVCSCMKFEHVGFLCSHALKVLDNRN 358
Query: 568 VVQIPSHFIMERWTKDANRG 587
+ +PS +I++RWTKDA G
Sbjct: 359 IKVVPSRYILKRWTKDARLG 418
>TC92024 similar to PIR|T05645|T05645 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (11%)
Length = 938
Score = 65.9 bits (159), Expect = 6e-11
Identities = 35/130 (26%), Positives = 62/130 (46%)
Frame = +3
Query: 523 NCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTK 582
N + + V ++ A C +++EF G+LCRHIL +F+ V+ +PSH+I++RWTK
Sbjct: 30 NLGEDHKAYNVRFNVLEMKATCSCQMFEFSGLLCRHILAVFRVTNVLTLPSHYILKRWTK 209
Query: 583 DANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAYNFIISEMKQTHK 642
+A + + + +R ++ EA D S E Y+ +++ K
Sbjct: 210 NAKSNVSLQEHSSHAYTYYLESHTVRYNTLRHEAFKFVDKGASSPETYDVAKDALQEAAK 389
Query: 643 SAAAMKTSEG 652
A + EG
Sbjct: 390 RVAQVMRKEG 419
>BE239532
Length = 645
Score = 63.9 bits (154), Expect = 2e-10
Identities = 44/167 (26%), Positives = 76/167 (45%), Gaps = 9/167 (5%)
Frame = +3
Query: 477 SKLEEHAASIYTRNIFAKFKGE-LEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDI 535
S+ HA+++YT F F E LE T +I + ++V + + V
Sbjct: 36 SETLRHASTVYTHKGFRIFLNEYLEGTGGSTSIEISQSNNLSNHEVMLNHKPNKKYAVTF 215
Query: 536 DLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDAN--------RG 587
D + C ++ MGILC H L I+ KG+++IP + ++RW+K+A RG
Sbjct: 216 DSSNMKINCSCRKFDSMGILCSHALRIYNIKGILRIPDQYFLKRWSKNARSVIYEHMPRG 395
Query: 588 LEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAYNFII 634
+ + N + D IL R + +L +++S+EA N +
Sbjct: 396 MGEDSTSNSV-INDDDAGILYRNAIMTSFYYLVLESQDSKEAQNIYV 533
>TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity to
Arabidopsis thaliana far-red impaired response protein
(GB:AAD51282.1), partial (25%)
Length = 673
Score = 39.7 bits (91), Expect(2) = 2e-07
Identities = 18/72 (25%), Positives = 36/72 (50%), Gaps = 2/72 (2%)
Frame = +2
Query: 95 RKCSTIRSGCEASLIVSRGTTK--SKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVAAK 152
R ++R GC+A + +S+ S+W + F+N HNH ++ V + ++K+ A +
Sbjct: 395 RDRKSVRCGCDAKMYLSKDVVDGVSQWTVLQFSNVHNHELLEDDQVRLLPAYRKIHEADQ 574
Query: 153 SLVEKFEEEGIP 164
+ + G P
Sbjct: 575 ERILLLSKAGFP 610
Score = 33.9 bits (76), Expect(2) = 2e-07
Identities = 27/84 (32%), Positives = 42/84 (49%), Gaps = 6/84 (7%)
Frame = +1
Query: 2 DDNGIGSSSKDFTTSILDTSDIGGNSKLEIGMEFESIEKVREFYNSFAKKNGFGVRVRSS 61
DD+ GS++ + + SDI +GM F+S ++ E Y +FA+K+GF +R S
Sbjct: 127 DDSDEGSAAPESMKADKTPSDI---ITPYVGMVFKSDDEAFEHYGNFARKHGFSMRKARS 297
Query: 62 K--PKRAV----LVCCNEGQHKVK 79
+ P+ V VC G VK
Sbjct: 298 RISPQLGVYKRDFVCYRSGFAPVK 369
>CB893278 weakly similar to PIR|T05645|T056 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (5%)
Length = 821
Score = 52.4 bits (124), Expect = 6e-07
Identities = 33/118 (27%), Positives = 57/118 (47%), Gaps = 8/118 (6%)
Frame = +3
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRA-------VLVCCNEG-QHKVKISR 82
IGM+F S E+ R FY+ + ++ GF VR+ ++ R VC EG + K + R
Sbjct: 321 IGMQFNSREEARGFYDGYGRRIGFTVRIHHNRRSRVNNELIGQDFVCSKEGFRAKKYVHR 500
Query: 83 TEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSY 140
+ + T R GC+A + ++ + KW++ F +H H ++SP V +
Sbjct: 501 KDRVLPPPPAT-------REGCQAMIRLAL-RDEGKWVVTKFVKEHTHKLMSPGEVPW 650
>TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (66%)
Length = 1169
Score = 48.5 bits (114), Expect = 9e-06
Identities = 40/130 (30%), Positives = 58/130 (43%), Gaps = 7/130 (5%)
Frame = +2
Query: 15 TSILDTSDIGGNSKLEIGMEFESIEKVREFYNSFAKKNGFGVRV-RSSKPKR------AV 67
TS++ T D + IG EF S + FYNS+A + GF +RV + S+ +R
Sbjct: 296 TSVMRTID-----EPYIGQEFVSEAEAHAFYNSYATRVGFVIRVSKLSRSRRDGSVIGRA 460
Query: 68 LVCCNEGQHKVKISRTEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNND 127
LVC EG + D + R+ + R GC A ++V R W I F +
Sbjct: 461 LVCNKEGFR---------MPDKREKIVRQRAETRVGCRAMIMV-RKLNSGLWSITKFVKE 610
Query: 128 HNHVMVSPKS 137
H H + KS
Sbjct: 611 HTHPLTPGKS 640
>CA922976 weakly similar to GP|9502168|gb|A contains simlarity to Arabidopsis
thaliana far-red impaired response protein
(GB:AAD51282.1), partial (11%)
Length = 827
Score = 43.9 bits (102), Expect = 2e-04
Identities = 29/112 (25%), Positives = 49/112 (42%), Gaps = 2/112 (1%)
Frame = -3
Query: 544 CGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDT 603
C + +E GILCRH L I K Q+P + + RW ++ + +ED +N+ +
Sbjct: 702 CSCKEFESSGILCRHALRILVTKNYFQLPEKYYLSRWHRERSLIVEDEHNNQSFIAE--- 532
Query: 604 LKILRRVHVQREASFLAD--LAEESEEAYNFIISEMKQTHKSAAAMKTSEGV 653
K + Q E F E S+ A ++ E+ + M ++GV
Sbjct: 531 -KWFQEYQSQAENLFSESSITKERSDYARKELMKELTRILNEVRDMSETDGV 379
>TC82992 homologue to PIR|T05645|T05645 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (7%)
Length = 354
Score = 43.5 bits (101), Expect = 3e-04
Identities = 24/50 (48%), Positives = 31/50 (62%), Gaps = 7/50 (14%)
Frame = +1
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKR---AVL----VCCNEG 74
GMEFES E + FYNS+A++ GF RV SS+ R A++ VC EG
Sbjct: 202 GMEFESEEAAKAFYNSYARRVGFSTRVSSSRRSRRDGAIIQRQXVCAKEG 351
>TC89448 similar to PIR|C84864|C84864 hypothetical protein At2g43280
[imported] - Arabidopsis thaliana, partial (80%)
Length = 1177
Score = 39.7 bits (91), Expect = 0.004
Identities = 32/99 (32%), Positives = 47/99 (47%), Gaps = 6/99 (6%)
Frame = +1
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRV----RSSKPKR--AVLVCCNEGQHKVKISRTEE 85
GMEF S + + FY+ +A+ GF +RV RS + R A CN+ H V
Sbjct: 310 GMEFVSEDAAKIFYDEYARHVGFVMRVMSCRRSXRDGRILARRFGCNKEGHCV------S 471
Query: 86 IQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSF 124
IQ ++ ++ R GC+A +I + KWMI F
Sbjct: 472 IQGKLGPVRKPRASTREGCKA-MIHIKYDQSGKWMITXF 585
>TC82323 similar to GP|7673677|gb|AAF66982.1| transposase {Zea mays},
partial (2%)
Length = 1165
Score = 39.3 bits (90), Expect = 0.006
Identities = 16/42 (38%), Positives = 24/42 (57%)
Frame = +1
Query: 543 KCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDA 584
+C L+E GI C H+ + + + V IP +M RWTK+A
Sbjct: 28 QCSCRLFESRGIPCCHVFFVMKEEDVDHIPKCLVMTRWTKNA 153
>TC81129 similar to GP|5764395|gb|AAD51282.1| far-red impaired response
protein {Arabidopsis thaliana}, partial (13%)
Length = 721
Score = 38.5 bits (88), Expect = 0.009
Identities = 25/93 (26%), Positives = 43/93 (45%)
Frame = +2
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDSTN 91
G+EFE+ E FY +AK GF +++S+ + + K SR +S
Sbjct: 422 GIEFETHEAAYSFYQEYAKSMGFTTSIKNSRRSKKTKEFIDA---KFACSRYGVTPESDG 592
Query: 92 QTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSF 124
++ ++ S ++ C+A + V R KW I F
Sbjct: 593 RSSQRSSVKKTDCKACMHVKR-NPDGKWNIFEF 688
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,608,427
Number of Sequences: 36976
Number of extensions: 309017
Number of successful extensions: 1592
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 1560
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1584
length of query: 742
length of database: 9,014,727
effective HSP length: 103
effective length of query: 639
effective length of database: 5,206,199
effective search space: 3326761161
effective search space used: 3326761161
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0091b.9


