

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.14
(487 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
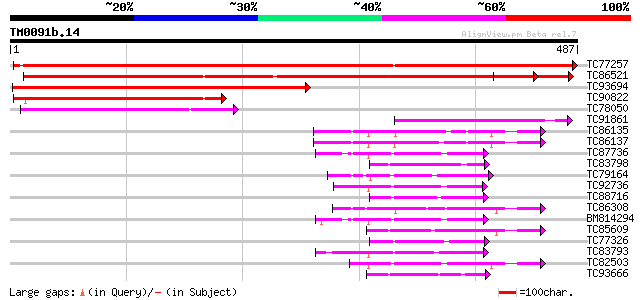
Score E
Sequences producing significant alignments: (bits) Value
TC77257 weakly similar to GP|14134199|gb|AAK54282.1 fatty acid 9... 752 0.0
TC86521 similar to GP|20160362|emb|CAD29735. allene oxide syntha... 459 e-130
TC93694 weakly similar to GP|14134199|gb|AAK54282.1 fatty acid 9... 345 3e-95
TC90822 similar to PIR|A56377|A56377 rubber particle cytochrome ... 182 2e-46
TC78050 homologue to GP|5830469|emb|CAB54849.1 hydroperoxide lya... 132 2e-31
TC91861 homologue to GP|5830467|emb|CAB54848.1 hydroperoxide lya... 122 2e-28
TC86135 similar to PIR|A96673|A96673 probable cytochrome P450 F1... 55 8e-08
TC86137 weakly similar to PIR|A96673|A96673 probable cytochrome ... 54 2e-07
TC87736 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P... 51 9e-07
TC83798 weakly similar to GP|10177750|dbj|BAB11063. cytochrome P... 51 1e-06
TC79164 similar to SP|O48928|C773_SOYBN Cytochrome P450 77A3 (EC... 49 3e-06
TC92736 weakly similar to SP|Q9STK9|C71O_ARATH Cytochrome P450 7... 48 7e-06
TC88716 weakly similar to GP|20258812|gb|AAM14017.1 putative cyt... 48 7e-06
TC86308 similar to SP|O48927|CP78_SOYBN Cytochrome P450 78A3 (EC... 48 7e-06
BM814294 weakly similar to GP|6002279|emb cytochrome P450 monoox... 48 1e-05
TC85609 weakly similar to SP|Q9LTL2|C72P_ARATH Cytochrome P450 7... 48 1e-05
TC77326 similar to GP|12004682|gb|AAG44132.1 cytochrome P450 {Pi... 47 1e-05
TC83793 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P... 47 2e-05
TC82503 weakly similar to GP|18252325|gb|AAL66194.1 cytochrome P... 46 4e-05
TC93666 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 7... 45 5e-05
>TC77257 weakly similar to GP|14134199|gb|AAK54282.1 fatty acid
9-hydroperoxide lyase {Cucumis melo}, partial (81%)
Length = 1989
Score = 752 bits (1941), Expect = 0.0
Identities = 365/484 (75%), Positives = 414/484 (85%)
Frame = +2
Query: 4 ASSDDTKNSSSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQGRDKFFATRVEKYNSTVIRT 63
ASS +T SS+NLPLKPIPGSYGLP G +HDRHDYFYNQGRDK+F TR+EKYNSTV++
Sbjct: 110 ASSSET--SSTNLPLKPIPGSYGLPIIGPLHDRHDYFYNQGRDKYFQTRIEKYNSTVLKL 283
Query: 64 NMPPGPGISSDSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDT 123
NMPPG I+ D KVIALLD ASFPILFDN+KVEKRDVLDGTFMPST F GGYR CAFQDT
Sbjct: 284 NMPPGGFIAPDPKVIALLDGASFPILFDNAKVEKRDVLDGTFMPSTDFFGGYRTCAFQDT 463
Query: 124 TEPSHKLLKTFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATF 183
EPSH LLK F +LSSKH+TF+PL +T L++HF+DL+ +LAGK KASFNTSI TF
Sbjct: 464 AEPSHSLLKRFIFHILSSKHDTFIPLFQTNLTEHFTDLEKELAGKHQKASFNTSIGGITF 643
Query: 184 NFLFLLLTDKHPSETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPFPA 243
NFLF L+TDK+PSET IGD GP LV WL AQLAPLAT GLPKIFNY ED LIRTIP PA
Sbjct: 644 NFLFKLITDKNPSETKIGDSGPTLVQTWLAAQLAPLATAGLPKIFNYLEDVLIRTIPIPA 823
Query: 244 WTVKWSYKKLCEGFASAGASLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIK 303
WTVK SY KL EG AG ++LDEAE++GIKR+EA HNLVF LGFNAFGGLTNQFPILIK
Sbjct: 824 WTVKSSYNKLYEGLMEAGTTVLDEAEKMGIKREEACHNLVFTLGFNAFGGLTNQFPILIK 1003
Query: 304 WIGLAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKA 363
W+GLAG +LH KLA+EIR +VRE G V+L ALDKMTLTKS VYE LRIEPAVPYQYAKA
Sbjct: 1004WVGLAGADLHKKLADEIRAIVREEGG-VNLYALDKMTLTKSTVYEALRIEPAVPYQYAKA 1180
Query: 364 REDVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYWS 423
RED+VV+SHDA+FEIKKGEMIFGYQPFATKD K+F+ PE+F+ RF+G+GEKLLK+V+WS
Sbjct: 1181REDLVVQSHDASFEIKKGEMIFGYQPFATKDAKIFDKPEDFIAERFIGDGEKLLKHVFWS 1360
Query: 424 NGRETEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKA 483
NGRET+E T DNK CPAKNLVVL+CRLY+VEFFL YDTFTF+FKP VLGP +TI+SL KA
Sbjct: 1361NGRETDEATPDNKICPAKNLVVLLCRLYLVEFFLNYDTFTFDFKPSVLGPTITIKSLVKA 1540
Query: 484 TTTV 487
++TV
Sbjct: 1541SSTV 1552
>TC86521 similar to GP|20160362|emb|CAD29735. allene oxide synthase {Solanum
tuberosum}, partial (84%)
Length = 2196
Score = 459 bits (1182), Expect = e-130
Identities = 227/473 (47%), Positives = 317/473 (66%), Gaps = 1/473 (0%)
Frame = +1
Query: 13 SSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQGRDKFFATRVEKYNSTVIRTNMPPGPGIS 72
++ LP++ IPG YG+PF DR DYFYNQGRD++F +R++KY ST+ RTN+PPGP I+
Sbjct: 379 TTKLPIRKIPGDYGIPFIQPYKDRLDYFYNQGRDEYFKSRIQKYQSTIFRTNVPPGPFIA 558
Query: 73 SDSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDTTEPSHKLLK 132
+ V+ LLD SFP+LFD SK++K DV GT+ PST TGGYR ++ D +EP H+ LK
Sbjct: 559 QNPNVVVLLDGKSFPVLFDASKIDKTDVFTGTYTPSTELTGGYRVLSYLDPSEPKHEQLK 738
Query: 133 TFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATFNFLFLLLTD 192
+L S+ +P ++ + F+ L++QLA ++G ASF + A FNFL L
Sbjct: 739 KLMFFLLKSRSRHVIPEFQSCYREFFNALENQLA-ENGHASFADNNDQAAFNFLNRALFG 915
Query: 193 KHPSETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPFPAWTVKWSYKK 252
+P +T +G GP +V W+ QL P+ LGLPK + ED +I P +K Y++
Sbjct: 916 VNPVDTELGLDGPKMVQKWVLFQLGPVLKLGLPK---FVEDSMIHNFRLPFRLIKKDYQR 1086
Query: 253 LCEGFASAGASLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIGLAGPEL 312
L + F ++ L+EAER+ + ++EA HNL+F FN+FGG+ FP L+KWIG G L
Sbjct: 1087LYDFFYASSGFALEEAERLDVSKEEACHNLLFATCFNSFGGMKLFFPNLMKWIGRGGVRL 1266
Query: 313 HAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESH 372
H KLA EIR VR + GE++++A++ M L KSVVYE RI+P VP Q+ +A+ D+V+E+H
Sbjct: 1267HTKLATEIREAVRSAGGEITMAAMENMPLMKSVVYEAFRIDPPVPLQFGRAKRDMVIENH 1446
Query: 373 DAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVG-EGEKLLKYVYWSNGRETEEP 431
+ F +KKGE++ GYQPFATKDPK+FE EEFV RFVG EGEKLLK+V WSNG E++ P
Sbjct: 1447ENGFLVKKGELLLGYQPFATKDPKIFERAEEFVADRFVGDEGEKLLKHVLWSNGPESQSP 1626
Query: 432 TVDNKQCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKAT 484
TV NKQC K+ L+ RL VVE FLRYD+F + LGP++T+ SL +++
Sbjct: 1627TVGNKQCAGKDFTTLISRLLVVELFLRYDSFEIQVGNSPLGPSITLTSLKRSS 1785
Score = 48.5 bits (114), Expect = 6e-06
Identities = 21/39 (53%), Positives = 28/39 (70%)
Frame = -2
Query: 416 LLKYVYWSNGRETEEPTVDNKQCPAKNLVVLMCRLYVVE 454
LL++V WSNG E++ PTV NKQC K+ L+ RL VV+
Sbjct: 179 LLRHVLWSNGPESQSPTVGNKQCAGKDFTTLISRLLVVK 63
>TC93694 weakly similar to GP|14134199|gb|AAK54282.1 fatty acid
9-hydroperoxide lyase {Cucumis melo}, partial (37%)
Length = 791
Score = 345 bits (884), Expect = 3e-95
Identities = 163/257 (63%), Positives = 204/257 (78%), Gaps = 1/257 (0%)
Frame = +2
Query: 3 AASSDDTKNSSSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQGRDKFFATRVEKYNSTVIR 62
A+S + +++ LPLK IPGSYGLPF G + DRHDYFYNQGRDKFF+TR++KYNST+ R
Sbjct: 20 ASSKQEQSSTNKELPLKQIPGSYGLPFIGPIFDRHDYFYNQGRDKFFSTRIQKYNSTIFR 199
Query: 63 TNMPPGPGISSDSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQD 122
TNMPPGP ISS+ +VIALLD+ASFPILFDN KVEK +VLDGTFMPST FTGGYR CA+ D
Sbjct: 200 TNMPPGPFISSNPRVIALLDAASFPILFDNKKVEKLNVLDGTFMPSTKFTGGYRVCAYLD 379
Query: 123 TTEPSHKLLKTFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAAT 182
TTEP+H L+K F++ L + +TF+PL +T LSD F++++ L+ KSGKA FN+ +S A+
Sbjct: 380 TTEPNHALIKGFYLNTLLLRKDTFIPLFKTILSDGFNEIEDGLSSKSGKADFNSMVSVAS 559
Query: 183 FNFLF-LLLTDKHPSETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPF 241
FNF+F L DK+PSET++GD+GP + WL QLAPLATLG PKIFNY ED L+RT+PF
Sbjct: 560 FNFMFKLFCDDKNPSETILGDQGPKMFDTWLLFQLAPLATLGPPKIFNYLEDILLRTVPF 739
Query: 242 PAWTVKWSYKKLCEGFA 258
PA + SYKKL E F+
Sbjct: 740 PACLTRSSYKKLYEAFS 790
>TC90822 similar to PIR|A56377|A56377 rubber particle cytochrome P450 -
guayule, partial (35%)
Length = 885
Score = 182 bits (463), Expect = 2e-46
Identities = 95/190 (50%), Positives = 124/190 (65%), Gaps = 7/190 (3%)
Frame = +2
Query: 4 ASSDDTKNS------SSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQG-RDKFFATRVEKY 56
AS DT +S LP++ IPG YGLPF G + DR +YFYN+G RDK+F +++ KY
Sbjct: 311 ASGTDTTSSLVSPEPPKKLPMRRIPGDYGLPFVGPIKDRLEYFYNEGGRDKYFRSKMHKY 490
Query: 57 NSTVIRTNMPPGPGISSDSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYR 116
ST+ R NMPPGP ISS+ VI LLD SFP+LFDNSKVEKRD+ GTFMPST TGGYR
Sbjct: 491 KSTIFRANMPPGPFISSNPNVIVLLDGKSFPVLFDNSKVEKRDLFTGTFMPSTELTGGYR 670
Query: 117 ACAFQDTTEPSHKLLKTFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNT 176
++ D TEP H LK +L S+ + F+P ++ ++ F+ L +LA K+G A F
Sbjct: 671 ILSYLDPTEPKHDQLKRLLFFLLKSRSSHFIPEFHSSYTNLFNTLXKELA-KTGNAIFAD 847
Query: 177 SISAATFNFL 186
+ A FN+L
Sbjct: 848 ANDQAAFNYL 877
>TC78050 homologue to GP|5830469|emb|CAB54849.1 hydroperoxide lyase
{Medicago sativa}, partial (45%)
Length = 697
Score = 132 bits (333), Expect = 2e-31
Identities = 76/189 (40%), Positives = 106/189 (55%), Gaps = 2/189 (1%)
Frame = +2
Query: 10 KNSSSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQGRDKFFATRVEKYNSTVIRTNMPPG- 68
K + LP++ IPGS+G P G + DR DYF+ Q + FF TR++KY STV RTN+PP
Sbjct: 98 KARPTELPIRQIPGSHGWPLLGPLSDRLDYFWFQKPENFFRTRMDKYKSTVFRTNVPPTF 277
Query: 69 PGISS-DSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDTTEPS 127
P ++ + +IA+LD SF LFD V+K+DVL G F+PS FTG R +QD +EP
Sbjct: 278 PFFTNVNPNIIAVLDCKSFSHLFDMDLVDKKDVLVGDFVPSVEFTGNIRVGVYQDVSEPQ 457
Query: 128 HKLLKTFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATFNFLF 187
H K+F M +L + ++P L + L D F D S S+ + + F FL
Sbjct: 458 HAKAKSFSMNILKQSSSIWVPELISNL-DIFLDQIEATLSNSSSVSYFSPLQQFLFTFLS 634
Query: 188 LLLTDKHPS 196
+L PS
Sbjct: 635 KVLARADPS 661
>TC91861 homologue to GP|5830467|emb|CAB54848.1 hydroperoxide lyase
{Medicago sativa}, partial (31%)
Length = 532
Score = 122 bits (307), Expect = 2e-28
Identities = 62/154 (40%), Positives = 90/154 (58%), Gaps = 1/154 (0%)
Frame = +2
Query: 331 VSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIKKGEMIFGYQPF 390
+ +L ++ L SVVYE LR+ P VP Q+ +AR+D + S+D+AF +KKGE++ G+Q
Sbjct: 20 LGFDSLKELELINSVVYETLRMNPPVPLQFGRARKDFQLSSYDSAFNVKKGELLCGFQKL 199
Query: 391 ATKDPKVFENPEEFVGHRFVGE-GEKLLKYVYWSNGRETEEPTVDNKQCPAKNLVVLMCR 449
+DP F+ PE+F RF E G +LL Y+YWSNG +T P+ NKQC K++V
Sbjct: 200 IMRDPVAFDEPEQFKPERFTKEKGAELLNYLYWSNGPQTGSPSGSNKQCAGKDIVTFTAA 379
Query: 450 LYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKA 483
L V RYD ++ G +I +L KA
Sbjct: 380 LIVAHLLRRYD--------LIKGDGSSITALRKA 457
>TC86135 similar to PIR|A96673|A96673 probable cytochrome P450 F13O11.25
[imported] - Arabidopsis thaliana, partial (62%)
Length = 1842
Score = 54.7 bits (130), Expect = 8e-08
Identities = 64/213 (30%), Positives = 92/213 (43%), Gaps = 14/213 (6%)
Frame = +2
Query: 262 ASLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAE 318
A L E E KR +E+ +V + GG T+ ++WI + PE+ +L E
Sbjct: 842 ADTLLELELPEEKRKLSENEMVNLCSEFLNGG-TDTTSTALQWIMANLVKYPEVQGRLVE 1018
Query: 319 EIRTV-VRESNGE---VSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKA-REDVVVESHD 373
EIR V V + NGE V L K+ K VV E LR P + A EDVV+ +
Sbjct: 1019 EIREVMVSDDNGEKEEVKEENLQKLRYLKCVVLEGLRRHPPGHFVLPHAVSEDVVLNGYL 1198
Query: 374 AAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGE------GEKLLKYVYWSNGRE 427
K G + F DP+V+E+P EF RF+ + G K +K + + GR
Sbjct: 1199 VP---KDGTVNFMVAEMGL-DPRVWEDPMEFKPERFLKDETFDITGSKEIKMMPFGAGR- 1363
Query: 428 TEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYD 460
+ CP NL +L +V +D
Sbjct: 1364 --------RICPGYNLALLHLEYFVANLVWNFD 1438
>TC86137 weakly similar to PIR|A96673|A96673 probable cytochrome P450
F13O11.25 [imported] - Arabidopsis thaliana, partial
(69%)
Length = 1695
Score = 53.5 bits (127), Expect = 2e-07
Identities = 64/213 (30%), Positives = 94/213 (44%), Gaps = 14/213 (6%)
Frame = +3
Query: 262 ASLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAE 318
A L E E KR +E+ +V + GG T+ ++WI + PE+ +L E
Sbjct: 891 ADTLLELEWPEEKRKLSENEMVNLCSEFLNGG-TDTTSTSLQWIMANVVKYPEVQGRLVE 1067
Query: 319 EIRTVV-RESNG---EVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKA-REDVVVESHD 373
EIR V+ + NG EV L K+ K VV E LR P+ + A +EDVV+ D
Sbjct: 1068 EIREVMGGDENGEKEEVKEEDLQKLRYLKCVVLEGLRRHPSGKFPLPHAVKEDVVL---D 1238
Query: 374 AAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGE------GEKLLKYVYWSNGRE 427
K G + F A D +V+E+P EF RF+ + G K +K + + GR
Sbjct: 1239 GYLVPKNGTVNFLLAEMAL-DRRVWEDPLEFKPERFLKDETFDITGSKEIKMMPFGAGR- 1412
Query: 428 TEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYD 460
+ CP NL +L +V +D
Sbjct: 1413 --------RICPGLNLALLHLEYFVANLVWNFD 1487
>TC87736 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P450
monooxygenase {Cicer arietinum}, partial (55%)
Length = 1367
Score = 51.2 bits (121), Expect = 9e-07
Identities = 44/153 (28%), Positives = 74/153 (47%), Gaps = 4/153 (2%)
Frame = +3
Query: 263 SLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEE 319
SLL E + + R++ +H L +L GG T+ ++W L P + +K+ +E
Sbjct: 423 SLLAEENKKELDREKIQHLLHDLL----VGG-TDTTTYTLEWAMAELLHNPNVMSKVKKE 587
Query: 320 IRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYA-KAREDVVVESHDAAFEI 378
+ + N + S + ++ ++++ E LR+ P P KA+EDV V + I
Sbjct: 588 LEETIGIGN-PIEESDVTRLPYLQAIIKETLRLHPIAPLLLPRKAKEDVEVNG----YLI 752
Query: 379 KKGEMIFGYQPFATKDPKVFENPEEFVGHRFVG 411
KG IF +DPKV++NP F RF+G
Sbjct: 753 PKGAQIFVNVWAIGRDPKVWDNPNLFSPERFLG 851
>TC83798 weakly similar to GP|10177750|dbj|BAB11063. cytochrome P450
{Arabidopsis thaliana}, partial (25%)
Length = 762
Score = 50.8 bits (120), Expect = 1e-06
Identities = 34/104 (32%), Positives = 53/104 (50%), Gaps = 1/104 (0%)
Frame = +1
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
P++ KL EI VV +N V S + KM +S V EVLR+ P P+ ++ ED +
Sbjct: 70 PKILKKLRAEINDVVG-TNRLVKESDIPKMPYLQSCVKEVLRLHPTAPFALRQSSEDCKI 246
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRF-VGE 412
+D + ++ +DP+ + NPEE++ RF VGE
Sbjct: 247 NGYDIKAHTRTLVNVYAIM----RDPQSWVNPEEYIPERFLVGE 366
>TC79164 similar to SP|O48928|C773_SOYBN Cytochrome P450 77A3 (EC 1.14.-.-).
[Soybean] {Glycine max}, partial (93%)
Length = 1592
Score = 49.3 bits (116), Expect = 3e-06
Identities = 41/146 (28%), Positives = 71/146 (48%), Gaps = 4/146 (2%)
Frame = +3
Query: 274 KRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIGLA----GPELHAKLAEEIRTVVRESNG 329
K + LV ++ GG T+ ++W G+A PE+ AKL +EI ++V +
Sbjct: 975 KSTPSNEELVSLISEFLNGG-TDTTATAVEW-GIAQLIDNPEIQAKLYQEINSIVGDK-- 1142
Query: 330 EVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIKKGEMIFGYQP 389
+V ++KM ++VV E+LR P + A V + ++I + Y
Sbjct: 1143 KVEEKDVEKMPYLQAVVKELLRKHPPTHFVLTHA---VTEPTTLGGYDIPIDANVEVYTA 1313
Query: 390 FATKDPKVFENPEEFVGHRFVGEGEK 415
+DPK++ NPE+F RF+ +GE+
Sbjct: 1314 GIGEDPKLWSNPEKFDPERFLSKGEE 1391
>TC92736 weakly similar to SP|Q9STK9|C71O_ARATH Cytochrome P450 71A24 (EC
1.14.-.-). [Mouse-ear cress] {Arabidopsis thaliana},
partial (34%)
Length = 781
Score = 48.1 bits (113), Expect = 7e-06
Identities = 36/136 (26%), Positives = 62/136 (45%), Gaps = 4/136 (2%)
Frame = +3
Query: 279 EHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEEIRTVVRESNGEVSLSA 335
+ ++ L + F T+ L++W L P + KL EE+R+V ++
Sbjct: 60 DKTVIKALLLDMFSAGTDTISTLLEWSMTELLRHPNIMKKLQEEVRSVAGNRT-HITEDD 236
Query: 336 LDKMTLTKSVVYEVLRIEPAVPYQYAKA-REDVVVESHDAAFEIKKGEMIFGYQPFATKD 394
L M ++VV E LR+ P +P + R+D+ V+ + IK G + +D
Sbjct: 237 LGNMKYLQAVVKETLRLHPPIPLLVPRENRQDITVKG----YHIKAGTRVIINAWAIARD 404
Query: 395 PKVFENPEEFVGHRFV 410
P ++ PEEF RF+
Sbjct: 405 PAYWDQPEEFKPERFL 452
>TC88716 weakly similar to GP|20258812|gb|AAM14017.1 putative cytochrome
P450 homolog {Arabidopsis thaliana}, partial (57%)
Length = 1228
Score = 48.1 bits (113), Expect = 7e-06
Identities = 32/102 (31%), Positives = 53/102 (51%)
Frame = +2
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
PE+ K+ EE+ TVV + N V S + K+T +V+ E LR+ PA+P +
Sbjct: 863 PEVMRKVQEELETVVGKDN-LVEESHIHKLTYLHAVMKETLRLHPALPLLVPHCPSET-- 1033
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVG 411
++ + I +G +F +DP V+ENP EF ++ +G
Sbjct: 1034-TNIGGYTIPEGSRVFINVWAIHRDPYVWENPLEFDPYKVLG 1156
>TC86308 similar to SP|O48927|CP78_SOYBN Cytochrome P450 78A3 (EC 1.14.-.-).
[Soybean] {Glycine max}, partial (88%)
Length = 2078
Score = 48.1 bits (113), Expect = 7e-06
Identities = 48/193 (24%), Positives = 90/193 (45%), Gaps = 10/193 (5%)
Frame = +3
Query: 278 AEHNLVFMLGFNAFGGLTNQFPILIKWIGLAGPELHAKLAEEIRTVVRE-SNGE---VSL 333
++ +++ +L F G T+ +LI+WI LA +H + ++++T + E ++GE ++
Sbjct: 1032 SDSDMIAVLWEMIFRG-TDTVAVLIEWI-LARLVIHPDVQKKVQTELDEVASGESCAITE 1205
Query: 334 SALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIKKGEMIFGYQPFATK 393
+ M +V+ EVLR+ P P + AR + + D + + G ++
Sbjct: 1206 EDVAAMVYLPAVIKEVLRLHPPGPL-LSWARLAITDTTIDG-YHVPAGTTAMVNMWAISR 1379
Query: 394 DPKVFENPEEFVGHRFVGEGEKL------LKYVYWSNGRETEEPTVDNKQCPAKNLVVLM 447
DP V+ NP EF RFV EG + L+ + +GR + CP KNL +
Sbjct: 1380 DPDVWRNPLEFNPERFVSEGAEFSVLGSDLRLAPFGSGR---------RSCPGKNLGLAT 1532
Query: 448 CRLYVVEFFLRYD 460
+V + ++
Sbjct: 1533 VTFWVAKLLHEFE 1571
>BM814294 weakly similar to GP|6002279|emb cytochrome P450 monooxygenase
{Cicer arietinum}, partial (37%)
Length = 793
Score = 47.8 bits (112), Expect = 1e-05
Identities = 46/155 (29%), Positives = 74/155 (47%), Gaps = 6/155 (3%)
Frame = +1
Query: 263 SLLD--EAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLA 317
SLLD E R + R++ EH L +L GG T+ ++W L P + +K+
Sbjct: 151 SLLDIPEENRKELDREKIEHLLHDLL----VGG-TDTTTYTLEWAMAELLHNPNIMSKVK 315
Query: 318 EEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYA-KAREDVVVESHDAAF 376
+E+ + N + S + ++ ++V+ E LR+ P P KA+EDV V +
Sbjct: 316 KELEDTIGIGN-PLEESDITRLPYLQAVIKETLRLHPIAPLLLPRKAKEDVEVNG----Y 480
Query: 377 EIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVG 411
I K IF +DP+V++NP F RF+G
Sbjct: 481 TIPKDAQIFVNVWAIGRDPEVWDNPYLFSPERFLG 585
>TC85609 weakly similar to SP|Q9LTL2|C72P_ARATH Cytochrome P450 71B25 (EC
1.14.-.-). [Mouse-ear cress] {Arabidopsis thaliana},
partial (33%)
Length = 1786
Score = 47.8 bits (112), Expect = 1e-05
Identities = 35/158 (22%), Positives = 70/158 (44%), Gaps = 4/158 (2%)
Frame = +1
Query: 307 LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKARED 366
+ P K+ EEIR V G + ++K+ K+V+ E +R+ P +P + +
Sbjct: 1009 MKNPRAMQKVQEEIRKVCA-GKGFIEEEDVEKLPYFKAVIKESMRLYPILPILLPR---E 1176
Query: 367 VVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKL----LKYVYW 422
+ + A ++I +++ +DP+V+++PEEF RF+G L + + +
Sbjct: 1177 TMTNCNIAGYDIPDKTLVYVNALAIHRDPEVWKDPEEFYPERFIGSDIDLKGQDFELIPF 1356
Query: 423 SNGRETEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYD 460
+GR + CP N+ + L + +D
Sbjct: 1357 GSGR---------RICPGLNMAIATIDLVLSNLLYSFD 1443
>TC77326 similar to GP|12004682|gb|AAG44132.1 cytochrome P450 {Pisum sativum},
complete
Length = 1864
Score = 47.4 bits (111), Expect = 1e-05
Identities = 30/104 (28%), Positives = 52/104 (49%), Gaps = 1/104 (0%)
Frame = +1
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAK-AREDVV 368
PE+ K EE+ V+ + V + + ++ E +R+ P P+ + ARED
Sbjct: 1033 PEIFKKATEELDRVIGKDRW-VEEKDIANLPYVYAIAKETMRLHPVAPFLVPREAREDCK 1209
Query: 369 VESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGE 412
V+ +D I KG ++ +D +V+ENP EF+ RF+G+
Sbjct: 1210 VDGYD----IPKGTIVLVNTWTIARDSEVWENPYEFMPERFLGK 1329
>TC83793 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P450
monooxygenase {Cicer arietinum}, partial (48%)
Length = 923
Score = 47.0 bits (110), Expect = 2e-05
Identities = 34/152 (22%), Positives = 69/152 (45%), Gaps = 3/152 (1%)
Frame = +2
Query: 263 SLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEE 319
++LD+ + G E + + L + F T+ ++W L P + +K E
Sbjct: 185 AMLDDEKNAG----EMYKDKIERLSVDLFVAGTDTVTSTLEWAMAELLHNPNIMSKAKSE 352
Query: 320 IRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIK 379
+ ++ + N V S + K+ ++++ E R+ PAVP + E ++ +++
Sbjct: 353 LNQIIGKGNS-VEESDIGKLPYLQAIIKETFRLHPAVPLLLPRKAE---IDLEINGYKVP 520
Query: 380 KGEMIFGYQPFATKDPKVFENPEEFVGHRFVG 411
KG + +D ++ENP EF+ RF+G
Sbjct: 521 KGAQVLINVWAIGRDSNLWENPNEFLPERFLG 616
>TC82503 weakly similar to GP|18252325|gb|AAL66194.1 cytochrome P450 {Pyrus
communis}, partial (39%)
Length = 806
Score = 45.8 bits (107), Expect = 4e-05
Identities = 48/176 (27%), Positives = 72/176 (40%), Gaps = 8/176 (4%)
Frame = +1
Query: 293 GLTNQFPILIKWIG---LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEV 349
G T+ I+W L P + K+ +E+ VV +V S L+K+ V+ E
Sbjct: 70 GATDTSATAIEWTISELLKNPRVMKKVQKELEIVVGMKR-KVEESDLEKLEYLNMVIKES 246
Query: 350 LRIEPAVPYQYA-KAREDVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHR 408
LR P VP ++ ED VE F I K I +DP + +PE+F R
Sbjct: 247 LRFHPVVPLSVPHQSMEDCTVEE----FFIPKNSRIMVNAWAIMRDPNSWTDPEKFWPER 414
Query: 409 FVGE----GEKLLKYVYWSNGRETEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYD 460
F G G + + + +GR + CP L + M RL V + +D
Sbjct: 415 FEGNNIDVGGQHFHLIPFGSGR---------RGCPGLQLGLTMVRLAVAQLVHCFD 555
>TC93666 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (37%)
Length = 675
Score = 45.4 bits (106), Expect = 5e-05
Identities = 29/107 (27%), Positives = 50/107 (46%)
Frame = +1
Query: 307 LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKARED 366
+ P + K +E+R V++ G V +++++ KSVV E LR+ P+VP +
Sbjct: 43 ITNPRIMKKAQDEVRDVIK-MKGRVDEKGINELSYLKSVVKETLRLHPSVPLLLPRECGK 219
Query: 367 VVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEG 413
E H +K +I + +DP + PE+F RF+ G
Sbjct: 220 AC-EIHGYHIPVKTKVIINAWA--IARDPNYWTEPEKFYPERFIDNG 351
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.137 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,327,417
Number of Sequences: 36976
Number of extensions: 199323
Number of successful extensions: 1050
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 1032
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1039
length of query: 487
length of database: 9,014,727
effective HSP length: 100
effective length of query: 387
effective length of database: 5,317,127
effective search space: 2057728149
effective search space used: 2057728149
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0091b.14


