

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.13
(267 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
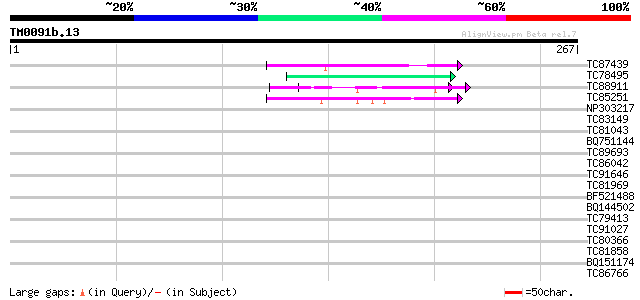
Score E
Sequences producing significant alignments: (bits) Value
TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in... 44 5e-05
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 43 1e-04
TC88911 weakly similar to GP|13926280|gb|AAK49610.1 AT5g14920/F2... 41 5e-04
TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean... 40 7e-04
NP303217 NP303217|AJ297721.1|CAC20726.1 putative repetitive prol... 39 0.002
TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk p... 39 0.002
TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 39 0.002
BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like ... 39 0.002
TC89693 weakly similar to GP|21104633|dbj|BAB93225. hypothetical... 39 0.003
TC86042 similar to GP|20466712|gb|AAM20673.1 putative SF16 prote... 38 0.004
TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 38 0.004
TC81969 similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich... 38 0.005
BF521488 weakly similar to GP|18447433|gb RE28280p {Drosophila m... 37 0.006
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 37 0.006
TC79413 similar to GP|21439800|emb|CAD35173. unnamed protein pro... 37 0.008
TC91027 similar to GP|14517502|gb|AAK62641.1 At2g38310/T19C21.20... 37 0.010
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 37 0.010
TC81858 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 36 0.014
BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xy... 35 0.023
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 35 0.030
>TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in cucumber
hypocotyls {Cucumis sativus}, partial (57%)
Length = 897
Score = 44.3 bits (103), Expect = 5e-05
Identities = 31/96 (32%), Positives = 41/96 (42%), Gaps = 4/96 (4%)
Frame = +3
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTD----SPKPPQIKGCQPKNVLPKADPPKSSPSNAA 177
TP++ PP +P+ A PK+ SPK P P A P S+P A
Sbjct: 153 TPVSSPVAAPPTTPTTPAPVASPKSSPPATSPKAAAPTATPPAASSPPAVTPVSTPPPAP 332
Query: 178 EAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+++PP AP VS+P VTP P T P
Sbjct: 333 VPVKSPPTPAP--------VSSPPAVTPVAAPTTTP 416
Score = 30.8 bits (68), Expect = 0.57
Identities = 27/107 (25%), Positives = 38/107 (35%), Gaps = 16/107 (14%)
Frame = +3
Query: 123 PITMIRYNPPFKKP-----------SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKS 171
P+ + +PP P + S PA +P P + P P + PP
Sbjct: 207 PVASPKSSPPATSPKAAAPTATPPAASSPPAVTPVSTPPPAPVPVKSPPTPAPVSSPPAV 386
Query: 172 SPSNAAEAIRAPPNTAPM--KKALSQH---VSAPKKVTPSKRPATQP 213
+P A A P AP KK +H +P + P PA P
Sbjct: 387 TPVAAPTTTPAVPAPAPSKGKKNKKKHGAPAPSPALLGPPAPPAGAP 527
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 42.7 bits (99), Expect = 1e-04
Identities = 23/80 (28%), Positives = 30/80 (36%)
Frame = +3
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP + P + P PP ++ P P A PP SP A PP P
Sbjct: 408 PPVQSSPPPVQQSPPPTPLTPPPVQSTPPPASPPPASPPPFSPPPATPPPATPPPATPPP 587
Query: 191 KALSQHVSAPKKVTPSKRPA 210
+S+P TP+ PA
Sbjct: 588 ALTPTPLSSPPATTPAPAPA 647
Score = 30.8 bits (68), Expect = 0.57
Identities = 28/91 (30%), Positives = 35/91 (37%), Gaps = 2/91 (2%)
Frame = +3
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKP--PQIKGCQPKNVLPKADPPKSSPSNAAEA 179
TP+T PP + P PA P SP P P P P PP +P+ +
Sbjct: 456 TPLTP----PPVQSTPP--PASPPPASPPPFSPPPATPPPATPPPATPPPALTPTPLSSP 617
Query: 180 IRAPPNTAPMKKALSQHVSAPKKVTPSKRPA 210
P AP K AP ++PS PA
Sbjct: 618 PATTPAPAPAKLKSKAPALAP-VLSPSDAPA 707
Score = 28.5 bits (62), Expect = 2.8
Identities = 22/96 (22%), Positives = 33/96 (33%)
Frame = +3
Query: 118 GVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAA 177
G++C + + S PA P + PP P V P +SSP
Sbjct: 198 GLICVVFSSVGAQQAPSTSPNSSPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSP---- 365
Query: 178 EAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+++ P A Q P + +P P T P
Sbjct: 366 PPLQSSPPPAQSTPPPVQSSPPPVQQSPPPTPLTPP 473
>TC88911 weakly similar to GP|13926280|gb|AAK49610.1 AT5g14920/F2G14_40
{Arabidopsis thaliana}, partial (11%)
Length = 1320
Score = 40.8 bits (94), Expect = 5e-04
Identities = 27/83 (32%), Positives = 34/83 (40%), Gaps = 10/83 (12%)
Frame = +2
Query: 137 SPSMPAQPKTDSPKPPQIKGCQPKNV----------LPKADPPKSSPSNAAEAIRAPPNT 186
+P+ P PK +P +K PK +PKA PK SP PP T
Sbjct: 167 APAKPLVPKVTTPTAAPVKPIVPKTTTPTSAPLKPPVPKAPAPKPSPVKPPVPKSKPPTT 346
Query: 187 APMKKALSQHVSAPKKVTPSKRP 209
AP+K V P K+ P K P
Sbjct: 347 APVKSP----VPDPPKLAPVKLP 403
Score = 40.4 bits (93), Expect = 7e-04
Identities = 29/97 (29%), Positives = 43/97 (43%), Gaps = 2/97 (2%)
Frame = +2
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
P T + P K P P PA PK KPP +PK+ PP ++P +
Sbjct: 233 PKTTTPTSAPLKPPVPKAPA-PKPSPVKPP----------VPKSKPPTTAP--VKSPVPD 373
Query: 183 PPNTAPMKKALSQHVSA--PKKVTPSKRPATQPLRRS 217
PP AP+K + + A PK +P +P P +++
Sbjct: 374 PPKLAPVKLPVPKVTPALSPKTPSPKIQPPPHPPKKA 484
Score = 34.7 bits (78), Expect = 0.039
Identities = 30/96 (31%), Positives = 42/96 (43%), Gaps = 12/96 (12%)
Frame = +2
Query: 132 PFKKPSPSMPAQPKTDSP-KPPQIKGCQP-----KNVLPKADPPKSSPSNAAEAIRAPPN 185
P+K P P A T +P KPP K P K ++PK P ++P P
Sbjct: 80 PWKSPVPK--ATTPTSAPVKPPVPKVATPTVAPAKPLVPKVTTPTAAPVKPIVPKTTTPT 253
Query: 186 TAPMKKALSQHVSAPK------KVTPSKRPATQPLR 215
+AP+K + + APK V SK P T P++
Sbjct: 254 SAPLKPPVPK-APAPKPSPVKPPVPKSKPPTTAPVK 358
Score = 28.9 bits (63), Expect = 2.2
Identities = 24/83 (28%), Positives = 34/83 (40%)
Frame = +3
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
P + P+P+ + K +P P + + P K PKSSP + + PP+ AP
Sbjct: 606 PAKEAPAPAPTHKKKAPAPAPDK-ETPAPAPTHKKKKAPKSSPVPSPAIL--PPSPAP-- 770
Query: 191 KALSQHVSAPKKVTPSKRPATQP 213
P TPS PA P
Sbjct: 771 --------TPAIDTPSSAPAPSP 815
>TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean, partial
(93%)
Length = 1554
Score = 40.4 bits (93), Expect = 7e-04
Identities = 36/113 (31%), Positives = 47/113 (40%), Gaps = 21/113 (18%)
Frame = -2
Query: 122 TPITMIRYNPPFKKPSPSMPAQPK-TDSPKPPQIKGCQPKNV---LPKADPP---KSSPS 174
T IT + PP SP + +QP T +PK + PK PK PP KSSP
Sbjct: 809 TAITPVTTQPPTVVASPPITSQPPVTVAPKSAPVTSPAPKIAPASSPKVPPPQPPKSSPV 630
Query: 175 N--------------AAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ + ++ PP AP+ KA AP K P+ PAT P
Sbjct: 629 STPTLPPPLPPPPKISPTPVQTPPAPAPV-KATPVPAPAPAKQAPTPAPATSP 474
>NP303217 NP303217|AJ297721.1|CAC20726.1 putative repetitive proline-rich
protein
Length = 525
Score = 39.3 bits (90), Expect = 0.002
Identities = 32/89 (35%), Positives = 38/89 (41%), Gaps = 4/89 (4%)
Frame = +1
Query: 129 YNPPFKKPSPS-MPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
YNPP KP P+ P K K P K K LPK PP P + I PP+
Sbjct: 157 YNPPIYKPPPTYKPPIKKQPINKSPNKKPLLLKPPLPK--PPHKEPPSRKRPINTPPDKK 330
Query: 188 P--MKKALSQHVSAPKKVTPS-KRPATQP 213
P +K + P+K PS KRP P
Sbjct: 331 PPLLKPPFPK---PPQKKPPSRKRPINTP 408
Score = 35.8 bits (81), Expect = 0.018
Identities = 22/61 (36%), Positives = 25/61 (40%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP KKP P PK KPP K +P N P PP P + PP+ P
Sbjct: 316 PPDKKPPLLKPPFPKPPQKKPPSRK--RPINTPPNKKPPLPKP-----PVNKPPHKKPSN 474
Query: 191 K 191
K
Sbjct: 475 K 477
Score = 31.6 bits (70), Expect = 0.33
Identities = 21/54 (38%), Positives = 21/54 (38%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPP 184
PPF KP P K PP K PK P PP PSN R PP
Sbjct: 346 PPFPKPPQKKPPSRKRPINTPPNKKPPLPKP--PVNKPPHKKPSNK----RPPP 489
>TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk protein
{Argiope trifasciata}, partial (3%)
Length = 1177
Score = 39.3 bits (90), Expect = 0.002
Identities = 25/91 (27%), Positives = 41/91 (44%), Gaps = 4/91 (4%)
Frame = +3
Query: 125 TMIRYNPPFKKPSPSMPAQPKTDSPKPPQIK-GCQPKNVLPKADPPKSSPSNAAEAIRAP 183
T +R PP + SP+ P+ P+T S P +P++ P PP P + ++P
Sbjct: 45 TPLRSTPPTRPSSPT-PSAPRTSSRSTPSTSPSRRPRSKTPLPPPPPPLPQHPKPPQKSP 221
Query: 184 PNTAPMKKALSQHVSA---PKKVTPSKRPAT 211
P K + S + P++ TP P+T
Sbjct: 222 PRMRRSKPSPSSPAATPLWPQRTTPPPSPST 314
Score = 36.6 bits (83), Expect = 0.010
Identities = 26/108 (24%), Positives = 38/108 (35%), Gaps = 16/108 (14%)
Frame = +3
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADP------------- 168
+P R P P P +P PK PP+++ +P P A P
Sbjct: 132 SPSRRPRSKTPLPPPPPPLPQHPKPPQKSPPRMRRSKPSPSSPAATPLWPQRTTPPPSPS 311
Query: 169 -PKSSPSNAAEAIRAP--PNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
P+ SP A +P P P + P+ +PS P +P
Sbjct: 312 TPRPSP*TPATPSTSPTAPLPTPPPRTTLPPALTPRPPSPSTPPTPKP 455
Score = 34.7 bits (78), Expect = 0.039
Identities = 31/93 (33%), Positives = 34/93 (36%)
Frame = +3
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
TP T P K P P P P PKPPQ P+ K SPS+ A
Sbjct: 120 TPSTSPSRRPRSKTPLPP-PPPPLPQHPKPPQKSP-------PRMRRSKPSPSSPAATPL 275
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQPL 214
P T P + S TPS P T PL
Sbjct: 276 WPQRTTPPPSPSTPRPSP*TPATPSTSP-TAPL 371
>TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (19%)
Length = 555
Score = 38.9 bits (89), Expect = 0.002
Identities = 26/79 (32%), Positives = 30/79 (37%)
Frame = +1
Query: 132 PFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKK 191
P K P+P P P +P PP I P A P SP + A APP P K
Sbjct: 238 PPKPPAPPSPGSPPAPAPAPPPIPPS------PPAPPSPGSPPSPPPASPAPPKPTPPK- 396
Query: 192 ALSQHVSAPKKVTPSKRPA 210
+P P K PA
Sbjct: 397 ------GSPLLPMPEKTPA 435
Score = 29.6 bits (65), Expect = 1.3
Identities = 19/43 (44%), Positives = 19/43 (44%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSP 173
PP PSP PA P SP P P PK PPK SP
Sbjct: 295 PPPIPPSP--PAPPSPGSPPSPPPASPAP----PKPTPPKGSP 405
>BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like protein
(clone pMG14) - common tobacco (fragment), partial (11%)
Length = 632
Score = 38.9 bits (89), Expect = 0.002
Identities = 27/87 (31%), Positives = 40/87 (45%), Gaps = 5/87 (5%)
Frame = +1
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIK---GCQPKNVLPKADPPK--SSPSNAAEAIRAP 183
++PP PS S P P T SP P +P P + PP+ SSPS + + P
Sbjct: 313 FSPPPPSPSRSRPPPPSTPSPLLPLPLLPLATEPSPFPPSSPPPRPASSPSLSPPVLLFP 492
Query: 184 PNTAPMKKALSQHVSAPKKVTPSKRPA 210
P+ +P + S+ P + P RP+
Sbjct: 493 PSASPPPWSPSR---PPSALAPPSRPS 564
Score = 33.9 bits (76), Expect = 0.067
Identities = 32/107 (29%), Positives = 45/107 (41%), Gaps = 12/107 (11%)
Frame = +1
Query: 137 SPSMPAQPKT-DSPKPPQIKGCQPKNVLPKADPPKSSPSN---------AAEAIRAPPNT 186
SPS P +P + SP PP +P PP S+PS A E PP++
Sbjct: 280 SPSSPTRPSSLFSPPPPSPSRSRP--------PPPSTPSPLLPLPLLPLATEPSPFPPSS 435
Query: 187 APMKKALSQHVSAPKKVTP--SKRPATQPLRRSNRLIWSNRKNLEQS 231
P + A S +S P + P + P P R + L +R +L S
Sbjct: 436 PPPRPASSPSLSPPVLLFPPSASPPPWSPSRPPSALAPPSRPSLPAS 576
>TC89693 weakly similar to GP|21104633|dbj|BAB93225. hypothetical
protein~similar to Arabidopsis thaliana chromosome 3
At3g46550, partial (53%)
Length = 1346
Score = 38.5 bits (88), Expect = 0.003
Identities = 20/73 (27%), Positives = 33/73 (44%), Gaps = 1/73 (1%)
Frame = +2
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLP-KADPPKSSPSNAAEAIRAPPNTAP 188
+PP P+ S SP P P N P + PP + P+ +A I P T
Sbjct: 272 SPPPPSPTSSATTSSSNSSPGPTSNTSLPPVNSSPLSSKPPAAPPTTSAPLISPTPRTLT 451
Query: 189 MKKALSQHVSAPK 201
+ +++ QH++ P+
Sbjct: 452 LLQSIPQHLTHPQ 490
>TC86042 similar to GP|20466712|gb|AAM20673.1 putative SF16 protein
{Arabidopsis thaliana}, partial (39%)
Length = 2305
Score = 38.1 bits (87), Expect = 0.004
Identities = 34/136 (25%), Positives = 53/136 (38%), Gaps = 6/136 (4%)
Frame = +1
Query: 72 KEDDAENHNIRPLLNDVDAMHLGAYAEYKKQNVEVFVEHDDEELPFGVLCTPITMIRYNP 131
K D+EN + + V + G + + +F E E FG ++ P
Sbjct: 466 KGKDSENKSTKEKKKGVGKLKHGETNSF----IPLFREPSSIEKIFGDFEREQQVLAIRP 633
Query: 132 PFKKPSPSMP--AQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNT--- 186
P P P P+ SP+PP K P++ P+A P+ + AA + +
Sbjct: 634 PTPPERPKTPPFVPPRVASPRPPSPKPPSPRDPSPRAASPRVTSPKAASSRNVHQHKEVR 813
Query: 187 -APMKKALSQHVSAPK 201
P +QHVSA K
Sbjct: 814 YRPEPTLQNQHVSATK 861
>TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (15%)
Length = 984
Score = 38.1 bits (87), Expect = 0.004
Identities = 30/106 (28%), Positives = 35/106 (32%), Gaps = 5/106 (4%)
Frame = +1
Query: 116 PFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKG-----CQPKNVLPKADPPK 170
P L P + PP SPS A P SP P + P LP++ P
Sbjct: 274 PLSPLRFPSRLPLTTPPSLTTSPSPTASPPPRSPPRPPPRSPPRQLMSPPRPLPRS--PS 447
Query: 171 SSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRR 216
P N E R P P S P P RP +P R
Sbjct: 448 PPPRNPPELSRTTPTPRPPMSPSPSPTSPPAATPPVSRPPPRPALR 585
Score = 26.9 bits (58), Expect = 8.2
Identities = 23/85 (27%), Positives = 30/85 (35%), Gaps = 4/85 (4%)
Frame = +1
Query: 131 PPFKKPSPSMPAQ--PKTDSPKPPQIKGCQPKNVLPKADPPKS--SPSNAAEAIRAPPNT 186
P P P P + T +P+PP P + P A PP S P A + PP
Sbjct: 433 PRSPSPPPRNPPELSRTTPTPRPPMSPSPSPTSP-PAATPPVSRPPPRPALRRLLPPPPL 609
Query: 187 APMKKALSQHVSAPKKVTPSKRPAT 211
P + + V P P T
Sbjct: 610 PPTLPSSMASLFLAASVLPPASPPT 684
>TC81969 similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (11%)
Length = 556
Score = 37.7 bits (86), Expect = 0.005
Identities = 25/92 (27%), Positives = 37/92 (40%)
Frame = -2
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
+P++ NP P P P P PP P LP + PP PS+ +
Sbjct: 390 SPVSSYSLNPSSSSPPPPSPPPP------PPSFSHPSPSPNLPFSPPPLEPPSSGSS--- 238
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PN + K + S + P +PS P+ +P
Sbjct: 237 --PNPSCSKVSQSTLLCIPPPQSPSCPPSPRP 148
>BF521488 weakly similar to GP|18447433|gb RE28280p {Drosophila
melanogaster}, partial (84%)
Length = 470
Score = 37.4 bits (85), Expect = 0.006
Identities = 24/57 (42%), Positives = 27/57 (47%)
Frame = -3
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNT 186
NPP P P +PK PKPP K +PK PK PPK + AA A NT
Sbjct: 237 NPP*GL*PPPKPGRPKPPPPKPPPPKPGRPKPP-PKPGPPKPNCGLAATATIKRANT 70
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 37.4 bits (85), Expect = 0.006
Identities = 20/54 (37%), Positives = 24/54 (44%)
Frame = -2
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPP 184
PP P+ S P +P P PPQ P + +PP S P A RAPP
Sbjct: 199 PPPPGPAASAPPRPPPPPPPPPQRPPPPPPPHTHQPEPPTSPPRRQPPAPRAPP 38
Score = 35.0 bits (79), Expect = 0.030
Identities = 24/88 (27%), Positives = 36/88 (40%), Gaps = 5/88 (5%)
Frame = -1
Query: 131 PPFKKPSPSMP-AQPKTDS--PKPPQIKGCQPKNVL--PKADPPKSSPSNAAEAIRAPPN 185
PP P PS P A P++++ P PP +G +P+ L P A P P + + P
Sbjct: 554 PPPVPPPPSPPRAPPQSEAAPPPPPPPRGARPRAPLRPPSAPDPAGHPRAPRQGVAVVPG 375
Query: 186 TAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ H + P P T+P
Sbjct: 374 PRSGNSPRAAHRGPEPRPLPRPPPPTRP 291
Score = 34.3 bits (77), Expect = 0.051
Identities = 28/100 (28%), Positives = 38/100 (38%), Gaps = 9/100 (9%)
Frame = -1
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSP----KPPQIKGCQPKNVLP----KADPPKS-SP 173
P + R PP + P+ P P P KPP +G P P + PP + +P
Sbjct: 329 PRPLPRPPPPTRPPAAPGPGPPPAPGPTPLSKPPAAEGIPPATAPPPRARRLRPPSAPAP 150
Query: 174 SNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
A P T P A + H+ P P+ RP P
Sbjct: 149 PPTPPAAAPPAPTPPHAPA*APHLPPP---PPAPRPPRPP 39
Score = 33.5 bits (75), Expect = 0.088
Identities = 28/95 (29%), Positives = 39/95 (40%), Gaps = 9/95 (9%)
Frame = -3
Query: 128 RYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAP---- 183
RY PP P P+ P P P I+G P P A PP+ P+ A + P
Sbjct: 582 RYIPPPTSPPPASPPAP-VPPQSPSTIRGRAP----PPAPPPRRPPARPPPAPQRPRPRR 418
Query: 184 ----PNTAPMKKALSQHVSAPKKVTP-SKRPATQP 213
P + + A ++ PK TP S+ P+ P
Sbjct: 417 PPARPPSGRRRGARAKVRKLPKGGTPGSRAPSPAP 313
Score = 33.1 bits (74), Expect = 0.11
Identities = 25/82 (30%), Positives = 28/82 (33%), Gaps = 1/82 (1%)
Frame = -2
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P P P P +P PP + P+A PP P A APP P Q
Sbjct: 307 PHPPGPPPPLDRAPPPPPAPLPLASRLRPRASPPPPPPP-PGPAASAPPRPPPPPPPPPQ 131
Query: 196 HVSAPKKV-TPSKRPATQPLRR 216
P T P T P RR
Sbjct: 130 RPPPPPPPHTHQPEPPTSPPRR 65
Score = 32.7 bits (73), Expect = 0.15
Identities = 28/108 (25%), Positives = 42/108 (37%), Gaps = 19/108 (17%)
Frame = -3
Query: 128 RYNPPFKKPSPSMP----AQPKTDS---PKPPQIKGCQPKNVLPKADPPKSSPSNAAEAI 180
R+ PP P PS P +P PP + +P +P P +SP A+
Sbjct: 702 RHPPPPPPPPPSSPPGMGTRPSCQENWWAAPPGARPSRPSRYIP----PPTSPPPASPPA 535
Query: 181 ------------RAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRR 216
RAPP P ++ ++ AP++ P + PA P R
Sbjct: 534 PVPPQSPSTIRGRAPPPAPPPRRPPARPPPAPQRPRPRRPPARPPSGR 391
Score = 27.3 bits (59), Expect = 6.3
Identities = 27/112 (24%), Positives = 39/112 (34%), Gaps = 19/112 (16%)
Frame = -2
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
TP+ R + PSP PA PPQ + + A P PS+ A
Sbjct: 826 TPLDRARACCARRPPSPHRPA-------PPPQAHPARERRTASLARTPPPPPSSPPPAPL 668
Query: 182 APPNT------------APMKKALSQHVS-------APKKVTPSKRPATQPL 214
PP +P +++ S AP + +P RP +PL
Sbjct: 667 LPPRNGNASLLPRKLVGSPARRSSLSSESLHTPAHLAPPRQSPRPRPPPEPL 512
Score = 26.9 bits (58), Expect = 8.2
Identities = 22/75 (29%), Positives = 29/75 (38%)
Frame = -1
Query: 142 AQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPK 201
A P S PP + + +P+A P AA + PP P + V K
Sbjct: 785 ALPAPPSATPPGAPCARAPHSIPRAHP-------AATPLLPPPRPPPPPPEW-ERVPLAK 630
Query: 202 KVTPSKRPATQPLRR 216
K+ RPA PL R
Sbjct: 629 KIGGQPRPALVPLVR 585
>TC79413 similar to GP|21439800|emb|CAD35173. unnamed protein product
{Glycine max}, partial (42%)
Length = 971
Score = 37.0 bits (84), Expect = 0.008
Identities = 26/96 (27%), Positives = 40/96 (41%), Gaps = 9/96 (9%)
Frame = +2
Query: 127 IRYNPPFKKPSPSMP--------AQPKTDSPKPPQIKGCQPKNVLPKADP-PKSSPSNAA 177
++ PP K PS P + P +P PP +K P P + P K+ P +
Sbjct: 278 VKPTPPIVKSPPSPPLVKSPPYQSPPIVKAPSPPLVKPTPPIVKSPPSPPLVKTPPYQSP 457
Query: 178 EAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
++APP P+ K + +P V P P+ P
Sbjct: 458 PIVKAPPTPPPIVK--TPPYQSPPIVKPPVAPSPPP 559
Score = 31.2 bits (69), Expect = 0.44
Identities = 31/118 (26%), Positives = 55/118 (46%), Gaps = 5/118 (4%)
Frame = +2
Query: 102 QNVEVFVEHDDEELPFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKN 161
+++++ V + + P + TP + ++ P P + + P +P PP +K ++
Sbjct: 104 EDLKIVVNYVNPTAPPSPIVTPPSPVKAPPT----PPLVKSPPIVKAPSPPLVKTPPYQS 271
Query: 162 VLPKADPP--KSSPSNAAEAIRAPP-NTAPMKKALSQHVSAPKKVTPS--KRPATQPL 214
K PP KS PS +++PP + P+ KA S + P TP K P + PL
Sbjct: 272 PPVKPTPPIVKSPPS--PPLVKSPPYQSPPIVKAPSPPLVKP---TPPIVKSPPSPPL 430
Score = 28.5 bits (62), Expect = 2.8
Identities = 17/65 (26%), Positives = 25/65 (38%)
Frame = +2
Query: 126 MIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN 185
+++ PP K PS P PP +K + K P +S P +PP
Sbjct: 380 LVKPTPPIVKSPPSPPLVKTPPYQSPPIVKAPPTPPPIVKTPPYQSPPIVKPPVAPSPPP 559
Query: 186 TAPMK 190
T +K
Sbjct: 560 TPIVK 574
>TC91027 similar to GP|14517502|gb|AAK62641.1 At2g38310/T19C21.20
{Arabidopsis thaliana}, partial (55%)
Length = 682
Score = 36.6 bits (83), Expect = 0.010
Identities = 30/86 (34%), Positives = 37/86 (42%), Gaps = 3/86 (3%)
Frame = +2
Query: 131 PPFKKPSPSM-PAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
PP PSP+ P Q S PP N+ P PP S PS+ A I P NT+
Sbjct: 317 PPSPTPSPATTPTQSPPTSVAPPS------SNISPHPSPP-SGPSSVASTIHKPTNTS-S 472
Query: 190 KKALSQHVSAPK--KVTPSKRPATQP 213
K A+S +A V PA+ P
Sbjct: 473 KAAMSSSETATSALSVKSVSSPASPP 550
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 36.6 bits (83), Expect = 0.010
Identities = 24/91 (26%), Positives = 32/91 (34%), Gaps = 6/91 (6%)
Frame = +3
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKS------SPSNAAEAIRAP 183
NPP PS P+ P T S PP P P PP SP + +P
Sbjct: 54 NPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPPTTPAISPPSGGGTTPSP 233
Query: 184 PNTAPMKKALSQHVSAPKKVTPSKRPATQPL 214
P+ P S + K P P++ +
Sbjct: 234 PSRTPPSSDDSPSPPSSKTPPPPSPPSSSSI 326
Score = 36.6 bits (83), Expect = 0.010
Identities = 23/84 (27%), Positives = 32/84 (37%)
Frame = +3
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+PP P+ P P P P P + P A PP S+PSN+ + P +P
Sbjct: 30 SPPPSTPANPPPVTPSAPPPSTPATPSSPPPSTTPSA-PPPSTPSNSPPSPPTTPAISPP 206
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
S P + PS + P
Sbjct: 207 SGG-GTTPSPPSRTPPSSDDSPSP 275
Score = 35.0 bits (79), Expect = 0.030
Identities = 25/78 (32%), Positives = 30/78 (38%)
Frame = +3
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P P+ P+ P +P P V P A P PS A PP+T P
Sbjct: 9 PPPATPSSPPPSTP-------ANPPPVTPSAPP----PSTPATPSSPPPSTTP------- 134
Query: 196 HVSAPKKVTPSKRPATQP 213
SAP TPS P + P
Sbjct: 135 --SAPPPSTPSNSPPSPP 182
>TC81858 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 748
Score = 36.2 bits (82), Expect = 0.014
Identities = 23/80 (28%), Positives = 33/80 (40%), Gaps = 1/80 (1%)
Frame = +1
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQP-KNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALS 194
P+P A P + + PP+ P + PK P S +AEA P +P
Sbjct: 187 PAPKATATPPSSTTTPPKSSATSPTSSPAPKVSSPPSPTPTSAEAPVESPTESP------ 348
Query: 195 QHVSAPKKVTPSKRPATQPL 214
P V+P+ PAT P+
Sbjct: 349 -----PAPVSPTVSPATSPV 393
>BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xylostella
granulovirus}, partial (42%)
Length = 909
Score = 35.4 bits (80), Expect = 0.023
Identities = 29/84 (34%), Positives = 36/84 (42%), Gaps = 1/84 (1%)
Frame = +2
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP +P PS P + +T P+ P K Q KN K +PPK EA PP P +
Sbjct: 11 PPNPQP-PSDPPKKRTPKPETPNRKKPQ-KN---KKEPPKKR-KTPPEASHPPPRPRPPR 172
Query: 191 KALSQHV-SAPKKVTPSKRPATQP 213
H P+K T P T P
Sbjct: 173 PPQPNHQHDKPQKTTHPHHPHTPP 244
Score = 34.3 bits (77), Expect = 0.051
Identities = 23/76 (30%), Positives = 29/76 (37%)
Frame = +1
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
NPP +P P P P+ KPP + PKN P PP +P + PP P
Sbjct: 124 NPPRGQPPPPPPPPPQAPPTKPPTRQ--TPKNNTP--PPPPHTPPDP-----TPPQPPPQ 276
Query: 190 KKALSQHVSAPKKVTP 205
H P+ P
Sbjct: 277 PPKPPHHEKRPRTPEP 324
Score = 30.4 bits (67), Expect = 0.74
Identities = 24/85 (28%), Positives = 29/85 (33%), Gaps = 3/85 (3%)
Frame = +1
Query: 131 PPFKKPSPSMPAQ--PKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
PP + P P + PK ++P PP P P PPK PP P
Sbjct: 160 PPPQAPPTKPPTRQTPKNNTPPPPPHTPPDPTPPQPPPQPPKPPHHEKRPRTPEPPGGRP 339
Query: 189 MKKALSQHVSAPKKVTPSK-RPATQ 212
P TP + RP TQ
Sbjct: 340 -----------PPPTTPKQARPTTQ 381
Score = 28.9 bits (63), Expect = 2.2
Identities = 22/81 (27%), Positives = 30/81 (36%), Gaps = 4/81 (4%)
Frame = +1
Query: 137 SPSMPAQPKTDSPKPPQIKGCQPKNVLPK----ADPPKSSPSNAAEAIRAPPNTAPMKKA 192
+P PA + K Q + QP+ K + K+ P PP AP K
Sbjct: 10 TPQPPAPKRPPKEKDTQARNPQPEEATKKQKRAPEKKKNPPRGQPPPPPPPPPQAPPTKP 189
Query: 193 LSQHVSAPKKVTPSKRPATQP 213
++ PK TP P T P
Sbjct: 190 PTR--QTPKNNTPPPPPHTPP 246
Score = 28.5 bits (62), Expect = 2.8
Identities = 24/85 (28%), Positives = 28/85 (32%), Gaps = 10/85 (11%)
Frame = +1
Query: 135 KPSPSMPAQPKTDSP----------KPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPP 184
K SP P T P PPQ K + P P AA + + PP
Sbjct: 580 KKSPGGRTPPPTPRPGGGGARGAPAPPPQKKQREGGKSGPPRAPGHEEKKPAATSRKRPP 759
Query: 185 NTAPMKKALSQHVSAPKKVTPSKRP 209
A K + KK TP K P
Sbjct: 760 RGAREKNPAAHAREREKKKTPKKNP 834
Score = 28.1 bits (61), Expect = 3.7
Identities = 20/75 (26%), Positives = 29/75 (38%), Gaps = 1/75 (1%)
Frame = +2
Query: 135 KPSPSMPAQPKTDSPKP-PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKAL 193
K P+ P+ +P P P+ +G K P PP P PP P K +
Sbjct: 503 KEGPATHGPPEKKTPPPTPKRRGKHTKRA-PGGGPPPPHPGRGGGGPGGPPPPPPKKNSE 679
Query: 194 SQHVSAPKKVTPSKR 208
+AP+ +KR
Sbjct: 680 RGGRAAPRGRPGTKR 724
Score = 27.3 bits (59), Expect = 6.3
Identities = 18/60 (30%), Positives = 23/60 (38%), Gaps = 2/60 (3%)
Frame = +1
Query: 131 PPFKKPSPSMPA--QPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
PP P P P + + +P+PP G P PK P + A R P AP
Sbjct: 256 PPQPPPQPPKPPHHEKRPRTPEPP--GGRPPPPTTPKQARPTTQERGGARGARPRPPPAP 429
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 35.0 bits (79), Expect = 0.030
Identities = 25/82 (30%), Positives = 34/82 (40%)
Frame = +2
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
++PP SP P+ P P PP C V P PP SSP+ PP+ +P
Sbjct: 239 HSPPPPVYSPPPPSPPPCVEPPPPPPPPC----VEP---PPPSSPAPHQTPYHPPPSPSP 397
Query: 189 MKKALSQHVSAPKKVTPSKRPA 210
+ + S P V S P+
Sbjct: 398 PPSPVYAYPSPPPPVYTSPPPS 463
Score = 33.1 bits (74), Expect = 0.11
Identities = 25/88 (28%), Positives = 31/88 (34%), Gaps = 3/88 (3%)
Frame = +2
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA-- 187
+PP SP+ P P P PP + P P PP SP E PP
Sbjct: 164 SPPPPPHSPTPPVYPYLSPPPPPPVHSPPP----PVYSPPPPSPPPCVEPPPPPPPPCVE 331
Query: 188 -PMKKALSQHVSAPKKVTPSKRPATQPL 214
P + + H P PS P P+
Sbjct: 332 PPPPSSPAPH-QTPYHPPPSPSPPPSPV 412
Score = 28.5 bits (62), Expect = 2.8
Identities = 23/86 (26%), Positives = 31/86 (35%), Gaps = 5/86 (5%)
Frame = +2
Query: 129 YNPPFKKPSPSM-----PAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAP 183
Y+PP P P + P P + P P Q P + P SP A + P
Sbjct: 260 YSPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPP 439
Query: 184 PNTAPMKKALSQHVSAPKKVTPSKRP 209
T+P + + S P V S P
Sbjct: 440 VYTSPPPSPVYAYPSPPPPVYSSPPP 517
Score = 27.3 bits (59), Expect = 6.3
Identities = 25/92 (27%), Positives = 30/92 (32%), Gaps = 9/92 (9%)
Frame = +2
Query: 131 PPFKKPSPSMPAQPKT---------DSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
PP P P PA + SP PP P P PP SP+
Sbjct: 41 PPNSPPPPPPPAPVFSPPPPVPYYYSSPPPPPAHSPPPPPXSP--PPPPHSPTPPVYPYL 214
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+PP P+ S P PS P +P
Sbjct: 215 SPPPPPPVHSPPPPVYSPP---PPSPPPCVEP 301
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.133 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,130,342
Number of Sequences: 36976
Number of extensions: 163927
Number of successful extensions: 3310
Number of sequences better than 10.0: 261
Number of HSP's better than 10.0 without gapping: 1481
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2216
length of query: 267
length of database: 9,014,727
effective HSP length: 94
effective length of query: 173
effective length of database: 5,538,983
effective search space: 958244059
effective search space used: 958244059
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0091b.13


