

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.10
(353 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
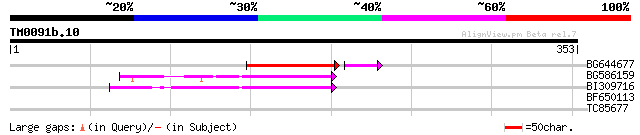
BG644677 weakly similar to GP|7682800|gb| Hypothetical protein T... 65 6e-11
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 57 8e-09
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 52 5e-07
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 35 0.033
TC85677 weakly similar to GP|6457331|gb|AAF09479.1| phytoalexin-... 31 0.82
>BG644677 weakly similar to GP|7682800|gb| Hypothetical protein T15F17.l
{Arabidopsis thaliana}, partial (3%)
Length = 539
Score = 64.7 bits (156), Expect(2) = 6e-11
Identities = 33/59 (55%), Positives = 44/59 (73%), Gaps = 1/59 (1%)
Frame = -3
Query: 148 DISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMSKLNLIGYADAGYLS 205
+I+F+INLL+RYSS+PT RH NG K + +YLKG +DM LF+ +LIGY +A YLS
Sbjct: 507 NITFSINLLSRYSSAPTMRH*NGIKHICKYLKGIIDMGLFYSKDCSPDLIGYVNA*YLS 331
Score = 19.6 bits (39), Expect(2) = 6e-11
Identities = 12/24 (50%), Positives = 13/24 (54%)
Frame = -1
Query: 209 KQVTCLHVEEQPFHGDL*KKL**P 232
KQVT HV + GDL L *P
Sbjct: 308 KQVTYSHVGILSYLGDLRNSLL*P 237
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 57.4 bits (137), Expect = 8e-09
Identities = 38/143 (26%), Positives = 76/143 (52%), Gaps = 8/143 (5%)
Frame = +1
Query: 69 IEYLKDE--VFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*-----EKN 121
+E +++E +++ QR Y+ +L+ F M+KS N++++P P ++N
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKS------------NLSRNPIAPRCKLIKDEN 144
Query: 122 EELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGA 180
+D Y + L+YLA+ TR D+ + ++L++R+ + PT+ H + K+V RYL G
Sbjct: 145 GVKVDA-TKYKQIVGCLMYLAA-TRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGT 318
Query: 181 LDMSLFFPFMSKLNLIGYADAGY 203
+++ + + L Y D+ Y
Sbjct: 319 INLGIMYKRNGSEKLEAYTDSDY 387
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 51.6 bits (122), Expect = 5e-07
Identities = 42/142 (29%), Positives = 66/142 (45%), Gaps = 1/142 (0%)
Frame = +2
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
+ L L + K + + QR Y ++L+ D L V +L + +
Sbjct: 287 YFLGLEVARSKQGILLNQRKYTLELLE----DSG----NLAVKSTLTPYDISLKLHNSDS 442
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNGK-QVFRYLKGAL 181
L + E Y I L+YL + TR DISFA+ L+++ S P Q H+ +V +YLK A
Sbjct: 443 PLYNDETQYRRLIGKLIYLTT-TRPDISFAVQQLSQFVSKPQQVHYQAAIRVLQYLKTAP 619
Query: 182 DMSLFFPFMSKLNLIGYADAGY 203
LF+ S L L +AD+ +
Sbjct: 620 AKGLFYSATSNLKLSSFADSDW 685
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 35.4 bits (80), Expect = 0.033
Identities = 16/62 (25%), Positives = 37/62 (58%), Gaps = 4/62 (6%)
Frame = +1
Query: 146 RLDISFAINLLARYSSSPTQRHW-NGKQVFRYLKGALDMSLFFPFMSK---LNLIGYADA 201
R DI +++++++++ P + H ++ RY++G ++ L FP+ +K LI Y+D+
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDS 300
Query: 202 GY 203
+
Sbjct: 301 DW 306
>TC85677 weakly similar to GP|6457331|gb|AAF09479.1| phytoalexin-deficient 4
protein {Arabidopsis thaliana}, partial (21%)
Length = 2227
Score = 30.8 bits (68), Expect = 0.82
Identities = 18/39 (46%), Positives = 21/39 (53%)
Frame = +2
Query: 184 SLFFPFMSKLNLIGYADAGYLSKYLKQVTCLHVEEQPFH 222
SLFFPF L AGY KY +++TC H PFH
Sbjct: 449 SLFFPFQ----LNPKPGAGYFRKYRRKITCDH---WPFH 544
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.357 0.160 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,487,423
Number of Sequences: 36976
Number of extensions: 137232
Number of successful extensions: 1533
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1153
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 371
Number of HSP's gapped (non-prelim): 1197
length of query: 353
length of database: 9,014,727
effective HSP length: 97
effective length of query: 256
effective length of database: 5,428,055
effective search space: 1389582080
effective search space used: 1389582080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0091b.10


