

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091a.5
(140 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
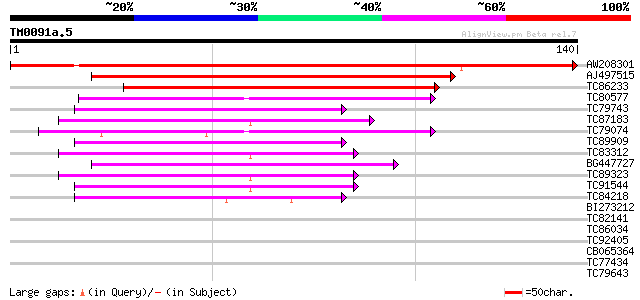
Sequences producing significant alignments: (bits) Value
AW208301 186 3e-48
AJ497515 similar to GP|21553738|gb| unknown {Arabidopsis thalian... 70 4e-13
TC86233 weakly similar to GP|21537084|gb|AAM61425.1 unknown {Ara... 55 7e-09
TC80577 50 3e-07
TC79743 similar to GP|21595139|gb|AAM66075.1 unknown {Arabidopsi... 49 5e-07
TC87183 similar to GP|21700773|gb|AAG38148.1 unknown {Glycine ma... 47 2e-06
TC79074 homologue to GP|21111803|gb|AAM40099.1 glycine rich prot... 47 3e-06
TC89909 similar to PIR|T00577|T00577 hypothetical protein At2g30... 46 4e-06
TC83312 similar to PIR|D96690|D96690 hypothetical protein F28G11... 45 1e-05
BG447727 weakly similar to GP|3152614|gb|A unknown protein {Arab... 43 4e-05
TC89323 similar to GP|21700771|gb|AAG38147.1 unknown {Glycine ma... 43 4e-05
TC91544 similar to GP|21700771|gb|AAG38147.1 unknown {Glycine ma... 42 6e-05
TC84218 similar to GP|15529248|gb|AAK97718.1 At1g29190/F28N24_12... 40 3e-04
BI273212 similar to GP|21553451|gb unknown {Arabidopsis thaliana... 32 0.084
TC82141 similar to PIR|B84889|B84889 hypothetical protein At2g45... 30 0.32
TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C5365... 28 0.93
TC92405 similar to GP|11994325|dbj|BAB02284. gene_id:MLM24.13~un... 27 2.1
CB065364 homologue to PIR|A83022|A83 LPS biosynthesis protein Rf... 27 2.7
TC77434 similar to GP|21554191|gb|AAM63270.1 unknown {Arabidopsi... 27 3.5
TC79643 similar to PIR|C86410|C86410 protein F3M18.18 [imported]... 27 3.5
>AW208301
Length = 434
Score = 186 bits (472), Expect = 3e-48
Identities = 94/141 (66%), Positives = 113/141 (79%), Gaps = 1/141 (0%)
Frame = +3
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MG CAS YA KGG + + VNIV +GKLQQ KEP+KAWHVLSQNPNHF+CSSESMYV
Sbjct: 15 MGSCASNLYASKGGEN-KSTFVNIVLSNGKLQQFKEPIKAWHVLSQNPNHFICSSESMYV 191
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHS-SSVFEPSSS 119
GSP PV+PN+ELQL+HIYFL+P SKS + LSL+DLC+LAIKAN AL + +S+ +PSS
Sbjct: 192 GSPFHPVLPNQELQLDHIYFLLPLSKSNVSLSLQDLCSLAIKANTALVNDPNSMLKPSSV 371
Query: 120 VPQINFKTHPVHSNVSLGYSH 140
NF+T+P+ SNVSLGYSH
Sbjct: 372 SEIRNFQTYPILSNVSLGYSH 434
>AJ497515 similar to GP|21553738|gb| unknown {Arabidopsis thaliana}, partial
(55%)
Length = 685
Score = 69.7 bits (169), Expect = 4e-13
Identities = 34/90 (37%), Positives = 55/90 (60%)
Frame = +1
Query: 21 TVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYF 80
T ++ DG+LQ+ PVK ++L Q P+ F+C+++ M +T V N+ LQ +YF
Sbjct: 34 TAKLILQDGRLQEFTSPVKVSYLLQQYPSSFICNADEMDFDDVVTAVDENQTLQPGQLYF 213
Query: 81 LVPRSKSRLPLSLEDLCALAIKANAALAHS 110
+P S R PL +++ ALA+KA++AL S
Sbjct: 214 ALPLSWLRQPLQAQEMAALAVKASSALMKS 303
>TC86233 weakly similar to GP|21537084|gb|AAM61425.1 unknown {Arabidopsis
thaliana}, partial (21%)
Length = 1129
Score = 55.5 bits (132), Expect = 7e-09
Identities = 23/78 (29%), Positives = 49/78 (62%)
Frame = +3
Query: 29 GKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSR 88
G+++Q +E +KA ++ +NPN+FL +S S+++ + +P+ +EEL+ ++Y P +
Sbjct: 363 GEVKQFREIMKAAELMLENPNYFLVNSRSLHISTRFSPLAADEELEFGNVYIFFPMRRLN 542
Query: 89 LPLSLEDLCALAIKANAA 106
++ D+ L + AN+A
Sbjct: 543 SVVTGADMAVLFLAANSA 596
>TC80577
Length = 640
Score = 50.1 bits (118), Expect = 3e-07
Identities = 24/88 (27%), Positives = 48/88 (54%)
Frame = +1
Query: 18 MQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNH 77
+++ + ++H +G ++ +P+ A V+ PN+F+C+S + + S P+ + +LQ
Sbjct: 19 LKNNIRVIHLNGFVEDFDQPISASQVIGNPPNYFVCTSIQL-LSSSYNPLKGDSQLQPGQ 195
Query: 78 IYFLVPRSKSRLPLSLEDLCALAIKANA 105
+YF++P S S DL LA + A
Sbjct: 196 LYFMLPYSILEDGFSPVDLACLAKRLTA 279
>TC79743 similar to GP|21595139|gb|AAM66075.1 unknown {Arabidopsis
thaliana}, partial (57%)
Length = 976
Score = 49.3 bits (116), Expect = 5e-07
Identities = 24/67 (35%), Positives = 35/67 (51%)
Frame = +2
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLN 76
G + IVH +G ++++ P+ A VL NPNH L S V + + P EL+
Sbjct: 161 GALDLIRIVHLNGYVEEITRPISAGEVLKANPNHVLSKPSSEGVVRRILILSPETELKRG 340
Query: 77 HIYFLVP 83
IYFL+P
Sbjct: 341 SIYFLIP 361
>TC87183 similar to GP|21700773|gb|AAG38148.1 unknown {Glycine max}, partial
(63%)
Length = 1101
Score = 47.4 bits (111), Expect = 2e-06
Identities = 26/79 (32%), Positives = 40/79 (49%), Gaps = 1/79 (1%)
Frame = +2
Query: 13 GGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNE 71
G G + T ++ DG+ +LK P+KA VL +P L SE++ + G + ++
Sbjct: 170 GNTFGSKKTTKVMKIDGETMKLKTPIKAGEVLKDHPGLVLLDSEAVKHYGVRAKVIEAHK 349
Query: 72 ELQLNHIYFLVPRSKSRLP 90
ELQ +YFLV K P
Sbjct: 350 ELQPKRLYFLVELPKETKP 406
>TC79074 homologue to GP|21111803|gb|AAM40099.1 glycine rich protein
{Xanthomonas campestris pv. campestris str. ATCC 33913},
partial (3%)
Length = 1209
Score = 46.6 bits (109), Expect = 3e-06
Identities = 29/105 (27%), Positives = 55/105 (51%), Gaps = 7/105 (6%)
Frame = -2
Query: 8 QYAGKGGNHGMQST-----VNIVHFDGKLQQLKEPVKAWHVLSQN--PNHFLCSSESMYV 60
Q A GG +S+ + +VH G ++ ++P+ V++ P HF+C+S + +
Sbjct: 779 QKAKMGGCFSSRSSCTSKKIRLVHLSGYVEDFEQPISVNQVINSRNPPTHFVCTSLQL-L 603
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANA 105
S P+ + +LQ ++YF++P S + +S DL +LA + A
Sbjct: 602 SSSSKPLKGDTQLQPGNVYFMLPYSILQADVSPVDLASLAKRLTA 468
>TC89909 similar to PIR|T00577|T00577 hypothetical protein At2g30230
[imported] - Arabidopsis thaliana, partial (57%)
Length = 871
Score = 46.2 bits (108), Expect = 4e-06
Identities = 23/67 (34%), Positives = 33/67 (48%)
Frame = +2
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLN 76
G + IVH +G + ++ + A VL NPNH L S V + + P EL+
Sbjct: 140 GALDLIRIVHLNGCVDEITHSITAGEVLQANPNHVLSKPSSQGVVRRILIISPETELKRG 319
Query: 77 HIYFLVP 83
IYFL+P
Sbjct: 320 SIYFLIP 340
>TC83312 similar to PIR|D96690|D96690 hypothetical protein F28G11.8
[imported] - Arabidopsis thaliana, partial (33%)
Length = 975
Score = 45.1 bits (105), Expect = 1e-05
Identities = 25/75 (33%), Positives = 38/75 (50%), Gaps = 1/75 (1%)
Frame = +1
Query: 13 GGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNE 71
G G I+ DG+ ++K P + V+ PNH L S+++ + G P+ PN+
Sbjct: 88 GNTIGKSKKAKIMKIDGETFKVKTPTTSNEVVKNYPNHVLLDSQAVKHFGLRAKPLEPNQ 267
Query: 72 ELQLNHIYFLVPRSK 86
EL+ IYFLV K
Sbjct: 268 ELKPKKIYFLVDLPK 312
>BG447727 weakly similar to GP|3152614|gb|A unknown protein {Arabidopsis
thaliana}, partial (27%)
Length = 349
Score = 43.1 bits (100), Expect = 4e-05
Identities = 21/76 (27%), Positives = 41/76 (53%)
Frame = +2
Query: 21 TVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYF 80
T +V DG +++ VK +VL P+ F+C S+ M + + +E L+L +YF
Sbjct: 119 TSKVVVHDGTMREFTYEVKVGYVLQLYPDCFICDSDEMGFDDVVLAMHEDEVLRLGQLYF 298
Query: 81 LVPRSKSRLPLSLEDL 96
+P ++ + PL + ++
Sbjct: 299 ALPLTRLKKPLPVVEM 346
>TC89323 similar to GP|21700771|gb|AAG38147.1 unknown {Glycine max}, partial
(73%)
Length = 1073
Score = 43.1 bits (100), Expect = 4e-05
Identities = 25/75 (33%), Positives = 40/75 (53%), Gaps = 1/75 (1%)
Frame = +2
Query: 13 GGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNE 71
G + G + + ++ DG+ +LK PVKA VL +P L SE + + G P+ ++
Sbjct: 215 GNSLGGKKSTKVMKIDGETFKLKTPVKAGEVLKDHPGLVLLESEEVKHYGVRAKPLEAHK 394
Query: 72 ELQLNHIYFLVPRSK 86
EL+ +YFLV K
Sbjct: 395 ELKPKRLYFLVELPK 439
>TC91544 similar to GP|21700771|gb|AAG38147.1 unknown {Glycine max}, partial
(34%)
Length = 1081
Score = 42.4 bits (98), Expect = 6e-05
Identities = 23/71 (32%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Frame = +3
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNEELQL 75
G ++ DG+ +LK P KA V+ P H L S+++ + G P+ P+++L+
Sbjct: 129 GRTKKAKVMKIDGETFKLKTPAKANDVVKHYPGHALLDSQAVKHFGLRAKPLEPHQQLKA 308
Query: 76 NHIYFLVPRSK 86
IYFLV K
Sbjct: 309 KKIYFLVELPK 341
>TC84218 similar to GP|15529248|gb|AAK97718.1 At1g29190/F28N24_12
{Arabidopsis thaliana}, partial (57%)
Length = 656
Score = 40.0 bits (92), Expect = 3e-04
Identities = 23/72 (31%), Positives = 38/72 (51%), Gaps = 5/72 (6%)
Frame = +2
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFL--CSSESMYVGSPMTPVV---PNE 71
G + IVH +G+++++ +KA ++ P H L SS S G + +V P+
Sbjct: 101 GALDVIRIVHCNGRVEEISGSIKASEIMKTYPKHVLKKPSSPSTQDGGVVPKIVVVPPDA 280
Query: 72 ELQLNHIYFLVP 83
+LQ IYFL+P
Sbjct: 281 DLQRGKIYFLMP 316
>BI273212 similar to GP|21553451|gb unknown {Arabidopsis thaliana}, partial
(62%)
Length = 663
Score = 32.0 bits (71), Expect = 0.084
Identities = 25/95 (26%), Positives = 40/95 (41%), Gaps = 18/95 (18%)
Frame = +1
Query: 6 STQYAGKGGNHGMQSTVNIV-HFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPM 64
S+ + G + + V ++ H GK+ +L PV A V+ NP H++ S+ + P
Sbjct: 28 SSSFFSMGNCQAVDAAVLVIQHPCGKIDRLYWPVTASEVMKTNPGHYV----SLIMPLPP 195
Query: 65 TP-----------------VVPNEELQLNHIYFLV 82
P + PNE L L H Y L+
Sbjct: 196 QPQEQNQEQKTVRFTRVKLLRPNETLNLGHAYRLI 300
>TC82141 similar to PIR|B84889|B84889 hypothetical protein At2g45320
[imported] - Arabidopsis thaliana, partial (36%)
Length = 885
Score = 30.0 bits (66), Expect = 0.32
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +1
Query: 11 GKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPN 49
G G HGM V +VHF G ++L +++W+ S P+
Sbjct: 289 GAGQFHGMPLDVKVVHFKGSRKRLM--LESWNFYSSTPD 399
>TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C53655)
AU056546(S20671) AU056545(S20671) correspond to a region
of the predicted, partial (42%)
Length = 2412
Score = 28.5 bits (62), Expect = 0.93
Identities = 19/64 (29%), Positives = 31/64 (47%), Gaps = 2/64 (3%)
Frame = -2
Query: 75 LNHIYFLVPRSKS--RLPLSLEDLCALAIKANAALAHSSSVFEPSSSVPQINFKTHPVHS 132
L+ +Y L+ + + RLP L I+ A+ H S + E + +IN +TH H
Sbjct: 1046 LSTLYRLINKERKTFRLPSRL-------IQPQWAIIHLSKLLEKLKQLLRINIRTHITHK 888
Query: 133 NVSL 136
N+ L
Sbjct: 887 NLPL 876
>TC92405 similar to GP|11994325|dbj|BAB02284. gene_id:MLM24.13~unknown
protein {Arabidopsis thaliana}, partial (69%)
Length = 864
Score = 27.3 bits (59), Expect = 2.1
Identities = 22/78 (28%), Positives = 37/78 (47%), Gaps = 8/78 (10%)
Frame = -1
Query: 57 SMYVGSP-MTPVVPNEELQLNHIYFLVPRSKSRL-------PLSLEDLCALAIKANAALA 108
S+ +G P P EE L +I F +P + SRL P S L A + ++
Sbjct: 438 SLPMGGPGRLPPRVREENALLYISFHLPCNLSRLSVRSTNPPASSSSLAAASAFCRSSSL 259
Query: 109 HSSSVFEPSSSVPQINFK 126
+ +F P +++ ++NFK
Sbjct: 258 AARPLFNPETALRRLNFK 205
>CB065364 homologue to PIR|A83022|A83 LPS biosynthesis protein RfaE PA4996
[imported] - Pseudomonas aeruginosa (strain PAO1),
partial (47%)
Length = 682
Score = 26.9 bits (58), Expect = 2.7
Identities = 12/33 (36%), Positives = 20/33 (60%)
Frame = -1
Query: 3 ICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLK 35
+ AS G+GG+ G+ T +I HF G +Q++
Sbjct: 262 VLASRDGHGEGGDDGIAGTGDIEHFTGPRRQVQ 164
>TC77434 similar to GP|21554191|gb|AAM63270.1 unknown {Arabidopsis
thaliana}, partial (71%)
Length = 1362
Score = 26.6 bits (57), Expect = 3.5
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = -1
Query: 56 ESMYVGSPMTPVVPNEELQLNHI 78
+S+ SP+ PV PN +QLN I
Sbjct: 813 DSLTGSSPLQPVQPNRSVQLNRI 745
>TC79643 similar to PIR|C86410|C86410 protein F3M18.18 [imported] -
Arabidopsis thaliana, partial (74%)
Length = 2150
Score = 26.6 bits (57), Expect = 3.5
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = -3
Query: 40 AWHVLSQNPNHFLCSSESMYVGSP 63
+W +L PNHFLC + + GSP
Sbjct: 1851 SWWIL-WTPNHFLCVDKIKHFGSP 1783
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.130 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,247,019
Number of Sequences: 36976
Number of extensions: 83599
Number of successful extensions: 454
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 445
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 449
length of query: 140
length of database: 9,014,727
effective HSP length: 86
effective length of query: 54
effective length of database: 5,834,791
effective search space: 315078714
effective search space used: 315078714
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0091a.5


