

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089a.1
(119 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
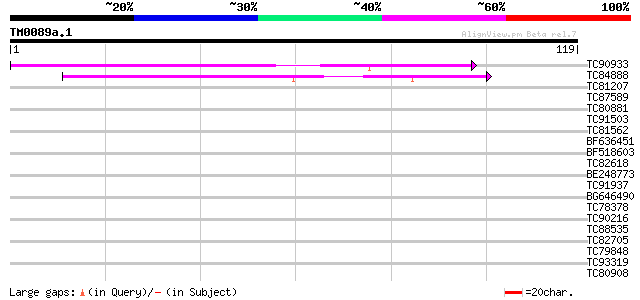
Score E
Sequences producing significant alignments: (bits) Value
TC90933 similar to SP|P27484|GRP2_NICSY Glycine-rich protein 2. ... 58 5e-10
TC84888 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 43 2e-05
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 36 0.002
TC87589 similar to GP|20268752|gb|AAM14079.1 unknown protein {Ar... 28 0.66
TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor ... 27 0.87
TC91503 similar to GP|20161013|dbj|BAB89946. putative ovule deve... 26 1.9
TC81562 weakly similar to GP|6671946|gb|AAF23206.1| putative ami... 25 3.3
BF636451 25 3.3
BF518603 similar to GP|4176424|dbj| ribosomal protein L12 {Oryza... 25 3.3
TC82618 similar to GP|9294614|dbj|BAB02953.1 DNA/RNA binding pro... 25 4.3
BE248773 similar to GP|22136352|gb| unknown protein {Arabidopsis... 25 4.3
TC91937 similar to GP|22136032|gb|AAM91598.1 GTP-binding protein... 25 4.3
BG646490 similar to GP|12323094|g unknown protein; 5864-31259 {A... 25 4.3
TC78378 similar to GP|10177426|dbj|BAB10711. GATA-binding transc... 25 5.6
TC90216 similar to GP|14326523|gb|AAK60306.1 AT4g20330/F9F13_2 {... 25 5.6
TC88535 weakly similar to GP|6686784|emb|CAB64719.1 AKIN beta2 {... 25 5.6
TC82705 similar to GP|22022522|gb|AAM83219.1 AT4g03080/T4I9_4 {A... 24 7.3
TC79848 similar to GP|10954048|gb|AAG25716.1 ovarian fibroin-lik... 24 7.3
TC93319 similar to GP|9759550|dbj|BAB11152.1 receptor protein ki... 24 9.6
TC80908 similar to GP|4966372|gb|AAD34703.1| ESTs gb|N38586 and ... 24 9.6
>TC90933 similar to SP|P27484|GRP2_NICSY Glycine-rich protein 2. [Wood
tobacco] {Nicotiana sylvestris}, partial (44%)
Length = 368
Score = 58.2 bits (139), Expect = 5e-10
Identities = 41/106 (38%), Positives = 55/106 (51%), Gaps = 8/106 (7%)
Frame = +3
Query: 1 WMMSVSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTI 60
++MS S + K+K F+DQKGF IT DDG +LF+ S+I++ GF LAE E V+
Sbjct: 51 FIMSGSKLTGKVKWFNDQKGFGFITPDDGSEELFVHQSQIQTDGFRSLAEGESVEY---- 218
Query: 61 LIPMAALRR*RD*R--------PD*API*GTCCGGNDSSGYGRGGG 98
+ D R PD A + G+ GG GY RGGG
Sbjct: 219 -----QIESDNDGRSKAVSVTGPDGASVQGSRRGGGGGGGYERGGG 341
>TC84888 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (58%)
Length = 612
Score = 42.7 bits (99), Expect = 2e-05
Identities = 35/103 (33%), Positives = 47/103 (44%), Gaps = 13/103 (12%)
Frame = +2
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL-------TILIPM 64
+K F+ KGF I+ D GG DLF+ + IK+ GF LAE E V+ ++ T I +
Sbjct: 71 VKWFNSTKGFGFISPDGGGEDLFVHQTSIKTDGFRSLAEGEAVEFIVEAGDDGRTKAIDV 250
Query: 65 AALRR*RD*RPD*API*GT------CCGGNDSSGYGRGGGYQE 101
P+ AP G GG G G GGGY +
Sbjct: 251 TG--------PNGAPPQGAPRQEMGYGGGGGRGGGGYGGGYDQ 355
Score = 24.6 bits (52), Expect = 5.6
Identities = 9/16 (56%), Positives = 10/16 (62%)
Frame = +2
Query: 86 GGNDSSGYGRGGGYQE 101
GG GYG GGGY +
Sbjct: 425 GGYXQGGYGGGGGYXQ 472
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 35.8 bits (81), Expect = 0.002
Identities = 18/44 (40%), Positives = 27/44 (60%)
Frame = +3
Query: 15 FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL 58
F++ KGF I DD DLF+ S I+S+GF L+E + V+ +
Sbjct: 75 FNNTKGFGFIKPDDDSEDLFVHQSDIRSEGFRSLSEGDRVEFTI 206
Score = 24.6 bits (52), Expect = 5.6
Identities = 7/14 (50%), Positives = 10/14 (71%)
Frame = +2
Query: 105 IRIWRNGRQKRWRW 118
+ WR +Q+RWRW
Sbjct: 500 LHAWRR*QQQRWRW 541
>TC87589 similar to GP|20268752|gb|AAM14079.1 unknown protein {Arabidopsis
thaliana}, partial (81%)
Length = 1615
Score = 27.7 bits (60), Expect = 0.66
Identities = 14/38 (36%), Positives = 19/38 (49%), Gaps = 2/38 (5%)
Frame = -2
Query: 83 TCCGGNDSSGYGRGGGYQEVVDIRIWRNGRQ--KRWRW 118
T G + GYGR G +E V +R+W +RW W
Sbjct: 330 TTVSGCINGGYGRTGS-EEAVGMRMWTTAGDDGERWEW 220
>TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (34%)
Length = 597
Score = 27.3 bits (59), Expect = 0.87
Identities = 16/42 (38%), Positives = 19/42 (45%), Gaps = 9/42 (21%)
Frame = -3
Query: 86 GGNDSSGYGRGGGYQ---------EVVDIRIWRNGRQKRWRW 118
GG D YG GG + V + IWR R +RWRW
Sbjct: 367 GGGDL*TYGGGGEGRWWRWRFVVIRWVFVDIWRWWRWRRWRW 242
>TC91503 similar to GP|20161013|dbj|BAB89946. putative ovule development
protein aintegumenta-like protein {Oryza sativa
(japonica cultivar-group), partial (27%)
Length = 1057
Score = 26.2 bits (56), Expect = 1.9
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = +2
Query: 86 GGNDSSGYGRGGGYQEVVDIRIWRNGRQ 113
GG D GYG GGY ++ + I +G Q
Sbjct: 905 GGGDHGGYGGNGGY--MIPMAIANDGNQ 982
>TC81562 weakly similar to GP|6671946|gb|AAF23206.1| putative amino acid
transporter protein {Arabidopsis thaliana}, partial
(36%)
Length = 673
Score = 25.4 bits (54), Expect = 3.3
Identities = 12/36 (33%), Positives = 19/36 (52%), Gaps = 5/36 (13%)
Frame = +1
Query: 88 NDSSGYGRGGGYQ-----EVVDIRIWRNGRQKRWRW 118
ND SG+G+ + + + ++R G KRWRW
Sbjct: 40 NDDSGFGKKSSFSGN*LTR*IWVVVYRTGN-KRWRW 144
>BF636451
Length = 648
Score = 25.4 bits (54), Expect = 3.3
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = -3
Query: 95 RGGGYQEVVDIRIWRNGRQKRWRW 118
+G G+QE+ ++IW G+ KR W
Sbjct: 334 QGNGFQELAKVKIWL-GQTKRKPW 266
>BF518603 similar to GP|4176424|dbj| ribosomal protein L12 {Oryza sativa
(japonica cultivar-group)}, partial (25%)
Length = 318
Score = 25.4 bits (54), Expect = 3.3
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = -3
Query: 86 GGNDSSGYGRGGG 98
GG++ SGYG GGG
Sbjct: 172 GGSEGSGYGGGGG 134
>TC82618 similar to GP|9294614|dbj|BAB02953.1 DNA/RNA binding protein-like
{Arabidopsis thaliana}, partial (35%)
Length = 850
Score = 25.0 bits (53), Expect = 4.3
Identities = 9/15 (60%), Positives = 10/15 (66%)
Frame = -3
Query: 84 CCGGNDSSGYGRGGG 98
CCGG + S G GGG
Sbjct: 209 CCGGGNVSECGEGGG 165
>BE248773 similar to GP|22136352|gb| unknown protein {Arabidopsis
thaliana}, partial (19%)
Length = 436
Score = 25.0 bits (53), Expect = 4.3
Identities = 10/15 (66%), Positives = 12/15 (79%), Gaps = 2/15 (13%)
Frame = -1
Query: 84 CCGGND--SSGYGRG 96
CCGG+D + GYGRG
Sbjct: 76 CCGGSDG*TIGYGRG 32
>TC91937 similar to GP|22136032|gb|AAM91598.1 GTP-binding protein obg-like
{Arabidopsis thaliana}, partial (45%)
Length = 1011
Score = 25.0 bits (53), Expect = 4.3
Identities = 15/45 (33%), Positives = 19/45 (41%), Gaps = 18/45 (40%)
Frame = +1
Query: 93 YGRGGGYQEVVDIRI-------WRNGRQKR-----------WRWF 119
+GRGGG+ + V R+ W RQ R WRWF
Sbjct: 172 FGRGGGHMDKVGCRLVLRGMMWW*RCRQGR*LGRLVGRR*FWRWF 306
>BG646490 similar to GP|12323094|g unknown protein; 5864-31259 {Arabidopsis
thaliana}, partial (4%)
Length = 702
Score = 25.0 bits (53), Expect = 4.3
Identities = 11/35 (31%), Positives = 20/35 (56%)
Frame = +2
Query: 85 CGGNDSSGYGRGGGYQEVVDIRIWRNGRQKRWRWF 119
C + +GY G G +D+R+ +GR+++ WF
Sbjct: 590 CCKHQGNGYSEGSGIF*YLDLRL--SGRREKGCWF 688
>TC78378 similar to GP|10177426|dbj|BAB10711. GATA-binding transcription
factor-like protein {Arabidopsis thaliana}, partial
(23%)
Length = 1284
Score = 24.6 bits (52), Expect = 5.6
Identities = 7/17 (41%), Positives = 13/17 (76%)
Frame = -1
Query: 100 QEVVDIRIWRNGRQKRW 116
+ V+ +R W+N R++RW
Sbjct: 471 ERVIRLRKWKNQRKRRW 421
>TC90216 similar to GP|14326523|gb|AAK60306.1 AT4g20330/F9F13_2 {Arabidopsis
thaliana}, partial (37%)
Length = 743
Score = 24.6 bits (52), Expect = 5.6
Identities = 9/14 (64%), Positives = 11/14 (78%)
Frame = -2
Query: 104 DIRIWRNGRQKRWR 117
D++I RNGR K WR
Sbjct: 103 DVKIKRNGRLKTWR 62
>TC88535 weakly similar to GP|6686784|emb|CAB64719.1 AKIN beta2 {Arabidopsis
thaliana}, partial (30%)
Length = 759
Score = 24.6 bits (52), Expect = 5.6
Identities = 9/25 (36%), Positives = 12/25 (48%)
Frame = -1
Query: 85 CGGNDSSGYGRGGGYQEVVDIRIWR 109
CGGN G ++E V + WR
Sbjct: 612 CGGNSGGGLLNSSSFKEYVIVTRWR 538
>TC82705 similar to GP|22022522|gb|AAM83219.1 AT4g03080/T4I9_4 {Arabidopsis
thaliana}, partial (26%)
Length = 849
Score = 24.3 bits (51), Expect = 7.3
Identities = 13/34 (38%), Positives = 16/34 (46%)
Frame = +2
Query: 85 CGGNDSSGYGRGGGYQEVVDIRIWRNGRQKRWRW 118
CGG DSSG + Y + + RNG W W
Sbjct: 296 CGGRDSSGTPQADAY----GLLMHRNG---TWEW 376
>TC79848 similar to GP|10954048|gb|AAG25716.1 ovarian fibroin-like
substance-1 {Cyprinus carpio}, partial (5%)
Length = 424
Score = 24.3 bits (51), Expect = 7.3
Identities = 10/20 (50%), Positives = 11/20 (55%)
Frame = +3
Query: 82 GTCCGGNDSSGYGRGGGYQE 101
G+ G GYG GGGY E
Sbjct: 15 GSGQGSGYGEGYGHGGGYGE 74
>TC93319 similar to GP|9759550|dbj|BAB11152.1 receptor protein kinase-like
protein {Arabidopsis thaliana}, partial (10%)
Length = 646
Score = 23.9 bits (50), Expect = 9.6
Identities = 6/13 (46%), Positives = 10/13 (76%)
Frame = +3
Query: 107 IWRNGRQKRWRWF 119
+W+ R KR++WF
Sbjct: 258 LWKTSRSKRFKWF 296
>TC80908 similar to GP|4966372|gb|AAD34703.1| ESTs gb|N38586 and gb|N38613
come from this gene. {Arabidopsis thaliana}, partial
(81%)
Length = 1442
Score = 23.9 bits (50), Expect = 9.6
Identities = 7/10 (70%), Positives = 9/10 (90%)
Frame = +2
Query: 110 NGRQKRWRWF 119
N R++RWRWF
Sbjct: 104 NHRRRRWRWF 133
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.359 0.166 0.627
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,547,621
Number of Sequences: 36976
Number of extensions: 41847
Number of successful extensions: 829
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 762
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 828
length of query: 119
length of database: 9,014,727
effective HSP length: 95
effective length of query: 24
effective length of database: 5,502,007
effective search space: 132048168
effective search space used: 132048168
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 50 (23.9 bits)
Lotus: description of TM0089a.1


