

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0087.21
(1215 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
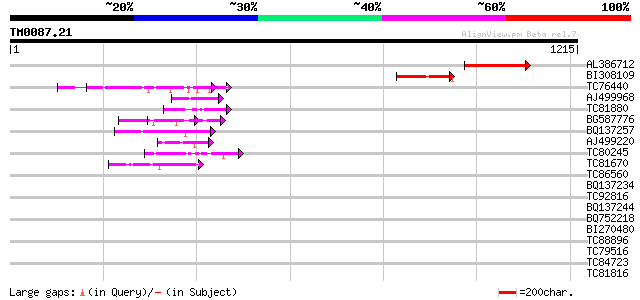
Score E
Sequences producing significant alignments: (bits) Value
AL386712 248 8e-66
BI308109 116 6e-26
TC76440 weakly similar to SP|Q9LD55|IF3A_ARATH Eukaryotic transl... 58 2e-08
AJ499968 weakly similar to GP|23495961|gb| hypothetical protein ... 50 7e-06
TC81880 similar to PIR|B85431|B85431 trichohyalin like protein [... 47 6e-05
BG587776 weakly similar to PIR|F86242|F86 unknown protein 98896... 46 8e-05
BQ137257 similar to GP|21536912|gb| unknown {Arabidopsis thalian... 45 2e-04
AJ499220 similar to SP|Q09863|YAF Hypothetical protein C29E6.10c... 44 3e-04
TC80245 similar to PIR|T48581|T48581 hypothetical protein T31B5.... 44 4e-04
TC81670 weakly similar to PIR|E86293|E86293 hypothetical protein... 44 5e-04
TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-... 42 0.001
BQ137234 similar to EGAD|35719|3709 hypothetical protein {Burkho... 42 0.002
TC92816 weakly similar to GP|18076679|emb|CAC84774. P70 protein ... 40 0.004
BQ137244 similar to GP|20161222|dbj Epstein-Barr virus EBNA-1-li... 40 0.004
BQ752218 similar to GP|19170914|emb hypothetical protein {Enceph... 40 0.005
BI270480 weakly similar to GP|9757727|db contains similarity to ... 39 0.016
TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown prot... 39 0.016
TC79516 similar to GP|8777406|dbj|BAA96996.1 gb|AAF13095.1~gene_... 39 0.016
TC84723 similar to GP|16945387|emb|CAD11800. conserved hypotheti... 37 0.036
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 37 0.046
>AL386712
Length = 445
Score = 248 bits (634), Expect = 8e-66
Identities = 124/145 (85%), Positives = 131/145 (89%), Gaps = 3/145 (2%)
Frame = +1
Query: 975 LKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESLSLLGHFALFHPGNQAVLRWGKSPT- 1033
LKMEIFHLM F+LSHCASRWK+PNDQVG LMLESLSLLGHFALFHPGNQAVLRWGKSPT
Sbjct: 1 LKMEIFHLMGFLLSHCASRWKSPNDQVGLLMLESLSLLGHFALFHPGNQAVLRWGKSPTP 180
Query: 1034 -ILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQQELSVDMLLSLLRSCKSA 1092
ILHKVCDLPF+FFSDPELMP+LAGTLVAACYGCEQNKF+VQQELSVDMLLSLLRSC++A
Sbjct: 181 TILHKVCDLPFVFFSDPELMPLLAGTLVAACYGCEQNKFMVQQELSVDMLLSLLRSCRNA 360
Query: 1093 APATQL-ITLDNSATDESSEPNQLG 1116
APATQL DN TDES NQ G
Sbjct: 361 APATQLNSNFDNIPTDESIGSNQSG 435
>BI308109
Length = 430
Score = 116 bits (290), Expect = 6e-26
Identities = 74/136 (54%), Positives = 85/136 (62%), Gaps = 11/136 (8%)
Frame = +2
Query: 829 LDETIKVNRNEESISIGKDCKLEHDSSDKLKNDEMEKIDDLDEPQKNQGGDIANSFVSQK 888
LDE KVNR+E+S +IGK C+ EHD+S KL N++ EKI DE QKNQ DIA S +SQ+
Sbjct: 2 LDEISKVNRDEKSFAIGKGCESEHDASVKLNNNDTEKIASSDESQKNQNEDIATSVISQR 181
Query: 889 DEKHTMVNITTQKNEKSSNLAQPVVFLLSALSETGL-VSLPSLLTAVLLQANNRSSSEP- 946
DEKH T QKNEK S LAQPV FLLSA+SET L VSL L Q N + P
Sbjct: 182 DEKH-----TAQKNEKESILAQPVAFLLSAVSETDLSVSLLF*LLFFCRQTINHPLNRPP 346
Query: 947 ---------VATGVLK 953
VATGVLK
Sbjct: 347 LSCHLILKEVATGVLK 394
>TC76440 weakly similar to SP|Q9LD55|IF3A_ARATH Eukaryotic translation
initiation factor 3 subunit 10 (eIF-3 theta), partial
(41%)
Length = 1953
Score = 58.2 bits (139), Expect = 2e-08
Identities = 91/416 (21%), Positives = 177/416 (41%), Gaps = 43/416 (10%)
Frame = +1
Query: 103 ESLLTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDKAPTEVINEKNPRSTDN 162
+ L+ +VD K + ++L ++ HL S A++L N
Sbjct: 193 QKFLSMKVDHMKNVVIFCKTSLEADGLRDHLASFAEQL---------------------N 309
Query: 163 LRRQMQLPEKDKEKRSTSLGKSMNAWKEKRNWEDILSSPFRVSSRMSYSPSLGRKSAERV 222
RQM P K+ + +L +++ E + R+ +R S + ++ E+
Sbjct: 310 KARQMICPPDRKQSKLGALLPTLS--------EVVAKEHKRLLARKSI---IEKRKEEQE 456
Query: 223 RTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWH 282
R L +K E K+ + +EAE++ R+ +E+E ++Q+LQR ++ +R E
Sbjct: 457 RQLLEKEREEESKRLRLLKIDEEAEQR-----RLATEIEQRKIQRLQREKEERDR-EEAE 618
Query: 283 AVRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQ 342
A+R L+ + R +R + Q+++ + + + V +VR + ++ +KL
Sbjct: 619 ALR-LEAEK----RLKRKGKKPVIEGGQISRESLMQLTLVEQVRERQEMEKKLQKLAKTM 783
Query: 343 KLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAE-----------KLQRLAEMQ 391
H +R E +I + ++ L E + ER + ++ E + +RL+ M
Sbjct: 784 D-HLERAKREEAAPLIDAAYQQRLVEERVLHEREQQLQVELSRQRHAGDLNEKERLSRMM 960
Query: 392 RKKEEAQIR--------------------------REEERKASSAAREARAIEQLRR--- 422
KE Q R R++ER+ + +E+ +R
Sbjct: 961 GNKEIYQERVVSHRQAEFNRLQRDRLDRISKILLSRKQEREKMRKLKYYLKVEEEKRQKL 1140
Query: 423 KEERAKAQQEEAELLAQKLAER---LNESEQRRKIYLEQIRERANLRDQSSPLLRR 475
+EE ++EEAE ++ AER L+E +++K L +I E+A R++ LL R
Sbjct: 1141REEEEAREREEAERKKKEQAERQAKLDEIAEKQKQRLREIEEKAE-REKREALLGR 1305
Score = 51.2 bits (121), Expect = 2e-06
Identities = 65/301 (21%), Positives = 130/301 (42%), Gaps = 22/301 (7%)
Frame = +1
Query: 165 RQMQLPEKDKEKRSTSLGKSMNAWKEKRNWEDILSSPFRVSSRMSYSPSLGRKSAERVRT 224
RQ+ E+++E + L K +++R +I ++ R+ AE +R
Sbjct: 457 RQLLEKEREEESKRLRLLKIDEEAEQRRLATEIEQR--KIQRLQREKEERDREEAEALRL 630
Query: 225 LHDKLMSPEKKKKTTS--DLKKEAEEKHARALRIRSELETER-VQKLQRTSQKLNRVSEW 281
+K + + KK + +E+ + ++R E E+ +QKL +T L R
Sbjct: 631 EAEKRLKRKGKKPVIEGGQISRESLMQLTLVEQVRERQEMEKKLQKLAKTMDHLERAKRE 810
Query: 282 HAVRHLKLREGMYAR--------HQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNE 333
A L + Y + H+R + Q + AGD + K R + +
Sbjct: 811 EAA---PLIDAAYQQRLVEERVLHEREQQLQVELSRQ--RHAGDLNEKERLSRMMGNKEI 975
Query: 334 ENKKLILRQ-----KLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLA 388
++++ + +L L R K+ + + +++E + + + L+ +E EK Q+L
Sbjct: 976 YQERVVSHRQAEFNRLQRDRLDRISKILLSRKQEREKMRKLKYYLK----VEEEKRQKL- 1140
Query: 389 EMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEER------AKAQQEEAELLAQKLA 442
R++EEA+ R E ERK A ++++ K+++ KA++E+ E L + A
Sbjct: 1141---REEEEAREREEAERKKKEQAERQAKLDEIAEKQKQRLREIEEKAEREKREALLGRPA 1311
Query: 443 E 443
E
Sbjct: 1312E 1314
>AJ499968 weakly similar to GP|23495961|gb| hypothetical protein {Plasmodium
falciparum 3D7}, partial (19%)
Length = 342
Score = 49.7 bits (117), Expect = 7e-06
Identities = 32/112 (28%), Positives = 63/112 (55%), Gaps = 1/112 (0%)
Frame = -3
Query: 348 ELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKA 407
E E+ + + +++E +A+E+ ER++ E E+L+ L + ++KEE + + +EER+
Sbjct: 340 EKEERERKEREEREEQERIAKEKEEQERKEREERERLE-LERIAKEKEERERKEKEEREE 164
Query: 408 SSAAREARAIEQLRRK-EERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQ 458
IE+ R++ EE+A +QEE LL + + ++ E E + K E+
Sbjct: 163 RERLEAVVRIEKERKEAEEKALKEQEEKALLKEGVIDKKEEEEPKSKATPEE 8
Score = 41.6 bits (96), Expect = 0.002
Identities = 31/123 (25%), Positives = 67/123 (54%)
Frame = -3
Query: 369 EEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAK 428
E+ ER++ E E+ +R+A ++KEE + + EER+ R A+ E+ RKE+ +
Sbjct: 340 EKEERERKEREEREEQERIA---KEKEEQERKEREERERLELERIAKEKEERERKEKEER 170
Query: 429 AQQEEAELLAQKLAERLNESEQRRKIYLEQIRERANLRDQSSPLLRRSLNKDGQGRSTPN 488
++E E + + ER E+E++ L++ E+A L++ ++ + ++ + ++TP
Sbjct: 169 EERERLEAVVRIEKER-KEAEEKA---LKEQEEKALLKE---GVIDKKEEEEPKSKATPE 11
Query: 489 NSS 491
+
Sbjct: 10 EEA 2
>TC81880 similar to PIR|B85431|B85431 trichohyalin like protein [imported] -
Arabidopsis thaliana, partial (4%)
Length = 1043
Score = 46.6 bits (109), Expect = 6e-05
Identities = 41/150 (27%), Positives = 78/150 (51%), Gaps = 5/150 (3%)
Frame = +2
Query: 330 SLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAE 389
S++++++K + ++L LR+ E+L+ + +QK+ +A + A+LE + E
Sbjct: 428 SVDKDDEKQKIERELEKERLRKIEELERERERQKDRMAVDRAMLEAER-----------E 574
Query: 390 MQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLA---ERLN 446
+R+K+ A + R A+ AR+ RA + R + ERA + AE + LA ERL
Sbjct: 575 REREKDRAAVDR-----ATFEARD-RAYAEGRERAERAAFDRATAEARQRALAEAKERLE 736
Query: 447 E--SEQRRKIYLEQIRERANLRDQSSPLLR 474
+ +E R K Y ++ A L+ + + + R
Sbjct: 737 KACAEARDKSYTDKATAEARLKAERAAVER 826
>BG587776 weakly similar to PIR|F86242|F86 unknown protein 98896-95855
[imported] - Arabidopsis thaliana, partial (23%)
Length = 608
Score = 46.2 bits (108), Expect = 8e-05
Identities = 42/177 (23%), Positives = 82/177 (45%), Gaps = 5/177 (2%)
Frame = +2
Query: 234 KKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGM 293
++ +T+ ++ + A+ R + + + +S L V + A+ + E
Sbjct: 86 RRSPSTTTWRRRSRSPTAKRYRRQRSRSSSLSPAHKSSSSSLGSVEQKTAIEKQRKEEEK 265
Query: 294 YARHQRSESRHEAF-----LAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSE 348
R Q +E + A + + ++ DES EVR E + L ++ ++
Sbjct: 266 KRRQQEAELKLIAEETAKRVEEAIRKRVDESLSSEEVRVEIQRRLEEGRKKLNDEV-AAQ 442
Query: 349 LRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEER 405
L++ ++ +I++KQKE+ AR+E R L E +++ E QR++ Q RREEER
Sbjct: 443 LQKEKEAALIEAKQKEEQARKEKEDLERML--EENRRKIEESQRREALEQQRREEER 607
Score = 43.5 bits (101), Expect = 5e-04
Identities = 43/172 (25%), Positives = 85/172 (49%), Gaps = 6/172 (3%)
Frame = +2
Query: 296 RHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRR-AEK 354
R QRS S + ++ SS + V T++ ++ K+ +++ +EL+ AE+
Sbjct: 149 RRQRSRS------SSLSPAHKSSSSSLGSVEQKTAIEKQRKEEEKKRRQQEAELKLIAEE 310
Query: 355 L-----QVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASS 409
+ I+ + E L+ EE +E +QR E RKK ++ + +++ +
Sbjct: 311 TAKRVEEAIRKRVDESLSSEEVRVE---------IQRRLEEGRKKLNDEVAAQLQKEKEA 463
Query: 410 AAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRE 461
A EA ++KEE+A+ ++E+ E + ++ ++ ES QRR+ +Q RE
Sbjct: 464 ALIEA------KQKEEQARKEKEDLERMLEENRRKIEES-QRREALEQQRRE 598
Score = 35.0 bits (79), Expect = 0.18
Identities = 27/101 (26%), Positives = 54/101 (52%), Gaps = 8/101 (7%)
Frame = +2
Query: 388 AEMQRKKEEAQIRREEERKASSAA----REARAIEQLRRKEERAKAQQE-EAELLAQKLA 442
A+ R++ K+SS++ + AIE+ R++EE+ + QQE E +L+A++ A
Sbjct: 137 AKRYRRQRSRSSSLSPAHKSSSSSLGSVEQKTAIEKQRKEEEKKRRQQEAELKLIAEETA 316
Query: 443 ERLNESEQRR---KIYLEQIRERANLRDQSSPLLRRSLNKD 480
+R+ E+ ++R + E++R R + R+ LN +
Sbjct: 317 KRVEEAIRKRVDESLSSEEVRVEIQRRLEEG---RKKLNDE 430
>BQ137257 similar to GP|21536912|gb| unknown {Arabidopsis thaliana}, partial
(15%)
Length = 1064
Score = 44.7 bits (104), Expect = 2e-04
Identities = 55/229 (24%), Positives = 94/229 (41%), Gaps = 11/229 (4%)
Frame = +2
Query: 224 TLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHA 283
++ L PEKKKK + + + R R+ TER + + S
Sbjct: 293 SIQPPLPVPEKKKKKKNKGTGQERTRREEDRRSRTHRTTERTTPRRGGNGPAQEASSER- 469
Query: 284 VRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQK 343
R RE RHQ+S R A + + +R+ E S + R T +E + R+
Sbjct: 470 -RRRTRRESGGTRHQQSAGRRHAIVPRGRRRSAAEESGQSSGRTETR-GDEWQHGRRREG 643
Query: 344 LHGSELRRAEKLQV--IKSKQKEDLAREEAVLER--------RKLIEAEKLQRLAEMQRK 393
E R E L +KQ + R+ R R + +R +QR+
Sbjct: 644 QDTDETGRREALNAAPTDAKQPQHEGRDTTAAARAESSATTSRDYTQTHARKRSDTIQRE 823
Query: 394 KEEAQIRREEERKASSAAREARAIE-QLRRKEERAKAQQEEAELLAQKL 441
+ Q +++E+RK ++ R+ E Q RR ERA +Q ++ + A+++
Sbjct: 824 QRSTQRQQDEKRKTANRRDGRRSAE*QTRRHTERA-SQSRDSTIRARRV 967
>AJ499220 similar to SP|Q09863|YAF Hypothetical protein C29E6.10c in
chromosome I. [Fission yeast] {Schizosaccharomyces
pombe}, partial (12%)
Length = 518
Score = 44.3 bits (103), Expect = 3e-04
Identities = 41/137 (29%), Positives = 68/137 (48%), Gaps = 17/137 (12%)
Frame = -3
Query: 316 GDESSKVNEVRFITSLNEENKKL--ILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVL 373
G+E ++ E R EE +++ I ++ + A + +V + +Q++ L E +
Sbjct: 402 GEEDNQTEEQRM-----EEGRRMFQIFAARMFEQRVLNAYREKVAQERQQKLL---EELE 247
Query: 374 ERRKLIEAEKLQRLAEMQRKK----------EEAQIRREEER----KASSAAREARAIEQ 419
E +L E +L++L E +RKK EE + RRE ER KA A RE + E+
Sbjct: 246 EENRLKEERELKKLKEKERKKAKNRALKQQKEEERARREAERLAEEKAQRAEREKKLEEE 67
Query: 420 -LRRKEERAKAQQEEAE 435
RR+EER K + E +
Sbjct: 66 RKRREEERLKKEAERRQ 16
Score = 42.0 bits (97), Expect = 0.001
Identities = 32/105 (30%), Positives = 61/105 (57%), Gaps = 4/105 (3%)
Frame = -3
Query: 368 REEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERA 427
RE+ ER++ + L+ L E R KEE ++++ +E++ A RA++Q +++EERA
Sbjct: 297 REKVAQERQQKL----LEELEEENRLKEERELKKLKEKERKKAKN--RALKQ-QKEEERA 139
Query: 428 KAQQEEAELLAQKLAERLNE----SEQRRKIYLEQIRERANLRDQ 468
+ EAE LA++ A+R E+R++ E++++ A R +
Sbjct: 138 R---REAERLAEEKAQRAEREKKLEEERKRREEERLKKEAERRQR 13
Score = 38.9 bits (89), Expect = 0.012
Identities = 32/120 (26%), Positives = 60/120 (49%), Gaps = 2/120 (1%)
Frame = -3
Query: 307 FLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQ--KLHGSELRRAEKLQVIKSKQKE 364
F +V ++ ++ + + + L EEN+ R+ KL E ++A K + +K +++E
Sbjct: 324 FEQRVLNAYREKVAQERQQKLLEELEEENRLKEERELKKLKEKERKKA-KNRALKQQKEE 148
Query: 365 DLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKE 424
+ AR EA +RLAE + ++ E + + EEERK R + E+ +RK+
Sbjct: 147 ERARREA-------------ERLAEEKAQRAEREKKLEEERKRREEERLKKEAERRQRKK 7
>TC80245 similar to PIR|T48581|T48581 hypothetical protein T31B5.160 -
Arabidopsis thaliana, partial (60%)
Length = 895
Score = 43.9 bits (102), Expect = 4e-04
Identities = 53/217 (24%), Positives = 102/217 (46%), Gaps = 6/217 (2%)
Frame = +2
Query: 290 REGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSEL 349
R + R +R SR + L+ R+ + K N+ +LN ++ K ++L E
Sbjct: 143 RRSRHHRRRRHRSRSSSSLSP-PPRSRTPTLKHNKKDHPPTLNNKSSKRQQDEELKLLEE 319
Query: 350 RRAEKLQ-VIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKAS 408
A +++ I+ +E L EE +E + + AE +++L + + +++ E+E++
Sbjct: 320 ETARRIEEAIRKNVEEKLNSEEVKVEIERRV-AEGVKKLFD------DVEVQLEKEKQ-- 472
Query: 409 SAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYL-----EQIRERA 463
A EAR E+ RKE +EE + + ++ R+ ES++R + L E+ RE
Sbjct: 473 DALTEARRKEEQARKE------REELDKMLEENRRRVEESQRREALELQRKEEERQRELE 634
Query: 464 NLRDQSSPLLRRSLNKDGQGRSTPNNSSDDSQTNIAS 500
+R Q RR +D + + NNS +++ S
Sbjct: 635 MIRRQKEEAARRKKLED-EEHANRNNSMGKNKSRANS 742
>TC81670 weakly similar to PIR|E86293|E86293 hypothetical protein AAF18488.1
[imported] - Arabidopsis thaliana, partial (41%)
Length = 1797
Score = 43.5 bits (101), Expect = 5e-04
Identities = 50/212 (23%), Positives = 96/212 (44%), Gaps = 9/212 (4%)
Frame = +2
Query: 213 SLGRKSAERVRTLHDKLMSPEK--KKKTTSDLKKEAEEKHARALRIRSELETERVQKLQR 270
SL R E+ R LH+ + + ++K ++++ EE+ ++R+EL+ E+++KL
Sbjct: 902 SLSRMLEEKDR-LHNAFVEESRSMQRKAREEVRRILEEQE----KLRNELD-EKMRKLDT 1063
Query: 271 TSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDES-------SKVN 323
S+ LN+ KL E + R+ES LA ++ DE+ K
Sbjct: 1064 WSRDLNKREVLTDQERQKLEEDKKKKDSRNES---LMLASKEQKIADENVFRLVEEQKRE 1234
Query: 324 EVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEK 383
+ + + + K+L +QKL KLQV+K +D + +E +K
Sbjct: 1235 KEEALNKILQLEKQLDAKQKLEMEIEELRGKLQVMKHLGDQDDTAIKKKMEEMNSELEDK 1414
Query: 384 LQRLAEMQRKKEEAQIRREEERKASSAAREAR 415
++ L +M+ ++ ER+++ +EAR
Sbjct: 1415 IESLEDMESMNSTLIVK---ERQSNDELQEAR 1501
>TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-like
{Arabidopsis thaliana}, partial (28%)
Length = 2450
Score = 42.0 bits (97), Expect = 0.001
Identities = 94/439 (21%), Positives = 167/439 (37%), Gaps = 71/439 (16%)
Frame = +2
Query: 103 ESLLTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKD--------KAPTEVINE 154
E + E + K P ++ + + + H+ +L LK+ KA V E
Sbjct: 416 EGAFSGERPIHKKPKAYSAERVLAKETQLHVAE--KELNKLKEQVKNAETTKAQALVELE 589
Query: 155 KNPRSTDNLRRQMQLPEKDKEK--RSTSLGKSM------------NAWKE---------- 190
+ R+ D+L ++++L + +E ++T KS AWKE
Sbjct: 590 RAQRTVDDLTQKLKLITESRESAVKATEAAKSQAKQKYGESDGVNGAWKEELENAVQRYA 769
Query: 191 --------------KRNWEDILSSPFRVSS--RMSYSPSLGRKSAERVRTLHDKLMSPEK 234
K E SS RVS+ R + + +++ ERV L ++ + ++
Sbjct: 770 SIMTELDAAKQDLRKTRQEYDSSSDARVSAVKRTEEAENAMKENTERVSELSKEISAVKE 949
Query: 235 KKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMY 294
+ T E++++ A L + L L+++ +KL + + K E
Sbjct: 950 SIEQTKLAYVESQQQQALVLTEKDALRQSYKATLEQSKKKLLALKKEFNPEITKSLEAQL 1129
Query: 295 AR--------HQRSESRHEAFLAQVAKRAGD-----ESSK--VNEVRFITSLNEENKKLI 339
A H E++ + L V + ES + V+E + SL E K +
Sbjct: 1130AETMNEIAALHTELENKRSSDLDSVKTVTSELDGAKESLQKVVDEENTLRSLVETLKVEL 1309
Query: 340 LRQKLHGSELRRAEK--------LQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQ 391
K SEL+ E L V K K +L A + + E + L+ +
Sbjct: 1310ENVKKEHSELKEKESELESTVGNLHVKLRKSKSELEACSADESKVRGASEEMILTLSRLT 1489
Query: 392 RKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQR 451
+ EEA RRE E + ++L+++ E K EEAE +KL E E+E
Sbjct: 1490SETEEA--RREVEDMKNKT-------DELKKEAEATKLALEEAE---KKLKEATEEAEAA 1633
Query: 452 RKIYLEQIRERANLRDQSS 470
+ I + L +++S
Sbjct: 1634KAAEASAIEQITVLTERTS 1690
>BQ137234 similar to EGAD|35719|3709 hypothetical protein {Burkholderia
cepacia}, partial (6%)
Length = 1184
Score = 41.6 bits (96), Expect = 0.002
Identities = 61/310 (19%), Positives = 122/310 (38%), Gaps = 13/310 (4%)
Frame = +1
Query: 135 SGADKLLSLKDKAPTEVINEKNPRSTDNLRRQMQLPEKDKEKRSTSLGKSMNAWKEKRNW 194
+G +K S + K E NE+ P++ + +K + +R + +E R
Sbjct: 250 AGRNKRRSQRQKERKE--NERKPKTKQKKGGEETRKKKKQPQRRRGEREEKGEGEEARKR 423
Query: 195 EDILSSPFRVSSRMSYSPSLGRKSAERVR---TLH-----DKLMSPEKKKKTTSDLKKEA 246
E R R ++ G+KS ++ R T H + +P +++ T+ + A
Sbjct: 424 EGP-GRTNRKQKRRTHKEESGKKSKQQQRSTSTAHNTQKRETTRAPARERAETARVTSNA 600
Query: 247 -EEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHE 305
++ A RIR + ++ + T + + + A RH+ R G ++ SR
Sbjct: 601 GDDSTGAAKRIREKTRRRAAERGRDTPRHDGQPTHREARRHVGER-GEERDNETEGSRAA 777
Query: 306 AFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQK---LHGSELR-RAEKLQVIKSK 361
+ +R + + N R E + +K G + R R +K SK
Sbjct: 778 TDADKKTRRRSKNTGRENRAREAAPARESERTRAQHEKHAEQRGEDARGRTDKSARTSSK 957
Query: 362 QKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLR 421
++ AR+E ++ A+ + + ++++ E + ERK R+ R R
Sbjct: 958 RQARGARDEKKQQKETTERAQISDKRKKREKRETEKDQQTTRERKKRQ*RRKQRRRAAER 1137
Query: 422 RKEERAKAQQ 431
RK R + ++
Sbjct: 1138RKRRRQQRRE 1167
Score = 35.8 bits (81), Expect = 0.10
Identities = 40/170 (23%), Positives = 70/170 (40%), Gaps = 16/170 (9%)
Frame = +1
Query: 350 RRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASS 409
R+ E + ++K++ +E ER+ K ++ E RKK++ RR ER+
Sbjct: 229 RKGEARRAGRNKRRSQRQKERKENERKP---KTKQKKGGEETRKKKKQPQRRRGEREEKG 399
Query: 410 AAREARAIE---QLRRKEERAKAQQEEA-ELLAQKLAERLNESEQRRKIYLEQIRERA-- 463
EAR E + RK++R ++E + Q+ + + Q+R+ RERA
Sbjct: 400 EGEEARKREGPGRTNRKQKRRTHKEESGKKSKQQQRSTSTAHNTQKRETTRAPARERAET 579
Query: 464 -----NLRDQSSPLLRRSLNKD-----GQGRSTPNNSSDDSQTNIASGLG 503
N D S+ +R K +GR TP + + +G
Sbjct: 580 ARVTSNAGDDSTGAAKRIREKTRRRAAERGRDTPRHDGQPTHREARRHVG 729
>TC92816 weakly similar to GP|18076679|emb|CAC84774. P70 protein {Nicotiana
tabacum}, partial (21%)
Length = 628
Score = 40.4 bits (93), Expect = 0.004
Identities = 35/148 (23%), Positives = 62/148 (41%), Gaps = 11/148 (7%)
Frame = +3
Query: 144 KDKAPTEVINEKNPRSTDNLRRQMQLPEKDKEKRSTSLGKSMNAWKEKRNWEDILSSPFR 203
+ AP E+ K+P +T RQ++ P S+S K+ SP
Sbjct: 180 RSSAP-EMSQRKSPAATPRTARQLKTPNSGSNSASSSPNPIRKTPKDM--------SPRV 332
Query: 204 VSSRMSYSPSLGRKSAERVRTLHDKLMSPEKKKKTTSD-----------LKKEAEEKHAR 252
R+S+SP +K +V+ L ++ ++ K+ D +++EAEE +
Sbjct: 333 NERRLSHSPISEKKRPSKVQELESQIAKLQEDLKSAKDQLNSSESWKRKVQEEAEEAKKQ 512
Query: 253 ALRIRSELETERVQKLQRTSQKLNRVSE 280
L + ELE R Q ++ + R+ E
Sbjct: 513 ILSLSKELEESRQQFSDLSASEETRLQE 596
>BQ137244 similar to GP|20161222|dbj Epstein-Barr virus EBNA-1-like protein
{Oryza sativa (japonica cultivar-group)}, partial (5%)
Length = 1050
Score = 40.4 bits (93), Expect = 0.004
Identities = 54/266 (20%), Positives = 101/266 (37%), Gaps = 27/266 (10%)
Frame = +3
Query: 216 RKSAERVR----TLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRT 271
R+ ER R + + K+ + KKK+ L + + + R R R+E +R + QRT
Sbjct: 207 RRGEERSRRSQVSENKKMRAGHKKKQDPKQLGEIKKRQGQRRRRARAEDRGKRESRRQRT 386
Query: 272 SQKLNR-------VSEWHAVRHLKLREG-------MYARHQRSESRHEAFLAQVAKRAGD 317
+ R ++ H +RE RH + +R AQ + D
Sbjct: 387 ATDAQRARPRTRETHKYTTCGHGTVREQARDATQCRRERHATTRNRDRERYAQTGRTTHD 566
Query: 318 ES---------SKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAR 368
+ S +E R + RQ+ + RR + + A+
Sbjct: 567 RADGTRRHSTQSSGDETRRARYGRARKTQPATRQRTTREKTRRGARAAARERAHARRRAQ 746
Query: 369 EEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAK 428
++ RR+ A + +R +RK+E A+ RE+ R+ ++ + + R ++ER
Sbjct: 747 QQRTRPRRERRGATREER---AERKRESARCNREQARRD*RNSKRSSETRRARTQDERET 917
Query: 429 AQQEEAELLAQKLAERLNESEQRRKI 454
+Q ER ++ +KI
Sbjct: 918 ERQRTTPSGDTNATERRRRADGEQKI 995
>BQ752218 similar to GP|19170914|emb hypothetical protein {Encephalitozoon
cuniculi}, partial (5%)
Length = 637
Score = 40.0 bits (92), Expect = 0.005
Identities = 32/116 (27%), Positives = 57/116 (48%), Gaps = 2/116 (1%)
Frame = -2
Query: 374 ERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEE 433
++R + EK ++ AE ++KK REE K + AREA + K+E K +EE
Sbjct: 633 KKRPKKKEEKAKKEAEKKKKKA-----REEAAKKAKKAREAAYKKAKEEKKEAEKKAKEE 469
Query: 434 AELLAQKLAERLNESEQRRKIYLEQIRERANLRDQSSPLLRRSL--NKDGQGRSTP 487
+K+ E E+E++ K + ++ + +D S L R+ +K+ + S P
Sbjct: 468 KRQAEKKIKEEKKEAEKKAK----KDKKEQDRKDAESALSRQDTEDSKESKDGSKP 313
>BI270480 weakly similar to GP|9757727|db contains similarity to guanylate
binding protein~gene_id:MCL19.12 {Arabidopsis thaliana},
partial (12%)
Length = 657
Score = 38.5 bits (88), Expect = 0.016
Identities = 42/158 (26%), Positives = 75/158 (46%), Gaps = 1/158 (0%)
Frame = +3
Query: 328 ITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRL 387
I+SL E K L R K ++ + E+ ++ ++K L ++ + +R
Sbjct: 90 ISSLRNEIKDLTDRMKSENAKAQSYEREAIVYQQEKNHLEQKYQ----------SEFKRF 239
Query: 388 AEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQ-KLAERLN 446
E+Q + + A+ +E +A+ A ARA + +K E+++ Q+ E LAQ + AER
Sbjct: 240 EEVQERCKTAE---KEAARATEVADRARAEAGMAQK-EKSEMQRLAMERLAQIERAERRI 407
Query: 447 ESEQRRKIYLEQIRERANLRDQSSPLLRRSLNKDGQGR 484
E+ R K LE +RA ++ + L L + Q R
Sbjct: 408 ETLGREKDNLEGELQRATDSEKDARLTVAKLEEKVQQR 521
>TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown protein
{Arabidopsis thaliana}, partial (27%)
Length = 1097
Score = 38.5 bits (88), Expect = 0.016
Identities = 69/320 (21%), Positives = 140/320 (43%), Gaps = 12/320 (3%)
Frame = +2
Query: 109 EVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDKAPTEVINEKNPRSTDNLRRQMQ 168
E+ + + LE S++L+ L G ++LL A +E+ + K + Q
Sbjct: 188 EMKVEEANQLERSASLSLETATKQL-EGKNELLH---DAESEISSLKEKLGMLEMTVGRQ 355
Query: 169 LPEKDKEKRSTSLGKSMNAWKEKRNWEDILSSPFRVSSRMSYSPSLGRKSAERVRTLHDK 228
+ + +R K N K+ E + S VS + + + + SA V+TL
Sbjct: 356 RGDLEDAERCLLAAKEENIEMSKKI-ESLESEIETVSKEKAQALNNEKLSASSVQTLL-- 526
Query: 229 LMSPEKKKKTTSDLK--KEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRH 286
E+K K ++L+ ++ EEK A+ + E + + T +KL H
Sbjct: 527 ----EEKNKLINELEICRDEEEKTKLAMDSLASALHEVSAEARDTKEKLLANQAEHESYE 694
Query: 287 LKLREGMYARHQRSESRHEAFLAQVAKRAGDESS--KVNEVRFITSLNEENKKL-ILRQK 343
++ E + + + S+ ++E+ L D S + ++ ++ + LN+ + ++ +L
Sbjct: 695 TQI-EDLKSDLEASKEKYESMLNDAHHEIEDLKSDLEASKEKYESMLNDAHHEIDVLTSS 871
Query: 344 LHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAE------KLQRLAEMQRK-KEE 396
+ S K+ ++ SK + + +E ++E K E E ++ RL + +K +EE
Sbjct: 872 IENS------KMDILNSKAEWE-QKEHDLVECIKRTEEENSSLGNEVNRLISLLKKTEEE 1030
Query: 397 AQIRREEERKASSAAREARA 416
A ++REEE + +E A
Sbjct: 1031ANVKREEETQLKENMKEVEA 1090
>TC79516 similar to GP|8777406|dbj|BAA96996.1
gb|AAF13095.1~gene_id:MIF21.5~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (29%)
Length = 1077
Score = 38.5 bits (88), Expect = 0.016
Identities = 35/146 (23%), Positives = 75/146 (50%), Gaps = 14/146 (9%)
Frame = +3
Query: 339 ILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQ 398
++++ + E EK+++ K + A + + ++ + EAE+L+ M+R+K+++Q
Sbjct: 222 VVQEAIRKMEFVADEKMRMFKKARLALEACDRELADKAR--EAEELK----MERQKKKSQ 383
Query: 399 IRREE-------------ERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERL 445
I E + KA+ A REA ++++ AK+ + E E + L ++L
Sbjct: 384 IEELERIVRLKNAEADMFQLKANEAKREAERLQRI----ALAKSDKSEEEYTSNYLKQKL 551
Query: 446 NESEQRRKIYLEQIRERANLR-DQSS 470
+E+E ++ E+I+ + + R QSS
Sbjct: 552 SEAEAEKQYLYEKIKLQESSRLSQSS 629
>TC84723 similar to GP|16945387|emb|CAD11800. conserved hypothetical protein
{Neurospora crassa}, partial (5%)
Length = 999
Score = 37.4 bits (85), Expect = 0.036
Identities = 30/126 (23%), Positives = 61/126 (47%), Gaps = 6/126 (4%)
Frame = +3
Query: 363 KEDLAREEAVLERR--KLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQL 420
+E+ E A+ E R K+ +A + + L + +EEA+++R + REAR E+
Sbjct: 486 REEYEAERALQELRELKISKAREREELRVQKEYREEAELQRAKRDLDEIRNREARISEEK 665
Query: 421 RRKEERAKAQQEEAELLAQKLAERLNES----EQRRKIYLEQIRERANLRDQSSPLLRRS 476
K + + +E LA+K +R E+ E+ +K E++ ++ + + +R
Sbjct: 666 HLKSQVELQKYQEEMRLAEKAKQREKEAADAVERFQKKEQERLAKQEKEKQEREKEFQRR 845
Query: 477 LNKDGQ 482
L ++ Q
Sbjct: 846 LQEERQ 863
Score = 35.8 bits (81), Expect = 0.10
Identities = 54/209 (25%), Positives = 89/209 (41%), Gaps = 4/209 (1%)
Frame = +3
Query: 257 RSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRAG 316
R E E ER + E ++ K RE R Q+ E R EA L Q AKR
Sbjct: 486 REEYEAERA------------LQELRELKISKAREREELRVQK-EYREEAEL-QRAKRDL 623
Query: 317 DE----SSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAV 372
DE ++++E + + S E K + E+R AEK +KQ+E A +
Sbjct: 624 DEIRNREARISEEKHLKSQVELQK--------YQEEMRLAEK-----AKQREKEAAD--A 758
Query: 373 LERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQE 432
+ER + ++ +RLA+ +++K+E R++E + QE
Sbjct: 759 VER---FQKKEQERLAKQEKEKQE-------------------------REKEFQRRLQE 854
Query: 433 EAELLAQKLAERLNESEQRRKIYLEQIRE 461
E + ++A+ E E+ K Y ++RE
Sbjct: 855 ERQKELDRIAKAKKEKEENEKEYQRRLRE 941
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 37.0 bits (84), Expect = 0.046
Identities = 30/180 (16%), Positives = 85/180 (46%)
Frame = +3
Query: 319 SSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKL 378
SSK+ E +++++ + K+ EK IKS +K+D +E+ E++
Sbjct: 57 SSKMEESNIPIKIDDKSAGDVKEDKVE------IEKDLEIKSVEKDDEKKEKKDKEKKDK 218
Query: 379 IEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLA 438
+ ++ + + ++KK+E ++EE K + + ++ ++KE++ K ++++ +
Sbjct: 219 TDVDEGKDKKDKEKKKKE---KKEENVKGEEEDGDEKKDKEKKKKEKKEKGKEDKDK--- 380
Query: 439 QKLAERLNESEQRRKIYLEQIRERANLRDQSSPLLRRSLNKDGQGRSTPNNSSDDSQTNI 498
+ E++ K E+ +++ D+ ++ NKD + + + ++ + ++
Sbjct: 381 -------DGEEKKSKKDKEKKKDKNEDDDEGEDGSKKKKNKDKKEKKKEEDEKEEGKVSV 539
Score = 30.4 bits (67), Expect = 4.3
Identities = 32/164 (19%), Positives = 67/164 (40%), Gaps = 13/164 (7%)
Frame = +1
Query: 216 RKSAERVRTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQKL 275
RK R+R KLM + + T +K+ K R LR++ ++ +R +++R ++
Sbjct: 181 RKRKRRIRRRRTKLM*MRVRIRRTKRRRKKKRRK--RMLRVKKKMVMKR--RIRRRRKRK 348
Query: 276 NRVSEWHAVRHLKLREGMYARHQRSESRHEAFL----------AQVAKRAGDESSKVNEV 325
R E + R + +R +R + + ++ KR + V
Sbjct: 349 RRRREKKTKTRMVKRRNLRKTRRRKRTRMKTMMKVRMEVRKRRIRIRKRKRRRRMRKKRV 528
Query: 326 RFITSL---NEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDL 366
+++ + + KK+ R++ + RR L+ IK+ L
Sbjct: 529 KYL*GILI*KKPRKKVKRRRRKKKTRKRRRRNLERIKTNDPSKL 660
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.129 0.350
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,316,549
Number of Sequences: 36976
Number of extensions: 382389
Number of successful extensions: 1964
Number of sequences better than 10.0: 87
Number of HSP's better than 10.0 without gapping: 1882
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1938
length of query: 1215
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1108
effective length of database: 5,058,295
effective search space: 5604590860
effective search space used: 5604590860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0087.21


