

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0087.11
(146 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
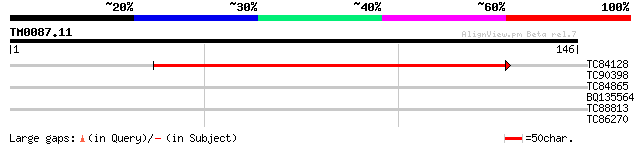
TC84128 homologue to GP|9279721|dbj|BAB01311.1 contains similari... 125 5e-30
TC90398 29 0.77
TC84865 weakly similar to GP|1684845|gb|AAB48303.1| pinin {Canis... 28 1.7
BQ135564 similar to PIR|S35942|S35 probable ATP synthase chain -... 27 3.8
TC88813 similar to GP|9828634|gb|AAG00257.1| F1N21.5 {Arabidopsi... 26 6.5
TC86270 similar to GP|15293161|gb|AAK93691.1 unknown protein {Ar... 25 8.5
>TC84128 homologue to GP|9279721|dbj|BAB01311.1 contains similarity to
unknown protein~gb|AAF15970.1~gene_id:MTE24.1
{Arabidopsis thaliana}, partial (56%)
Length = 542
Score = 125 bits (315), Expect = 5e-30
Identities = 61/92 (66%), Positives = 75/92 (81%)
Frame = +3
Query: 38 SPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCG 97
SPRNLLLA++GEVGELSEIFQWKGEV RGLP++ ++K HL EELSDVLLYLVRL+D+CG
Sbjct: 30 SPRNLLLAMIGEVGELSEIFQWKGEVQRGLPDFKEEEKVHLGEELSDVLLYLVRLSDICG 209
Query: 98 LDLGQAALAKIVKNAQKYPVTSAINNGNSKTG 129
+DLG+AAL K+ NA KYP ++ N + G
Sbjct: 210 VDLGKAALRKVELNAIKYPAKASKEVANKEDG 305
>TC90398
Length = 1434
Score = 28.9 bits (63), Expect = 0.77
Identities = 16/80 (20%), Positives = 38/80 (47%), Gaps = 5/80 (6%)
Frame = +3
Query: 69 NWSCDDKEHLEEELSDVLLYLVRLADVCG-----LDLGQAALAKIVKNAQKYPVTSAINN 123
N C+D + ++ ELS++ L ++ D+ G +D +++ +K+ + +
Sbjct: 327 NSHCEDSKMIKNELSELTSVLEKILDILGANNWVVDRQSKGFRELLPPLEKHIIAILSES 506
Query: 124 GNSKTGEKTTAVVKIDNQRI 143
NSK+ + I +Q++
Sbjct: 507 ENSKSKAHELGIKLIGSQKV 566
>TC84865 weakly similar to GP|1684845|gb|AAB48303.1| pinin {Canis
familiaris}, partial (4%)
Length = 559
Score = 27.7 bits (60), Expect = 1.7
Identities = 18/60 (30%), Positives = 31/60 (51%)
Frame = -2
Query: 42 LLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLG 101
L+++ + V E+ I W E+ G+ NW+CD LSD+ ++ + A +C LG
Sbjct: 240 LIISRLPRVFEVCSIL-WLREILIGVLNWNCD------FGLSDI*IFNLMAALLCDFMLG 82
>BQ135564 similar to PIR|S35942|S35 probable ATP synthase chain - soybean,
partial (69%)
Length = 712
Score = 26.6 bits (57), Expect = 3.8
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = -3
Query: 54 SEIFQWKGEVARGLPNWSC 72
S IF+WK + LPN SC
Sbjct: 302 SLIFRWKSPICSSLPNQSC 246
>TC88813 similar to GP|9828634|gb|AAG00257.1| F1N21.5 {Arabidopsis
thaliana}, partial (2%)
Length = 1511
Score = 25.8 bits (55), Expect = 6.5
Identities = 22/83 (26%), Positives = 36/83 (42%), Gaps = 1/83 (1%)
Frame = +2
Query: 56 IFQWKGEVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKY 115
IFQ E+ R LP W+ + HL++E +CG + G L ++ +
Sbjct: 485 IFQM--ELLRILPTWTLKVRNHLKKE-----------RRICGSETGFNHLKLLLVGMKMI 625
Query: 116 PVTS-AINNGNSKTGEKTTAVVK 137
PV I G+ K G K +++
Sbjct: 626 PVKDILIYQGDPKGGVKRLLLLR 694
>TC86270 similar to GP|15293161|gb|AAK93691.1 unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 2204
Score = 25.4 bits (54), Expect = 8.5
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +2
Query: 33 WDQYHSPRNLLLALV 47
W + HSP N+LLAL+
Sbjct: 1955 WKEIHSPENVLLALL 1999
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.132 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,740,526
Number of Sequences: 36976
Number of extensions: 43819
Number of successful extensions: 153
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 153
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 153
length of query: 146
length of database: 9,014,727
effective HSP length: 87
effective length of query: 59
effective length of database: 5,797,815
effective search space: 342071085
effective search space used: 342071085
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0087.11


