

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.5
(176 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
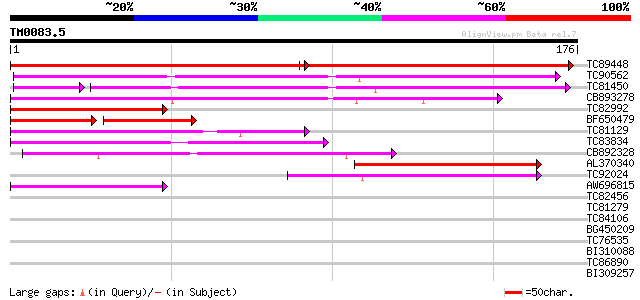
Score E
Sequences producing significant alignments: (bits) Value
TC89448 similar to PIR|C84864|C84864 hypothetical protein At2g43... 172 2e-82
TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 135 8e-33
TC81450 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 116 1e-27
CB893278 weakly similar to PIR|T05645|T056 hypothetical protein ... 112 6e-26
TC82992 homologue to PIR|T05645|T05645 hypothetical protein F20D... 66 8e-12
BF650479 homologue to PIR|T06643|T066 hypothetical protein T20K1... 45 2e-11
TC81129 similar to GP|5764395|gb|AAD51282.1| far-red impaired re... 59 8e-10
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 55 1e-08
CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sin... 49 1e-06
AL370340 similar to PIR|T05644|T056 hypothetical protein F20D10.... 47 3e-06
TC92024 similar to PIR|T05645|T05645 hypothetical protein F20D10... 47 4e-06
AW696815 homologue to GP|11994736|dbj far-red impaired response ... 42 2e-04
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 34 0.034
TC81279 similar to PIR|T47949|T47949 hypothetical protein F2A19.... 31 0.29
TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity ... 29 0.84
BG450209 similar to GP|10177604|db gb|AAD41995.1~gene_id:K21H1.1... 28 1.4
TC76535 similar to SP|Q9BBN6|YCF1_LOTJA Hypothetical 214.8 kDa p... 28 1.4
BI310088 26 7.2
TC86890 weakly similar to GP|4406801|gb|AAD20110.1| unknown prot... 26 7.2
BI309257 similar to PIR|T47436|T474 protein kinase-like protein ... 26 9.3
>TC89448 similar to PIR|C84864|C84864 hypothetical protein At2g43280
[imported] - Arabidopsis thaliana, partial (80%)
Length = 1177
Score = 172 bits (437), Expect(2) = 2e-82
Identities = 79/93 (84%), Positives = 87/93 (92%)
Frame = +1
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEF SEDAAK+FYDEYARHVGFVMRVMSCRRS RDGRILARR GCNKEGHC++I+GKLG
Sbjct: 313 MEFVSEDAAKIFYDEYARHVGFVMRVMSCRRSXRDGRILARRFGCNKEGHCVSIQGKLGP 492
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVK 93
+R+PRASTREGCKAMIHIKYD+SGKW+IT F K
Sbjct: 493 VRKPRASTREGCKAMIHIKYDQSGKWMITXFCK 591
Score = 149 bits (377), Expect(2) = 2e-82
Identities = 71/85 (83%), Positives = 78/85 (91%)
Frame = +2
Query: 91 FVKDHNHPLVVSPREARQTMDEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSD 150
FVKDHNHPLVVSPREARQTMDEKDKKIQELT E+RIKKRLC TYQEQL +M IVEE+SD
Sbjct: 584 FVKDHNHPLVVSPREARQTMDEKDKKIQELTVELRIKKRLCVTYQEQLKCYMNIVEEYSD 763
Query: 151 KLSVKIHHIVNNLKEFEPIEELLHQ 175
KLS +IHHI++NLKE+E IEELLHQ
Sbjct: 764 KLSARIHHILDNLKEYESIEELLHQ 838
>TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (66%)
Length = 1169
Score = 135 bits (340), Expect = 8e-33
Identities = 75/175 (42%), Positives = 104/175 (58%), Gaps = 5/175 (2%)
Frame = +2
Query: 2 EFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSI 61
EF SE A FY+ YA VGFV+RV RS RDG ++ R L CNKEG + K I
Sbjct: 338 EFVSEAEAHAFYNSYATRVGFVIRVSKLSRSRRDGSVIGRALVCNKEG--FRMPDKREKI 511
Query: 62 RRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREAR-----QTMDEKDKK 116
R RA TR GC+AMI ++ SG W ITKFVK+H HPL +P ++R + +
Sbjct: 512 VRQRAETRVGCRAMIMVRKLNSGLWSITKFVKEHTHPL--TPGKSRRDFVYEQYPSGHHR 685
Query: 117 IQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFEPIEE 171
++ELT ++ I+K+ TY+ L + +EEH++ +S KI HIV ++KE E E+
Sbjct: 686 VRELTQQLAIEKKRAETYKRNLDLLYECIEEHNEAVSKKIQHIVESVKEMEAKEQ 850
>TC81450 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (59%)
Length = 832
Score = 116 bits (291), Expect(2) = 1e-27
Identities = 68/155 (43%), Positives = 93/155 (59%), Gaps = 6/155 (3%)
Frame = +2
Query: 26 VMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIRRPRASTREGCKAMIHIKYDKSGK 85
V RS +DG + R L CNKEG+ + K I R RA TR GC+AMI ++ SGK
Sbjct: 125 VSKLSRSRKDGTAIGRALVCNKEGY--RMPDKREKIVRQRAETRVGCRAMIMMRKINSGK 298
Query: 86 WVITKFVKDHNHPLVVSPREARQTMDE------KDKKIQELTAEIRIKKRLCATYQEQLS 139
WVITKFVK+H HPL +P +R+ M E + KI+EL+ ++ I+K+ TY+ QL
Sbjct: 299 WVITKFVKEHTHPL--NPGSSRRDMFEIYPSQNEHDKIRELSQQLAIEKKRSVTYKRQLE 472
Query: 140 SFMKIVEEHSDKLSVKIHHIVNNLKEFEPIEELLH 174
+EEH+ LS K+ IV+++KE EP EE H
Sbjct: 473 VIFDYIEEHNVGLSRKMQRIVDSVKEMEPKEEEEH 577
Score = 22.7 bits (47), Expect(2) = 1e-27
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = +3
Query: 2 EFESEDAAKVFYDEYARHVGFV 23
EFESE AA FY+ G++
Sbjct: 48 EFESEAAAHAFYNSIXAARGWI 113
>CB893278 weakly similar to PIR|T05645|T056 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (5%)
Length = 821
Score = 112 bits (281), Expect = 6e-26
Identities = 62/166 (37%), Positives = 97/166 (58%), Gaps = 13/166 (7%)
Frame = +3
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEG-HCLNIRGKLG 59
M+F S + A+ FYD Y R +GF +R+ RRS + ++ + C+KEG +
Sbjct: 327 MQFNSREEARGFYDGYGRRIGFTVRIHHNRRSRVNNELIGQDFVCSKEGFRAKKYVHRKD 506
Query: 60 SIRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREA--------RQTMD 111
+ P +TREGC+AMI + GKWV+TKFVK+H H L +SP E + D
Sbjct: 507 RVLPPPPATREGCQAMIRLALRDEGKWVVTKFVKEHTHKL-MSPGEVPWRGSGKHLVSED 683
Query: 112 EKDKKIQELTAEIRIK----KRLCATYQEQLSSFMKIVEEHSDKLS 153
EKD++I+EL+ E+ + KR CA Y+EQL++ + +E+H++ +S
Sbjct: 684 EKDRRIRELSLELHNERQKYKRRCAAYEEQLNTILNDLEKHTEHIS 821
>TC82992 homologue to PIR|T05645|T05645 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (7%)
Length = 354
Score = 65.9 bits (159), Expect = 8e-12
Identities = 32/49 (65%), Positives = 36/49 (73%)
Frame = +1
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEG 49
MEFESE+AAK FY+ YAR VGF RV S RRS RDG I+ R+ C KEG
Sbjct: 205 MEFESEEAAKAFYNSYARRVGFSTRVSSSRRSRRDGAIIQRQXVCAKEG 351
>BF650479 homologue to PIR|T06643|T066 hypothetical protein T20K18.200 -
Arabidopsis thaliana, partial (13%)
Length = 580
Score = 44.7 bits (104), Expect(2) = 2e-11
Identities = 20/27 (74%), Positives = 24/27 (88%)
Frame = +2
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVM 27
MEF+SE+AA+ FY EYAR VGFV+RVM
Sbjct: 404 MEFDSEEAARKFYAEYARRVGFVVRVM 484
Score = 39.7 bits (91), Expect(2) = 2e-11
Identities = 19/29 (65%), Positives = 21/29 (71%)
Frame = +3
Query: 30 RRSERDGRILARRLGCNKEGHCLNIRGKL 58
RRS DGR LARRLGCNK+G N +G L
Sbjct: 492 RRSGIDGRTLARRLGCNKQGFSPNSKGTL 578
>TC81129 similar to GP|5764395|gb|AAD51282.1| far-red impaired response
protein {Arabidopsis thaliana}, partial (13%)
Length = 721
Score = 59.3 bits (142), Expect = 8e-10
Identities = 31/94 (32%), Positives = 52/94 (54%), Gaps = 1/94 (1%)
Frame = +2
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
+EFE+ +AA FY EYA+ +GF + + RRS++ + + C++ G G+
Sbjct: 425 IEFETHEAAYSFYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVTPESDGRSSQ 604
Query: 61 IRRPRASTRE-GCKAMIHIKYDKSGKWVITKFVK 93
R+S ++ CKA +H+K + GKW I +F K
Sbjct: 605 ----RSSVKKTDCKACMHVKRNPDGKWNIFEFYK 694
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 55.5 bits (132), Expect = 1e-08
Identities = 34/99 (34%), Positives = 47/99 (47%)
Frame = +3
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEFES D A +Y YA+ VGF +RV + L C+ +G +
Sbjct: 225 MEFESYDDAYSYYICYAKEVGFCVRVKNSWFKRNSKEKYGAVLCCSSQGF-----KRTKD 389
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPL 99
+ R TR GC AMI +K +S +W I + +HNH L
Sbjct: 390 VNNLRKETRTGCPAMIRMKLVESQRWRICEVTLEHNHVL 506
>CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sinorhizobium
meliloti}, partial (1%)
Length = 828
Score = 48.5 bits (114), Expect = 1e-06
Identities = 35/122 (28%), Positives = 51/122 (41%), Gaps = 6/122 (4%)
Frame = +1
Query: 5 SEDAAKVFYDEYARHVGFVMRV-MSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIRR 63
S+D A Y E+A GF +R + + C+K+G N G +
Sbjct: 178 SKDEAYDLYQEHAFKTGFSVRKGKELYYDNEKKKTRLKDFYCSKQGFKNNEPD--GEVAY 351
Query: 64 PRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSP-----REARQTMDEKDKKIQ 118
RA +R C AM+ K G W +TK + DHNH V R R + K+ +I+
Sbjct: 352 KRADSRTNCLAMVRFNVTKEGVWKVTKLILDHNHEFVPLQQRYLLRSMRNMSNLKEDRIK 531
Query: 119 EL 120
L
Sbjct: 532 SL 537
>AL370340 similar to PIR|T05644|T056 hypothetical protein F20D10.290 -
Arabidopsis thaliana, partial (10%)
Length = 508
Score = 47.4 bits (111), Expect = 3e-06
Identities = 24/58 (41%), Positives = 37/58 (63%)
Frame = +1
Query: 108 QTMDEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKE 165
Q++ EK KKI+ELTAE+ + C Y+ L + +K +EE KLSVK+ + ++KE
Sbjct: 73 QSVAEKQKKIRELTAELETPNQRCNVYRANLLAVLKDMEEQKLKLSVKVQNARLSMKE 246
>TC92024 similar to PIR|T05645|T05645 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (11%)
Length = 938
Score = 47.0 bits (110), Expect = 4e-06
Identities = 27/84 (32%), Positives = 43/84 (51%), Gaps = 5/84 (5%)
Frame = +3
Query: 87 VITKFVKDHNHPLVVSPREARQ-----TMDEKDKKIQELTAEIRIKKRLCATYQEQLSSF 141
V + + D NH + S + D+ DK I +LT E+ R C Y+ L S
Sbjct: 444 VRSHLLNDENHAIYSSGCQEESLGEHMNEDDMDKHITKLTDELECANRKCEMYRSNLLSV 623
Query: 142 MKIVEEHSDKLSVKIHHIVNNLKE 165
+K VE+H +LSVK+ +I ++K+
Sbjct: 624 LKAVEDHKLELSVKVENIKISMKD 695
>AW696815 homologue to GP|11994736|dbj far-red impaired response protein;
Mutator-like transposase-like protein; phytochrome A
signaling, partial (7%)
Length = 394
Score = 41.6 bits (96), Expect = 2e-04
Identities = 19/49 (38%), Positives = 27/49 (54%)
Frame = +2
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEG 49
MEFES A FY EYAR +GF + + RRS+ + + C++ G
Sbjct: 227 MEFESHGEAYSFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYG 373
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 33.9 bits (76), Expect = 0.034
Identities = 26/102 (25%), Positives = 46/102 (44%), Gaps = 3/102 (2%)
Frame = +1
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
M F SE+ +Y YA VGF + +S + ++ + L CN C R + +
Sbjct: 220 MTFRSEEEVIRYYMNYANRVGFGVTKISSKNADDGKKYFT--LACN----CA--RRYVST 375
Query: 61 IRRPRA---STREGCKAMIHIKYDKSGKWVITKFVKDHNHPL 99
+ P +++ C+A ++ G I++ V +HNH L
Sbjct: 376 SKNPSKQYLTSKTQCRARLNACVALDGSSTISRIVLEHNHEL 501
>TC81279 similar to PIR|T47949|T47949 hypothetical protein F2A19.170 -
Arabidopsis thaliana, partial (31%)
Length = 1205
Score = 30.8 bits (68), Expect = 0.29
Identities = 22/75 (29%), Positives = 38/75 (50%), Gaps = 6/75 (8%)
Frame = +2
Query: 105 EARQTMDEKDKKIQELTAEIRIKKRLCATYQEQ---LSSFMKIVEEHSDK---LSVKIHH 158
+ ++T E+DK +QELT R+K+ L E+ + K++EE D L +I H
Sbjct: 422 DLKKTQQERDKAVQELT---RLKQHLLEKENEESEKMDEDTKVIEELRDSNNYLRAQISH 592
Query: 159 IVNNLKEFEPIEELL 173
+ L++ +E L
Sbjct: 593 LERALEQATSDQEKL 637
>TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity to
Arabidopsis thaliana far-red impaired response protein
(GB:AAD51282.1), partial (25%)
Length = 673
Score = 29.3 bits (64), Expect = 0.84
Identities = 14/41 (34%), Positives = 21/41 (51%), Gaps = 3/41 (7%)
Frame = +2
Query: 63 RPRASTREGCKAMIHIK---YDKSGKWVITKFVKDHNHPLV 100
R R S R GC A +++ D +W + +F HNH L+
Sbjct: 395 RDRKSVRCGCDAKMYLSKDVVDGVSQWTVLQFSNVHNHELL 517
>BG450209 similar to GP|10177604|db gb|AAD41995.1~gene_id:K21H1.13~similar to
unknown protein {Arabidopsis thaliana}, partial (40%)
Length = 680
Score = 28.5 bits (62), Expect = 1.4
Identities = 21/69 (30%), Positives = 31/69 (44%)
Frame = +1
Query: 103 PREARQTMDEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNN 162
P + R +D +I E+TA+ R A E EEH S+ + HI++N
Sbjct: 208 PSQFRSQVDALGLRIVEVTADGNCFFRALADQLEGNE------EEHQKYRSMVVKHILDN 369
Query: 163 LKEFEPIEE 171
+ FEP E
Sbjct: 370 REMFEPFIE 396
>TC76535 similar to SP|Q9BBN6|YCF1_LOTJA Hypothetical 214.8 kDa protein ycf1.
{Lotus japonicus}, partial (16%)
Length = 1658
Score = 28.5 bits (62), Expect = 1.4
Identities = 16/41 (39%), Positives = 24/41 (58%), Gaps = 1/41 (2%)
Frame = +3
Query: 127 KKRLCATYQEQLSSFMKIVEEHSDKLSV-KIHHIVNNLKEF 166
K+R+ TY LSSF+KI+E+ D + KI + N L +
Sbjct: 1323 KERISFTYPPDLSSFLKIMEKKMDLFTKDKISYNDNELSNY 1445
>BI310088
Length = 541
Score = 26.2 bits (56), Expect = 7.2
Identities = 10/35 (28%), Positives = 18/35 (50%)
Frame = +3
Query: 23 VMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGK 57
+ R RR E++ ++ L CN+ G C+ G+
Sbjct: 78 ITRCFLFRREEKENEVMFLLLCCNRVGSCIKNHGE 182
>TC86890 weakly similar to GP|4406801|gb|AAD20110.1| unknown protein
{Arabidopsis thaliana}, partial (54%)
Length = 2446
Score = 26.2 bits (56), Expect = 7.2
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = -3
Query: 104 REARQTMDEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLS 153
R+ R+T EKD ++ I + + C TY L+S E +K+S
Sbjct: 1769 RKERKTKGEKDT*VKSEER*IILYRLACKTYLTSLNSCQSKHSEEEEKIS 1620
>BI309257 similar to PIR|T47436|T474 protein kinase-like protein -
Arabidopsis thaliana, partial (19%)
Length = 628
Score = 25.8 bits (55), Expect = 9.3
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = -1
Query: 29 CRRSERDGRILARRLGCNKEG 49
CRRS GR RR GC+++G
Sbjct: 352 CRRSRDGGRSGGRRGGCDEKG 290
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.135 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,449,252
Number of Sequences: 36976
Number of extensions: 69742
Number of successful extensions: 390
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 376
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 381
length of query: 176
length of database: 9,014,727
effective HSP length: 90
effective length of query: 86
effective length of database: 5,686,887
effective search space: 489072282
effective search space used: 489072282
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0083.5


