

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.2
(84 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
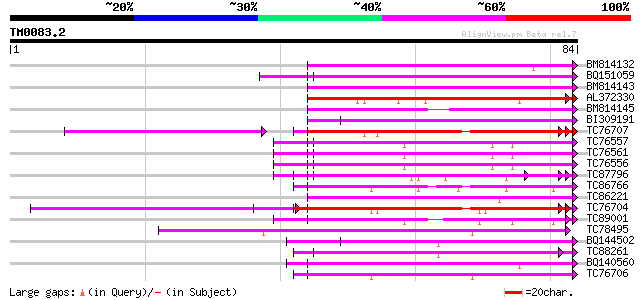
Sequences producing significant alignments: (bits) Value
BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein ... 51 7e-08
BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melano... 50 1e-07
BM814143 GP|21322711|e pherophorin-dz1 protein {Volvox carteri f... 50 1e-07
AL372330 similar to GP|21322752|dbj cold shock protein-1 {Tritic... 47 1e-06
BM814145 homologue to GP|21322711|em pherophorin-dz1 protein {Vo... 47 1e-06
BI309191 46 2e-06
TC76707 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 44 3e-06
TC76557 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 45 5e-06
TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 45 5e-06
TC76556 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 45 5e-06
TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear prot... 44 8e-06
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 44 1e-05
TC86221 similar to PIR|T09608|T09608 environmental stress-induce... 42 3e-05
TC76704 homologue to SP|Q09134|GRPA_MEDFA Abscisic acid and envi... 42 3e-05
TC89001 similar to GP|5726567|gb|AAD48471.1| glycine-rich RNA-bi... 42 5e-05
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 41 7e-05
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 41 9e-05
TC88261 similar to PIR|S22697|S22697 extensin - Volvox carteri (... 40 1e-04
BQ140560 weakly similar to GP|15128223|db hypothetical protein~s... 40 1e-04
TC76706 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 39 4e-04
>BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein
{Mycobacterium tuberculosis CDC1551}, partial (7%)
Length = 164
Score = 51.2 bits (121), Expect = 7e-08
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 28 GGGGGGGGGGGGRGGGGGGGGGGAGGGGGGGGGGGGGGGG 147
Score = 50.8 bits (120), Expect = 9e-08
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 44 GGGGGGGGGGGGGGGGGGRGGGGGGGGGGGGGGGGGGGGG 163
Score = 50.8 bits (120), Expect = 9e-08
Identities = 22/40 (55%), Positives = 23/40 (57%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG+ GG GG GGG GG G GGGGG
Sbjct: 33 GGGGGGGGGGAGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 152
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 23/40 (57%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG+G GGGGG
Sbjct: 4 GGGGGGGGGGGGGGGGGGGGRGGGGGGGGGGAGGGGGGGG 123
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 41 GGGGGGGGGGGGGGGGGGGRGGGGGGGGGGGGGGGGGGGG 160
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 17 GGGGGGGGGGGGGGGGGGGGGGGGGGGRGGGGGGGGGGGG 136
Score = 50.1 bits (118), Expect = 2e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 38 GGGGGGGGGGGGGGGGGGGGRGGGGGGGGGGGGGGGGGGG 157
Score = 50.1 bits (118), Expect = 2e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 14 GGGGGGGGGGGGGGGGGGGGGGGGGGGGRGGGGGGGGGGG 133
Score = 50.1 bits (118), Expect = 2e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 27 GGGGGGGGGGGGAGGGGGGGGGGGGGGGGGGGGGGGGGGG 146
Score = 50.1 bits (118), Expect = 2e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 24 GGGGGGGGGGGGGAGGGGGGGGGGGGGGGGGGGGGGGGGG 143
Score = 49.7 bits (117), Expect = 2e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 21 GGGGGGGGGGGGGGAGGGGGGGGGGGGGGGGGGGGGGGGG 140
Score = 49.7 bits (117), Expect = 2e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 3 GGGRGGGGGGGGGGGGGGGGAGGGGGGGGGGGGGGGGGGG 122
Score = 48.5 bits (114), Expect = 4e-07
Identities = 21/40 (52%), Positives = 21/40 (52%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG G G GGGGG
Sbjct: 10 GGGGGGGGGGGGGGGGGGRGGGGGGGGGGAGGGGGGGGGG 129
Score = 48.5 bits (114), Expect = 4e-07
Identities = 21/40 (52%), Positives = 21/40 (52%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
G GGGGG GG GG GGG GG G GGGGG
Sbjct: 9 GRGGGGGGGGGGGGGGGGAGGGGGGGGGGGGGGGGGGGGG 128
Score = 48.1 bits (113), Expect = 6e-07
Identities = 21/40 (52%), Positives = 21/40 (52%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGG GG GG GGG GG G GGGGG
Sbjct: 36 GGGGGGGGGAGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 155
Score = 45.8 bits (107), Expect = 3e-06
Identities = 22/42 (52%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGS--GDNGGGGG 84
GG + GGGGG GG GG GGG GG G GGGGG
Sbjct: 2 GGGAGGGGGGGGGGGGGGGGGGGGGGGGGGGGRGGGGGGGGG 127
>BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melanogaster},
partial (26%)
Length = 308
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 36 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 155
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 111 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 230
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 40 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 159
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 45 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 164
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 38 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 157
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 50 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 169
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 54 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 173
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 55 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 174
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 59 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 178
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 61 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 180
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 64 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 183
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 48 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 167
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 72 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 191
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 73 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 192
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 78 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 197
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 79 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 198
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 84 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 203
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 87 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 206
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 89 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 208
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 92 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 211
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 94 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 213
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 99 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 218
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 100 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 219
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 105 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 224
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 108 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 227
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 68 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 187
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 112 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 231
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 115 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 234
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 118 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 237
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 121 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 240
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 124 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 243
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 128 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 247
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 131 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 250
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 133 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 252
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 138 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 257
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 140 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 259
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 142 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 261
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 145 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 264
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 149 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 268
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 152 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 271
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 154 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 273
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 159 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 278
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 161 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 280
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 164 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 283
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 167 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 286
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 170 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 289
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 172 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 291
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 176 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 295
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 180 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 299
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 183 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 302
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 184 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 303
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 188 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 307
Score = 49.3 bits (116), Expect = 3e-07
Identities = 21/39 (53%), Positives = 23/39 (58%)
Frame = +2
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
G++ GGGGG GG GG GGG GG G GGGGG
Sbjct: 23 GENPGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 139
Score = 48.9 bits (115), Expect = 3e-07
Identities = 21/47 (44%), Positives = 25/47 (52%)
Frame = +2
Query: 38 VAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
+ ++ + G GGGGG GG GG GGG GG G GGGGG
Sbjct: 11 IPLSGENPGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 151
>BM814143 GP|21322711|e pherophorin-dz1 protein {Volvox carteri f.
nagariensis}, partial (4%)
Length = 132
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 6 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 125
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 7 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 126
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 10 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 129
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 13 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 132
Score = 50.4 bits (119), Expect = 1e-07
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 1 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 120
>AL372330 similar to GP|21322752|dbj cold shock protein-1 {Triticum
aestivum}, partial (33%)
Length = 435
Score = 47.4 bits (111), Expect = 1e-06
Identities = 25/43 (58%), Positives = 27/43 (62%), Gaps = 3/43 (6%)
Frame = +2
Query: 45 GGDSSGGGGGSS---DDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG SSGG GGSS ++GG GG GGG SGG G G GGG
Sbjct: 107 GGGSSGGYGGSSGGYGGSSGGYGGGGYGGGASGGYGGGGYGGG 235
Score = 45.1 bits (105), Expect = 5e-06
Identities = 22/41 (53%), Positives = 24/41 (57%), Gaps = 1/41 (2%)
Frame = +2
Query: 45 GGDSSG-GGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG S G GGGG +GG GG GGG+ GG G G GGG
Sbjct: 152 GGSSGGYGGGGYGGGASGGYGGGGYGGGSGGGYGGGGYGGG 274
Score = 43.5 bits (101), Expect = 1e-05
Identities = 22/40 (55%), Positives = 24/40 (60%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG +SGG GG GG + GG GGG GG G GGGGG
Sbjct: 188 GGGASGGYGGGG---YGGGSGGGYGGGGYGGGGYGGGGGG 298
Score = 42.7 bits (99), Expect = 2e-05
Identities = 22/41 (53%), Positives = 26/41 (62%), Gaps = 1/41 (2%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNN-GGINDGGIGGGNSGGSGDNGGGGG 84
GG SSGG GGSS + + GG GGG+SGG G + GG G
Sbjct: 32 GGSSSGGYGGSSSSSYPASSSGGGYGGGSSGGYGGSSGGYG 154
Score = 42.4 bits (98), Expect = 3e-05
Identities = 21/40 (52%), Positives = 22/40 (54%), Gaps = 1/40 (2%)
Frame = +2
Query: 45 GGDSSG-GGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG S G GGGG + GG GG GGG GG G G GG
Sbjct: 191 GGASGGYGGGGYGGGSGGGYGGGGYGGGGYGGGGGGGYGG 310
Score = 39.3 bits (90), Expect = 3e-04
Identities = 23/44 (52%), Positives = 24/44 (54%), Gaps = 4/44 (9%)
Frame = +2
Query: 45 GGDSSGG-GGGSSDDNNGGINDGGIGGGNSG---GSGDNGGGGG 84
GG GG GGG+S GG GG GGG G G G GGGGG
Sbjct: 164 GGYGGGGYGGGASGGYGGGGYGGGSGGGYGGGGYGGGGYGGGGG 295
Score = 35.8 bits (81), Expect = 0.003
Identities = 19/37 (51%), Positives = 21/37 (56%)
Frame = +2
Query: 48 SSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
SSGGG G G + GG G G+SGG G G GGG
Sbjct: 89 SSGGGYGGGSSGGYGGSSGGYG-GSSGGYGGGGYGGG 196
Score = 33.5 bits (75), Expect = 0.015
Identities = 18/38 (47%), Positives = 19/38 (49%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
GG S GG GG GG GG GGG GG G + G
Sbjct: 227 GGGSGGGYGG------GGYGGGGYGGGGGGGYGGDKMG 322
Score = 30.0 bits (66), Expect = 0.17
Identities = 16/36 (44%), Positives = 16/36 (44%), Gaps = 1/36 (2%)
Frame = +2
Query: 45 GGDSSG-GGGGSSDDNNGGINDGGIGGGNSGGSGDN 79
GG G GGGG GG GG GG G G N
Sbjct: 230 GGSGGGYGGGGYGGGGYGGGGGGGYGGDKMGALGAN 337
>BM814145 homologue to GP|21322711|em pherophorin-dz1 protein {Volvox
carteri f. nagariensis}, partial (3%)
Length = 111
Score = 47.0 bits (110), Expect = 1e-06
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 1 GGGGGGGGGGGGGGGGGG---GGGGGGGGGGGGGGGGGGG 111
Score = 36.6 bits (83), Expect = 0.002
Identities = 19/43 (44%), Positives = 19/43 (44%)
Frame = +2
Query: 42 VDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
V GG GGGGG GGG GG G GGGGG
Sbjct: 20 VGGGGGGGGGGGGG-------------GGGGGGGGGGGGGGGG 109
>BI309191
Length = 109
Score = 46.2 bits (108), Expect = 2e-06
Identities = 20/35 (57%), Positives = 20/35 (57%)
Frame = +2
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GGGGG GG GG GGG GG G GGGGG
Sbjct: 2 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 106
Score = 46.2 bits (108), Expect = 2e-06
Identities = 20/35 (57%), Positives = 20/35 (57%)
Frame = +2
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GGGGG GG GG GGG GG G GGGGG
Sbjct: 5 GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG 109
Score = 45.4 bits (106), Expect = 4e-06
Identities = 22/40 (55%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG GG GGG GG G GGGGG
Sbjct: 1 GGGGGGGGGG----GGGGGGGGGGGGGGGGGGGGGGGGGG 108
>TC76707 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (87%)
Length = 780
Score = 43.9 bits (102), Expect(2) = 3e-06
Identities = 24/43 (55%), Positives = 26/43 (59%), Gaps = 3/43 (6%)
Frame = +1
Query: 45 GGDSSGGGG---GSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG ++GGGG G N GG N GG GG N GG G N GGGG
Sbjct: 268 GGYNNGGGGYNHGGGGYNGGGYNHGG-GGYNHGGGGYNHGGGG 393
Score = 43.5 bits (101), Expect = 1e-05
Identities = 23/43 (53%), Positives = 27/43 (62%), Gaps = 3/43 (6%)
Frame = +1
Query: 45 GGDSSGG---GGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG + GG GGG ++ GG N GG GG N+GG G N GGGG
Sbjct: 190 GGYNGGGYNHGGGGYNNGGGGYNHGG-GGYNNGGGGYNHGGGG 315
Score = 42.0 bits (97), Expect = 4e-05
Identities = 20/40 (50%), Positives = 23/40 (57%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
+ GG GGG + GG N+GG GG N GG G NGGG
Sbjct: 214 NHGGGGYNNGGGGYNHGGGGYNNGG-GGYNHGGGGYNGGG 330
Score = 42.0 bits (97), Expect = 4e-05
Identities = 23/40 (57%), Positives = 25/40 (62%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG + GGGG N GG N GG GG N+GG G N GGGG
Sbjct: 166 GGGYNHGGGGY---NGGGYNHGG-GGYNNGGGGYNHGGGG 273
Score = 40.8 bits (94), Expect = 9e-05
Identities = 20/41 (48%), Positives = 22/41 (52%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
+ GG GGG + GG N GG G G GG G NGGGG
Sbjct: 334 NHGGGGYNHGGGGYNHGGGGYNGGG-GHGGHGGGGYNGGGG 453
Score = 40.0 bits (92), Expect = 2e-04
Identities = 20/42 (47%), Positives = 23/42 (54%), Gaps = 3/42 (7%)
Frame = +1
Query: 45 GGDSSGG---GGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG + GG GGG + GG N GG G GG G +GGGG
Sbjct: 310 GGYNGGGYNHGGGGYNHGGGGYNHGGGGYNGGGGHGGHGGGG 435
Score = 35.8 bits (81), Expect = 0.003
Identities = 19/39 (48%), Positives = 21/39 (53%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG + GGGG NGG GG GGG G G +GG G
Sbjct: 367 GGYNHGGGG-----YNGGGGHGGHGGGGYNGGGGHGGHG 468
Score = 35.8 bits (81), Expect = 0.003
Identities = 19/42 (45%), Positives = 21/42 (49%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
+ GG S GGGS GG N GG GG+ G G GG G
Sbjct: 544 NHGGGSYNHGGGSYHHGGGGYNHGG--GGHGGHGGGGHGGHG 663
Score = 35.0 bits (79), Expect = 0.005
Identities = 24/60 (40%), Positives = 30/60 (50%), Gaps = 5/60 (8%)
Frame = +1
Query: 30 AAVAVVVEVAIAVDEGGDSSGGGG----GSSDDNNGGINDGGIGGGNSGGSGDN-GGGGG 84
AA +V V+ +E D+ GGG G S ++ GG G GG N GG G GGGG
Sbjct: 469 AAESVAVQTEEKTNEVNDAKYGGGYNHGGGSYNHGGGSYHHGGGGYNHGGGGHGGHGGGG 648
Score = 34.3 bits (77), Expect = 0.009
Identities = 21/43 (48%), Positives = 24/43 (54%), Gaps = 3/43 (6%)
Frame = +1
Query: 45 GGDSSGGGG---GSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG + GGGG G + GG N+GG GG N GG G GGG
Sbjct: 169 GGYNHGGGGYNGGGYNHGGGGYNNGG-GGYNHGGGG-YNNGGG 291
Score = 30.0 bits (66), Expect = 0.17
Identities = 13/30 (43%), Positives = 15/30 (49%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGN 72
+ GG GGGG GG N GG GG+
Sbjct: 376 NHGGGGYNGGGGHGGHGGGGYNGGGGHGGH 465
Score = 25.0 bits (53), Expect = 5.3
Identities = 12/44 (27%), Positives = 23/44 (52%)
Frame = -1
Query: 4 IKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAVDEGGD 47
+ V + VVV S+VV +TMVV ++ +++ + G+
Sbjct: 612 VVVTTSTVVVTTSSMVVTTSTMVVTTTIFGIIYFISLFLSLHGN 481
Score = 25.0 bits (53), Expect = 5.3
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = -1
Query: 6 VVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAI 40
V ++ +VV +VVV TMVV + V V +
Sbjct: 318 VTSSTMVVATSTVVVTTTTMVVTTSTVVVTTTAVV 214
Score = 21.2 bits (43), Expect(2) = 3e-06
Identities = 11/30 (36%), Positives = 15/30 (49%)
Frame = +2
Query: 9 AVVVVVWRSVVVVAATMVVLAAAVAVVVEV 38
A+V V VVV+ ++ A V VEV
Sbjct: 161 AMVEVTTTVVVVIMVEVITTAVVVTTTVEV 250
>TC76557 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 912
Score = 45.1 bits (105), Expect = 5e-06
Identities = 20/40 (50%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG + G GG GGG GG G+ GGGG
Sbjct: 532 GGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYGERRGGGG 651
Score = 39.7 bits (91), Expect = 2e-04
Identities = 20/42 (47%), Positives = 22/42 (51%), Gaps = 2/42 (4%)
Frame = +1
Query: 45 GGDSSGGGGGSSD--DNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG + GG + GG GGG GG GGG G
Sbjct: 595 GGYGGGGGGGYGERRGGGGGYSRGGGGGGYGGGGYSRGGGDG 720
Score = 38.5 bits (88), Expect = 5e-04
Identities = 21/42 (50%), Positives = 21/42 (50%), Gaps = 3/42 (7%)
Frame = +1
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGG---GNSGGSGDNGGGGG 84
G GGGGG GG GG GG G GG G GGGGG
Sbjct: 499 GSGGGGGGGRGGGGYGG-GGGGYGGERRGYGGGGGYGGGGGG 621
Score = 37.7 bits (86), Expect = 8e-04
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Frame = +1
Query: 40 IAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNS----GGSGDNGGGGG 84
I V++ GGGG GG GG G G GG G GGGGG
Sbjct: 472 ITVNQAQSRGSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGGGG 618
Score = 35.8 bits (81), Expect = 0.003
Identities = 18/37 (48%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +1
Query: 50 GGGGGSSDDNNGGIND--GGIGGGNSGGSGDNGGGGG 84
GGGGG GG + GG GG + GG G GGGG
Sbjct: 586 GGGGGYGGGGGGGYGERRGGGGGYSRGGGGGGYGGGG 696
Score = 33.5 bits (75), Expect = 0.015
Identities = 20/53 (37%), Positives = 22/53 (40%), Gaps = 3/53 (5%)
Frame = +1
Query: 34 VVVEVAIAVDEGGDSSGGGGGSSDDNNGGINDG---GIGGGNSGGSGDNGGGG 83
+ V A + GG GG GG GG G G GGG G G GG G
Sbjct: 472 ITVNQAQSRGSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYG 630
>TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 1054
Score = 45.1 bits (105), Expect = 5e-06
Identities = 20/40 (50%), Positives = 22/40 (55%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG + G GG GGG GG G+ GGGG
Sbjct: 628 GGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYGERRGGGG 747
Score = 39.7 bits (91), Expect = 2e-04
Identities = 20/42 (47%), Positives = 22/42 (51%), Gaps = 2/42 (4%)
Frame = +1
Query: 45 GGDSSGGGGGSSD--DNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG + GG + GG GGG GG GGG G
Sbjct: 691 GGYGGGGGGGYGERRGGGGGYSRGGGGGGYGGGGYSRGGGDG 816
Score = 38.5 bits (88), Expect = 5e-04
Identities = 21/42 (50%), Positives = 21/42 (50%), Gaps = 3/42 (7%)
Frame = +1
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGG---GNSGGSGDNGGGGG 84
G GGGGG GG GG GG G GG G GGGGG
Sbjct: 595 GSGGGGGGGRGGGGYGG-GGGGYGGERRGYGGGGGYGGGGGG 717
Score = 37.7 bits (86), Expect = 8e-04
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Frame = +1
Query: 40 IAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNS----GGSGDNGGGGG 84
I V++ GGGG GG GG G G GG G GGGGG
Sbjct: 568 ITVNQAQSRGSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGGGG 714
Score = 35.8 bits (81), Expect = 0.003
Identities = 18/37 (48%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +1
Query: 50 GGGGGSSDDNNGGIND--GGIGGGNSGGSGDNGGGGG 84
GGGGG GG + GG GG + GG G GGGG
Sbjct: 682 GGGGGYGGGGGGGYGERRGGGGGYSRGGGGGGYGGGG 792
Score = 33.5 bits (75), Expect = 0.015
Identities = 20/53 (37%), Positives = 22/53 (40%), Gaps = 3/53 (5%)
Frame = +1
Query: 34 VVVEVAIAVDEGGDSSGGGGGSSDDNNGGINDG---GIGGGNSGGSGDNGGGG 83
+ V A + GG GG GG GG G G GGG G G GG G
Sbjct: 568 ITVNQAQSRGSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYG 726
>TC76556 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 754
Score = 45.1 bits (105), Expect = 5e-06
Identities = 20/40 (50%), Positives = 22/40 (55%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG + G GG GGG GG G+ GGGG
Sbjct: 374 GGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYGERRGGGG 493
Score = 39.7 bits (91), Expect = 2e-04
Identities = 20/42 (47%), Positives = 22/42 (51%), Gaps = 2/42 (4%)
Frame = +2
Query: 45 GGDSSGGGGGSSD--DNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG + GG + GG GGG GG GGG G
Sbjct: 437 GGYGGGGGGGYGERRGGGGGYSRGGGGGGYGGGGYSRGGGDG 562
Score = 38.5 bits (88), Expect = 5e-04
Identities = 21/42 (50%), Positives = 21/42 (50%), Gaps = 3/42 (7%)
Frame = +2
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGG---GNSGGSGDNGGGGG 84
G GGGGG GG GG GG G GG G GGGGG
Sbjct: 341 GSGGGGGGGRGGGGYGG-GGGGYGGERRGYGGGGGYGGGGGG 463
Score = 37.7 bits (86), Expect = 8e-04
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Frame = +2
Query: 40 IAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNS----GGSGDNGGGGG 84
I V++ GGGG GG GG G G GG G GGGGG
Sbjct: 314 ITVNQAQSRGSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGGGG 460
Score = 35.8 bits (81), Expect = 0.003
Identities = 18/37 (48%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +2
Query: 50 GGGGGSSDDNNGGIND--GGIGGGNSGGSGDNGGGGG 84
GGGGG GG + GG GG + GG G GGGG
Sbjct: 428 GGGGGYGGGGGGGYGERRGGGGGYSRGGGGGGYGGGG 538
Score = 33.5 bits (75), Expect = 0.015
Identities = 20/53 (37%), Positives = 22/53 (40%), Gaps = 3/53 (5%)
Frame = +2
Query: 34 VVVEVAIAVDEGGDSSGGGGGSSDDNNGGINDG---GIGGGNSGGSGDNGGGG 83
+ V A + GG GG GG GG G G GGG G G GG G
Sbjct: 314 ITVNQAQSRGSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYG 472
>TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear protein.
[strain B95-8 Human herpesvirus 4] {Epstein-barr
virus}, partial (23%)
Length = 431
Score = 44.3 bits (103), Expect = 8e-06
Identities = 21/45 (46%), Positives = 25/45 (54%), Gaps = 5/45 (11%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDN-----NGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG+ + +GG GG GGG+ GG G GG GG
Sbjct: 34 GGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGG 168
Score = 42.7 bits (99), Expect = 2e-05
Identities = 24/52 (46%), Positives = 27/52 (51%), Gaps = 12/52 (23%)
Frame = +1
Query: 45 GGDSSGGGGGSSDD------NNGGINDGGIGGGN------SGGSGDNGGGGG 84
GG S GGGGG + +GG GG GGG+ GGSG GGGGG
Sbjct: 31 GGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGGGG 186
Score = 41.6 bits (96), Expect = 5e-05
Identities = 24/51 (47%), Positives = 25/51 (48%), Gaps = 9/51 (17%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGI---------NDGGIGGGNSGGSGDNGGGGG 84
D GG GGGGGS GG + GG GGG GGS GGGGG
Sbjct: 7 DHGGGY-GGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGG 156
Score = 40.8 bits (94), Expect = 9e-05
Identities = 19/39 (48%), Positives = 19/39 (48%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG S GGGGG GG G GG GG G GGG
Sbjct: 121 GGGSPGGGGGGGGSGGGGGGGGAHGGAYGGGIGGGEGGG 237
Score = 40.0 bits (92), Expect = 2e-04
Identities = 19/39 (48%), Positives = 20/39 (50%)
Frame = +1
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
G G GGG+ GG GG GGG SGG G GG G
Sbjct: 79 GVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGGGGAHG 195
Score = 39.7 bits (91), Expect = 2e-04
Identities = 17/33 (51%), Positives = 18/33 (54%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSG 77
GG GGGG +GG GGIGGG GG G
Sbjct: 145 GGGGGSGGGGGGGGAHGGAYGGGIGGGEGGGHG 243
Score = 38.5 bits (88), Expect = 5e-04
Identities = 19/45 (42%), Positives = 21/45 (46%)
Frame = +1
Query: 40 IAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
+ GG + GG GG S GG GG GGG GG G GG
Sbjct: 82 VGYGSGGGTGGGYGGGSPGGGGG--GGGSGGGGGGGGAHGGAYGG 210
Score = 38.1 bits (87), Expect = 6e-04
Identities = 19/40 (47%), Positives = 19/40 (47%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG G GGGS GG GG GGG G GGG G
Sbjct: 100 GGTGGGYGGGSPGGGGGGGGSGGGGGGGGAHGGAYGGGIG 219
Score = 38.1 bits (87), Expect = 6e-04
Identities = 19/38 (50%), Positives = 20/38 (52%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
GG SGGGGG GG + G GGG GG G GG
Sbjct: 148 GGGGSGGGGGG-----GGAHGGAYGGGIGGGEGGGHGG 246
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 43.9 bits (102), Expect = 1e-05
Identities = 22/47 (46%), Positives = 23/47 (48%), Gaps = 5/47 (10%)
Frame = -1
Query: 43 DEGGDSSGGG-----GGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
D GGD GGG G+ DD GG GG GGG G GGGG
Sbjct: 405 DGGGDGDGGG**GV*CGAGDDGGGGSTQGGGGGGGGSTQGGGDGGGG 265
Score = 41.2 bits (95), Expect = 7e-05
Identities = 21/43 (48%), Positives = 25/43 (57%), Gaps = 3/43 (6%)
Frame = -1
Query: 45 GGDSSGGGGGSSDDN-NGGINDGGIGGGNSGGSGDN--GGGGG 84
GG+ +GGGGG GG+ + G GGG GG G GGGGG
Sbjct: 249 GGECTGGGGGGDKYGYTGGVGE*GGGGGEXGGGGGE*AGGGGG 121
Score = 40.8 bits (94), Expect = 9e-05
Identities = 21/40 (52%), Positives = 24/40 (59%)
Frame = -1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG + GGGGG GG DGG GG +GG G+ GGGG
Sbjct: 336 GGSTQGGGGGGGGSTQGG-GDGG-GGE*TGGGGECTGGGG 223
Score = 38.1 bits (87), Expect = 6e-04
Identities = 21/46 (45%), Positives = 27/46 (58%), Gaps = 6/46 (13%)
Frame = -1
Query: 45 GGDSSGGGGGSSDDNNGGIND--GGIGGGN----SGGSGDNGGGGG 84
GG + GGG G + GG + GG GGG+ +GG G+ GGGGG
Sbjct: 303 GGSTQGGGDGGGGE*TGGGGECTGGGGGGDKYGYTGGVGE*GGGGG 166
Score = 37.0 bits (84), Expect = 0.001
Identities = 19/42 (45%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Frame = -1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGN--SGGSGDNGGGGG 84
GG+ +GGGG + GG G GG GG G+ GGGGG
Sbjct: 270 GGE*TGGGGECTGGGGGGDKYGYTGGVGE*GGGGGEXGGGGG 145
Score = 36.6 bits (83), Expect = 0.002
Identities = 21/51 (41%), Positives = 24/51 (46%), Gaps = 9/51 (17%)
Frame = -1
Query: 43 DEGGDSSGGGGGSS---DDNNGGINDGG------IGGGNSGGSGDNGGGGG 84
D GG+ GGGG + GG DGG G G+ GG G GGGG
Sbjct: 462 DGGGEVYTGGGGEG*AYTGDGGGDGDGGG**GV*CGAGDDGGGGSTQGGGG 310
Score = 36.2 bits (82), Expect = 0.002
Identities = 20/47 (42%), Positives = 23/47 (48%), Gaps = 7/47 (14%)
Frame = -1
Query: 45 GGDSSGGGGGSSDDNNG-----GINDGG--IGGGNSGGSGDNGGGGG 84
GG+ GGGG + G G GG G G GG G+ GGGGG
Sbjct: 171 GGEXGGGGGE*AGGGGGEL***GTGGGGENTGAGGGGGGGEFGGGGG 31
Score = 32.3 bits (72), Expect = 0.033
Identities = 21/52 (40%), Positives = 25/52 (47%), Gaps = 12/52 (23%)
Frame = -1
Query: 45 GGDS---SGG----GGGSSDDNNGGINDGGIGGG-----NSGGSGDNGGGGG 84
GGD +GG GGG + GG G GGG +GG G+N G GG
Sbjct: 222 GGDKYGYTGGVGE*GGGGGEXGGGGGE*AGGGGGEL***GTGGGGENTGAGG 67
Score = 32.3 bits (72), Expect = 0.033
Identities = 19/46 (41%), Positives = 22/46 (47%), Gaps = 9/46 (19%)
Frame = -1
Query: 45 GGDSSGGGGGS----SDDNNGGINDGGIG-----GGNSGGSGDNGG 81
GGD GGGG + D G + GG G G+ GG GD GG
Sbjct: 513 GGDEYTGGGGEGYAYTGDGGGEVYTGGGGEG*AYTGDGGGDGDGGG 376
Score = 31.6 bits (70), Expect = 0.057
Identities = 17/39 (43%), Positives = 21/39 (53%)
Frame = -1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
G +GGGGG D+ GG +G G+ GG GGGG
Sbjct: 537 GPSYTGGGGG--DEYTGGGGEGYAYTGDGGGEVYTGGGG 427
Score = 29.3 bits (64), Expect = 0.28
Identities = 14/33 (42%), Positives = 15/33 (45%)
Frame = -1
Query: 51 GGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GGGG + GG G GGG G GG G
Sbjct: 99 GGGGENTGAGGGGGGGEFGGGGGGDLTQYGGVG 1
Score = 29.3 bits (64), Expect = 0.28
Identities = 17/36 (47%), Positives = 19/36 (52%), Gaps = 2/36 (5%)
Frame = -1
Query: 50 GGGGGSSDD--NNGGINDGGIGGGNSGGSGDNGGGG 83
GGGGG + + GGI GGG GG GGGG
Sbjct: 585 GGGGGDAYEIPKTGGIGPSYTGGG--GGDEYTGGGG 484
Score = 25.8 bits (55), Expect = 3.1
Identities = 19/49 (38%), Positives = 22/49 (44%), Gaps = 9/49 (18%)
Frame = -1
Query: 45 GGDS-----SGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGG----GGG 84
GGD+ +GG G S GG G GG +GD GG GGG
Sbjct: 576 GGDAYEIPKTGGIGPSYTGGGGGDEYTGGGGEGYAYTGDGGGEVYTGGG 430
>TC86221 similar to PIR|T09608|T09608 environmental stress-induced protein -
alfalfa (fragment), partial (25%)
Length = 655
Score = 42.4 bits (98), Expect = 3e-05
Identities = 21/43 (48%), Positives = 25/43 (57%), Gaps = 3/43 (6%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGG---NSGGSGDNGGGGG 84
GG ++GG + DD+N G D GG N GG G NGGGGG
Sbjct: 187 GGSNNGGDSNNGDDSNNGDGDVHEKGGHVYNHGGEGYNGGGGG 315
Score = 29.6 bits (65), Expect = 0.22
Identities = 14/54 (25%), Positives = 24/54 (43%)
Frame = +1
Query: 27 VLAAAVAVVVEVAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNG 80
+L + +V ++ + E D N G + G +GG N+GG +NG
Sbjct: 61 ILMLGLLAMVLISSVISETSTDPNWDSVEQTDLNYGKDGGYVGGSNNGGDSNNG 222
Score = 24.3 bits (51), Expect = 9.1
Identities = 15/68 (22%), Positives = 31/68 (45%), Gaps = 3/68 (4%)
Frame = +1
Query: 20 VVAATMVVLAAAVAVVVEVAIAVDEGGDSSGGGGGSSDDN---NGGINDGGIGGGNSGGS 76
+V+ +++ +A+V+ ++ + D + +D N +GG G GG+S
Sbjct: 43 MVSKKAILMLGLLAMVLISSVISETSTDPNWDSVEQTDLNYGKDGGYVGGSNNGGDSNNG 222
Query: 77 GDNGGGGG 84
D+ G G
Sbjct: 223 DDSNNGDG 246
>TC76704 homologue to SP|Q09134|GRPA_MEDFA Abscisic acid and environmental
stress inducible protein. [Sickle medic] {Medicago
falcata}, partial (81%)
Length = 771
Score = 42.4 bits (98), Expect = 3e-05
Identities = 22/44 (50%), Positives = 28/44 (63%), Gaps = 3/44 (6%)
Frame = +2
Query: 43 DEGGDSSGGG---GGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
+ GG +GGG GG +N GG N GG GG N+GG ++GGGG
Sbjct: 218 NHGGGYNGGGYNHGGGGYNNGGGYNHGG-GGYNNGGGYNHGGGG 346
Score = 40.4 bits (93), Expect = 1e-04
Identities = 23/45 (51%), Positives = 26/45 (57%), Gaps = 7/45 (15%)
Frame = +2
Query: 45 GGDSSGGG---GGSSDDNNGGINDGGIG----GGNSGGSGDNGGG 82
GG ++GGG GG +N GG N GG G G N GG G NGGG
Sbjct: 263 GGYNNGGGYNHGGGGYNNGGGYNHGGGGYNGGGYNHGGGGYNGGG 397
Score = 38.9 bits (89), Expect = 4e-04
Identities = 23/53 (43%), Positives = 28/53 (52%), Gaps = 5/53 (9%)
Frame = +2
Query: 37 EVAIAVDEGGDSSGGGG--GSSDDNNGGINDGGI---GGGNSGGSGDNGGGGG 84
EV +E D+ GGG G ++ GG N GG GGG + G G N GGGG
Sbjct: 149 EVVEKTNEVNDAKYGGGYNGGGYNHGGGYNGGGYNHGGGGYNNGGGYNHGGGG 307
Score = 38.9 bits (89), Expect(2) = 7e-05
Identities = 23/46 (50%), Positives = 25/46 (54%), Gaps = 4/46 (8%)
Frame = +2
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIG----GGNSGGSGDNGGGGG 84
+ GG + GGGG N GG N GG G G N GG G N GGGG
Sbjct: 311 NNGGGYNHGGGGY---NGGGYNHGGGGYNGGGYNHGGGGYNHGGGG 439
Score = 38.5 bits (88), Expect = 5e-04
Identities = 21/41 (51%), Positives = 24/41 (58%), Gaps = 2/41 (4%)
Frame = +2
Query: 45 GGDSSGGG--GGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG + GGG GG + GG N+GG G N GG G N GGG
Sbjct: 209 GGYNHGGGYNGGGYNHGGGGYNNGG--GYNHGGGGYNNGGG 325
Score = 38.1 bits (87), Expect = 6e-04
Identities = 21/46 (45%), Positives = 26/46 (55%), Gaps = 7/46 (15%)
Frame = +2
Query: 45 GGDSSGGGG----GSSDDNNGGINDGGI---GGGNSGGSGDNGGGG 83
GG + GGGG G + GG N GG GGG +GG ++GGGG
Sbjct: 281 GGYNHGGGGYNNGGGYNHGGGGYNGGGYNHGGGGYNGGGYNHGGGG 418
Score = 32.3 bits (72), Expect = 0.033
Identities = 21/52 (40%), Positives = 23/52 (43%), Gaps = 4/52 (7%)
Frame = +2
Query: 37 EVAIAVDEGGDSSGGGGGSSDDNNGGINDGG----IGGGNSGGSGDNGGGGG 84
EV A GG + GG N GG N GG GGG + G G GGG
Sbjct: 170 EVNDAKYGGGYNGGGYNHGGGYNGGGYNHGGGGYNNGGGYNHGGGGYNNGGG 325
Score = 27.7 bits (60), Expect = 0.82
Identities = 12/60 (20%), Positives = 30/60 (50%)
Frame = +2
Query: 23 ATMVVLAAAVAVVVEVAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
A +++ A+ +++ ++ + +++ + +ND GGG +GG ++GGG
Sbjct: 53 AILILGLLAMVLLISSEVSARDLTETTSDAKKEVVEKTNEVNDAKYGGGYNGGGYNHGGG 232
Score = 21.6 bits (44), Expect(2) = 7e-05
Identities = 10/40 (25%), Positives = 17/40 (42%)
Frame = +3
Query: 4 IKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAVD 43
++V VVV + V ++ VVV+ + VD
Sbjct: 207 VEVTTTVVVTMEEVTTTVGVDTTMVEVTTTVVVDTTMVVD 326
>TC89001 similar to GP|5726567|gb|AAD48471.1| glycine-rich RNA-binding
protein {Glycine max}, partial (90%)
Length = 488
Score = 41.6 bits (96), Expect = 5e-05
Identities = 23/46 (50%), Positives = 23/46 (50%), Gaps = 6/46 (13%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGI----GGGNSGGSGDN--GGGGG 84
GG GGGG GG DGG GGG GG GD GGGGG
Sbjct: 333 GGGGGYGGGGGGYGGGGGRRDGGYSRSGGGGGYGGGGDRGYGGGGG 470
Score = 38.5 bits (88), Expect = 5e-04
Identities = 21/42 (50%), Positives = 22/42 (52%), Gaps = 3/42 (7%)
Frame = +3
Query: 45 GGDSSGGGGGSSD---DNNGGINDGGIGGGNSGGSGDNGGGG 83
GG GGGGG D +GG GG GGG G G GGGG
Sbjct: 357 GGGGYGGGGGRRDGGYSRSGG--GGGYGGGGDRGYGGGGGGG 476
Score = 38.5 bits (88), Expect = 5e-04
Identities = 20/43 (46%), Positives = 21/43 (48%), Gaps = 3/43 (6%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNS---GGSGDNGGGGG 84
GG GGGGG G GG GGG GG +GGGGG
Sbjct: 300 GGGRGGGGGGYGGGGGYGGGGGGYGGGGGRRDGGYSRSGGGGG 428
Score = 38.1 bits (87), Expect = 6e-04
Identities = 21/45 (46%), Positives = 22/45 (48%)
Frame = +3
Query: 40 IAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
I V+E GGGG GG G GGG GG G GGGGG
Sbjct: 264 ITVNEAQSRGSGGGG----RGGGGGGYGGGGGYGGGGGGYGGGGG 386
Score = 32.7 bits (73), Expect = 0.025
Identities = 18/35 (51%), Positives = 18/35 (51%)
Frame = +3
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSG 77
D G SGGGGG GG D G GGG GG G
Sbjct: 393 DGGYSRSGGGGG-----YGGGGDRGYGGGGGGGYG 482
Score = 30.8 bits (68), Expect = 0.097
Identities = 19/48 (39%), Positives = 22/48 (45%), Gaps = 2/48 (4%)
Frame = +3
Query: 39 AIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNS--GGSGDNGGGGG 84
AI G D G ++ + G GG GGG GG G GGGGG
Sbjct: 222 AIEEMNGQDIDGRNITVNEAQSRGSGGGGRGGGGGGYGGGGGYGGGGG 365
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 41.2 bits (95), Expect = 7e-05
Identities = 28/66 (42%), Positives = 34/66 (51%), Gaps = 5/66 (7%)
Frame = -2
Query: 23 ATMVVLAAAVAVVV----EVAIAVDEGGDSSGGGGGSSDDNNGGINDGG-IGGGNSGGSG 77
A ++ LA A A VV E + V GG +GGG GG N GG GGG++GG
Sbjct: 664 ALLLSLAGAGAGVVAGGDESGVGVSAGGGVAGGGVAGGGVAGGGENGGGEAGGGDAGGGV 485
Query: 78 DNGGGG 83
D GGG
Sbjct: 484 D*TGGG 467
Score = 35.0 bits (79), Expect = 0.005
Identities = 22/50 (44%), Positives = 29/50 (58%), Gaps = 4/50 (8%)
Frame = -2
Query: 38 VAIAVDEGGDSSGGG--GGSSDDNNGGINDGGIGGGN--SGGSGDNGGGG 83
VA + GG +GGG GG D GG++ G+GGG+ +GG D GGG
Sbjct: 547 VAGGGENGGGEAGGGDAGGGVD*TGGGVS--GVGGGDC*TGGGED*TGGG 404
Score = 32.3 bits (72), Expect = 0.033
Identities = 20/44 (45%), Positives = 23/44 (51%), Gaps = 4/44 (9%)
Frame = -2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGI---GGGNSG-GSGDNGGGGG 84
GG +GGG + GG GG+ GGG SG G GD GGG
Sbjct: 556 GGGVAGGGENGGGEAGGGDAGGGVD*TGGGVSGVGGGDC*TGGG 425
Score = 30.4 bits (67), Expect = 0.13
Identities = 17/58 (29%), Positives = 25/58 (42%), Gaps = 21/58 (36%)
Frame = -2
Query: 46 GDSSGGGGGS---------------SDDN------NGGINDGGIGGGNSGGSGDNGGG 82
G+S+G G+ D++ GG+ GG+ GG G G+NGGG
Sbjct: 691 GESTGANAGALLLSLAGAGAGVVAGGDESGVGVSAGGGVAGGGVAGGGVAGGGENGGG 518
Score = 28.5 bits (62), Expect = 0.48
Identities = 19/44 (43%), Positives = 21/44 (47%), Gaps = 7/44 (15%)
Frame = -2
Query: 45 GGDSSGGG----GGSSDDNNGGINDGGIGG---GNSGGSGDNGG 81
G D GGG GG D GG G +GG G GG+GD G
Sbjct: 385 GDD*RGGGEDWTGGGED*TGGGEA*GVVGGVLAGGVGGAGDEFG 254
Score = 28.1 bits (61), Expect = 0.63
Identities = 18/44 (40%), Positives = 20/44 (44%), Gaps = 5/44 (11%)
Frame = -2
Query: 45 GGDSSGGGG-----GSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG S GGG G D GG++ G G GG D GGG
Sbjct: 472 GGVSGVGGGDC*TGGGED*TGGGVD*AGGGDD*RGGGEDWTGGG 341
Score = 25.0 bits (53), Expect = 5.3
Identities = 19/44 (43%), Positives = 22/44 (49%), Gaps = 5/44 (11%)
Frame = -2
Query: 45 GGDS-SGGG----GGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GGD +GGG GG D GG + G G +GG D GGG
Sbjct: 451 GGDC*TGGGED*TGGGVD*AGGGDD*RGGGEDWTGGGED*TGGG 320
Score = 24.3 bits (51), Expect = 9.1
Identities = 14/35 (40%), Positives = 18/35 (51%), Gaps = 1/35 (2%)
Frame = -2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGG-GNSGGSGD 78
GG GGG + GG+ GG+GG G+ G D
Sbjct: 349 GGGED*TGGGEA*GVVGGVLAGGVGGAGDEFGEVD 245
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 40.8 bits (94), Expect = 9e-05
Identities = 18/35 (51%), Positives = 19/35 (53%)
Frame = +3
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GGGGG GG GG GG + G G GGGGG
Sbjct: 108 GGGGGGGRCGGGGGGGGGRGGAEAAGPGGGGGGGG 212
Score = 37.7 bits (86), Expect = 8e-04
Identities = 20/47 (42%), Positives = 23/47 (48%), Gaps = 4/47 (8%)
Frame = +3
Query: 42 VDEGGDSSGGGGGSSDDNNGG----INDGGIGGGNSGGSGDNGGGGG 84
+ EGG + G GG GG + GG GGG GG G GGG G
Sbjct: 27 IPEGGGARGAGGWRRGGEVGGSGWCVWGGGGGGGRCGGGGGGGGGRG 167
Score = 37.0 bits (84), Expect = 0.001
Identities = 17/33 (51%), Positives = 21/33 (63%)
Frame = +1
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
GGGGG++ D G + GG GGG +GG G G G
Sbjct: 481 GGGGGAASDCGGAL--GGDGGGGTGGGGRGGRG 573
Score = 35.4 bits (80), Expect = 0.004
Identities = 20/51 (39%), Positives = 20/51 (39%), Gaps = 12/51 (23%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSG------------GSGDNGGGG 83
GG GGGG GG G GGG G GSG GGGG
Sbjct: 117 GGGGRCGGGGGGGGGRGGAEAAGPGGGGGGGGDALGRRRLAKGSGAGGGGG 269
Score = 34.3 bits (77), Expect = 0.009
Identities = 18/48 (37%), Positives = 23/48 (47%), Gaps = 9/48 (18%)
Frame = +2
Query: 46 GDSSGGGGGSSDDNNGGI---------NDGGIGGGNSGGSGDNGGGGG 84
G +GGGG ++ GG+ G + GG GG G GGGGG
Sbjct: 38 GGGAGGGGLAAGGGGGGLRLVRVGGWGRGGPLRGGWGGGRGPRGGGGG 181
Score = 33.1 bits (74), Expect = 0.020
Identities = 16/40 (40%), Positives = 18/40 (45%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG +GGGGG G G GG +GG G G G
Sbjct: 49 GGRGAGGGGGRWGAQAGACGGVGAGGAAAGGVGGGAGAEG 168
Score = 31.2 bits (69), Expect = 0.074
Identities = 17/48 (35%), Positives = 21/48 (43%)
Frame = +1
Query: 37 EVAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
E+ I +GG G G G G G GG +GG+ G GGG
Sbjct: 13 ELLIIYQKGGGRGGRGAGGG-GGRWGAQAGACGGVGAGGAAAGGVGGG 153
Score = 30.4 bits (67), Expect = 0.13
Identities = 18/53 (33%), Positives = 21/53 (38%), Gaps = 14/53 (26%)
Frame = +3
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNS--------------GGSGDNGGGGG 84
G GGGGG G GG GGG++ GG + GGGG
Sbjct: 135 GGGGGGGGGRGGAEAAGPGGGGGGGGDALGRRRLAKGSGAGGGGGARSRGGGG 293
Score = 30.0 bits (66), Expect = 0.17
Identities = 15/41 (36%), Positives = 18/41 (43%)
Frame = +3
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
+ G GGGGG + G G G GG+ GGGG
Sbjct: 174 EAAGPGGGGGGGGDALGRRRLAKGS-GAGGGGGARSRGGGG 293
Score = 28.5 bits (62), Expect = 0.48
Identities = 14/34 (41%), Positives = 14/34 (41%)
Frame = +3
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG GG G GG GGG G GGG
Sbjct: 429 GGAGGPEGGARAGAAGGGRGGGRGLGLWRGSGGG 530
Score = 27.7 bits (60), Expect = 0.82
Identities = 14/40 (35%), Positives = 19/40 (47%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG +GG ++ G+ G GG G +G GGGG
Sbjct: 193 GGAVAGGMPSAAGGLLRGVGPGAGGGPGPGAAGGRVGGGG 312
Score = 26.9 bits (58), Expect = 1.4
Identities = 16/40 (40%), Positives = 17/40 (42%), Gaps = 2/40 (5%)
Frame = +3
Query: 46 GDSSGGGGGSSDDNN--GGINDGGIGGGNSGGSGDNGGGG 83
G GGGG + G GG GG S G G GG G
Sbjct: 189 GGGGGGGGDALGRRRLAKGSGAGGGGGARSRGGGGPGGWG 308
Score = 26.6 bits (57), Expect = 1.8
Identities = 15/38 (39%), Positives = 18/38 (46%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
G G GGG+ G+ GG G G G G+ GGG
Sbjct: 464 GRRGGGAGGGARPRIVEGL-WGGTGAGGLAGGGEVGGG 574
Score = 26.2 bits (56), Expect = 2.4
Identities = 13/36 (36%), Positives = 17/36 (47%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNG 80
GG+ GGGG + D G GG +G G+ G
Sbjct: 681 GGEEGGGGGVRARDAVRRSRAGCAWGGGAGRCGEGG 788
Score = 26.2 bits (56), Expect(2) = 0.30
Identities = 10/18 (55%), Positives = 11/18 (60%)
Frame = +2
Query: 67 GIGGGNSGGSGDNGGGGG 84
G GGG +GG G GGG
Sbjct: 440 GAGGGRAGGRRGGGAGGG 493
Score = 26.2 bits (56), Expect = 2.4
Identities = 16/42 (38%), Positives = 20/42 (47%), Gaps = 2/42 (4%)
Frame = +2
Query: 45 GGDSSGG--GGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG +GG GGG+ I +G GG +GG G GG
Sbjct: 446 GGGRAGGRRGGGAGGGARPRIVEGLWGGTGAGGLAGGGEVGG 571
Score = 26.2 bits (56), Expect = 2.4
Identities = 15/39 (38%), Positives = 17/39 (43%)
Frame = +3
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
G +GGGGG+ GGG GG G G G G
Sbjct: 243 GSGAGGGGGARSR----------GGGGPGGWGRAGQGTG 329
Score = 24.6 bits (52), Expect = 6.9
Identities = 15/38 (39%), Positives = 17/38 (44%)
Frame = +3
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
G + G GG+ GG GG G G GSG G G
Sbjct: 429 GGAGGPEGGARAGAAGGGRGGGRGLGLWRGSGGGRGRG 542
Score = 24.3 bits (51), Expect = 9.1
Identities = 16/49 (32%), Positives = 19/49 (38%), Gaps = 11/49 (22%)
Frame = +2
Query: 45 GGDSSGGG-----------GGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
GG GGG GG++ + I GG GG G GGG
Sbjct: 551 GGGEVGGGM*RLGREGRAPGGAAHQFSWQEGRVPIPGGEEGGGGGGGGG 697
Score = 21.6 bits (44), Expect(2) = 0.30
Identities = 10/28 (35%), Positives = 12/28 (42%)
Frame = +2
Query: 44 EGGDSSGGGGGSSDDNNGGINDGGIGGG 71
E G GGG G + +GG G G
Sbjct: 245 EWGRGRGGGPVQGRRGAGWVGEGGAGDG 328
>TC88261 similar to PIR|S22697|S22697 extensin - Volvox carteri (fragment),
partial (7%)
Length = 1516
Score = 40.4 bits (93), Expect = 1e-04
Identities = 18/42 (42%), Positives = 21/42 (49%)
Frame = -2
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
D+ G + GG + G N GG GGG G D GGGGG
Sbjct: 498 DD*GTNDTGGAEDGKERGNGCNIGGGGGGGGGDGDDGGGGGG 373
Score = 38.9 bits (89), Expect = 4e-04
Identities = 19/41 (46%), Positives = 21/41 (50%), Gaps = 4/41 (9%)
Frame = -2
Query: 46 GDSSGGGGGSSDDNNGG----INDGGIGGGNSGGSGDNGGG 82
G GGGGG DD GG GG+GGG S G+G G
Sbjct: 429 GGGGGGGGGDGDDGGGGGGGEEGKGGVGGGESRGNGMEPSG 307
Score = 35.4 bits (80), Expect = 0.004
Identities = 18/42 (42%), Positives = 21/42 (49%), Gaps = 1/42 (2%)
Frame = -2
Query: 43 DEGGDSSGGGGGSSDDNNG-GINDGGIGGGNSGGSGDNGGGG 83
D G + +GG + NG I GG GGG G G GGGG
Sbjct: 495 D*GTNDTGGAEDGKERGNGCNIGGGGGGGGGDGDDGGGGGGG 370
Score = 27.7 bits (60), Expect = 0.82
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = -2
Query: 53 GGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
G +S D+ G + GG G G+G N GGGG
Sbjct: 513 GRASPDD*GTNDTGGAEDGKERGNGCNIGGGG 418
>BQ140560 weakly similar to GP|15128223|db hypothetical protein~similar to
Oryza sativa chromosome 1 P0489A01.2, partial (5%)
Length = 215
Score = 40.4 bits (93), Expect = 1e-04
Identities = 23/50 (46%), Positives = 25/50 (50%), Gaps = 7/50 (14%)
Frame = -1
Query: 42 VDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSG-------GSGDNGGGGG 84
VD G S GGGG + GG +GG GGG G G D GGGGG
Sbjct: 197 VDAGRYSXGGGGAPWNGGGGGGIEGGGGGGGGGRGAPLDAGRYDGGGGGG 48
Score = 40.4 bits (93), Expect = 1e-04
Identities = 18/40 (45%), Positives = 21/40 (52%)
Frame = -1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG ++ G GG GG+ NGGGGG
Sbjct: 140 GGGIEGGGGGGGGGRGAPLDAGRYDGGGGGGAP*NGGGGG 21
Score = 35.0 bits (79), Expect = 0.005
Identities = 18/38 (47%), Positives = 21/38 (54%)
Frame = -1
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
G GGGGG + G DGG GGG + +G GGGG
Sbjct: 125 GGGGGGGGGRGAPLDAGRYDGG-GGGGAP*NGGGGGGG 15
Score = 32.3 bits (72), Expect = 0.033
Identities = 15/35 (42%), Positives = 15/35 (42%)
Frame = -1
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GGGG D GG GG G GGGGG
Sbjct: 215 GGGGAPVDAGRYSXGGGGAPWNGGGGGGIEGGGGG 111
>TC76706 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (94%)
Length = 1034
Score = 38.9 bits (89), Expect = 4e-04
Identities = 19/42 (45%), Positives = 24/42 (56%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
+ GG + GGG N+GG G GG ++GG G N GGGG
Sbjct: 196 NHGGYNHGGGYNGGGYNHGGGYHNGGGGYHNGGGGYNHGGGG 321
Score = 38.5 bits (88), Expect = 5e-04
Identities = 22/47 (46%), Positives = 24/47 (50%), Gaps = 7/47 (14%)
Frame = +1
Query: 45 GGDSSGGG----GGSSDDNNGGINDGG---IGGGNSGGSGDNGGGGG 84
GG + GGG GG + GG N GG GGG GG G GGGG
Sbjct: 235 GGYNHGGGYHNGGGGYHNGGGGYNHGGGGYNGGGGHGGHGGYNGGGG 375
Score = 37.0 bits (84), Expect = 0.001
Identities = 22/41 (53%), Positives = 23/41 (55%), Gaps = 2/41 (4%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGG--SGDNGGGG 83
GG + GGGG NGG GG GG N GG G NGGGG
Sbjct: 295 GGYNHGGGG-----YNGGGGHGGHGGYNGGGGHGGYNGGGG 402
Score = 36.6 bits (83), Expect = 0.002
Identities = 17/39 (43%), Positives = 20/39 (50%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG GGG + GG N GG GG+ G +G G GG
Sbjct: 268 GGGGYHNGGGGYNHGGGGYNGGGGHGGHGGYNGGGGHGG 384
Score = 36.2 bits (82), Expect = 0.002
Identities = 19/41 (46%), Positives = 21/41 (50%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
+ GG S GGGS GG N GG GG G +GGGG
Sbjct: 514 NHGGGSYNHGGGSYHHGGGGYNHGG------GGHGGHGGGG 618
Score = 35.8 bits (81), Expect = 0.003
Identities = 28/84 (33%), Positives = 37/84 (43%), Gaps = 3/84 (3%)
Frame = +1
Query: 3 AIKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAVDE---GGDSSGGGGGSSDDN 59
AI ++ + + + SV+ A VVE V++ GG + GG N
Sbjct: 52 AILMLGLLAMALISSVMSARELTETSTDAKKEVVEKTNEVNDAKYGGYNHGGYNHGGGYN 231
Query: 60 NGGINDGGIGGGNSGGSGDNGGGG 83
GG N GG G N GG NGGGG
Sbjct: 232 GGGYNHGG-GYHNGGGGYHNGGGG 300
Score = 35.4 bits (80), Expect = 0.004
Identities = 21/54 (38%), Positives = 29/54 (52%)
Frame = +1
Query: 30 AAVAVVVEVAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
AA +V V+ +E D+ GGG S ++ G N GG + GGS +GGGG
Sbjct: 418 AAESVAVQTEEKTNEVNDAKYGGG--SYNHGGSYNHGGGSYNHGGGSYHHGGGG 573
Score = 35.4 bits (80), Expect = 0.004
Identities = 21/45 (46%), Positives = 26/45 (57%), Gaps = 3/45 (6%)
Frame = +1
Query: 43 DEGGDSSGGG---GGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
+ GG +GGG GG + GG ++GG GG N GG G GGGG
Sbjct: 211 NHGGGYNGGGYNHGGGYHNGGGGYHNGG-GGYNHGGGG-YNGGGG 339
Score = 32.7 bits (73), Expect = 0.025
Identities = 16/39 (41%), Positives = 19/39 (48%)
Frame = +1
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
G S GGGS + G + GG G + GG GGGG
Sbjct: 502 GGSYNHGGGSYNHGGGSYHHGGGGYNHGGGGHGGHGGGG 618
Score = 28.1 bits (61), Expect(2) = 0.007
Identities = 20/62 (32%), Positives = 23/62 (36%), Gaps = 24/62 (38%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGG----------------------INDGGIGGG--NSGGSGDNG 80
GG GGG NGG +ND GGG N GGS ++G
Sbjct: 343 GGHGGYNGGGGHGGYNGGGGHGGHGAAESVAVQTEEKTNEVNDAKYGGGSYNHGGSYNHG 522
Query: 81 GG 82
GG
Sbjct: 523 GG 528
Score = 25.4 bits (54), Expect(2) = 0.007
Identities = 16/43 (37%), Positives = 21/43 (48%)
Frame = +2
Query: 1 MVAIKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAVD 43
+V V VVV +VVVV T+ V+ V VV V A +
Sbjct: 218 VVVTMVEVITTVVVTTTVVVVTTTVEVVTTMVEVVTMVVEATE 346
Score = 24.6 bits (52), Expect = 6.9
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = -3
Query: 4 IKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEV 38
+ V + VVV S+VV +TMVV + V +
Sbjct: 582 VVVTTSTVVVTTSSMVVTTSTMVVTSTMVVTTTTI 478
Score = 24.3 bits (51), Expect = 9.1
Identities = 13/37 (35%), Positives = 18/37 (48%)
Frame = -3
Query: 6 VVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAV 42
VV VVV + VVV T+V+ + V V + V
Sbjct: 306 VVTTSTVVVTTTTVVVTTTVVITSTIVTTTVVITTMV 196
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.136 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,825,855
Number of Sequences: 36976
Number of extensions: 29438
Number of successful extensions: 4010
Number of sequences better than 10.0: 513
Number of HSP's better than 10.0 without gapping: 1091
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2563
length of query: 84
length of database: 9,014,727
effective HSP length: 60
effective length of query: 24
effective length of database: 6,796,167
effective search space: 163108008
effective search space used: 163108008
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0083.2


