

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.15
(489 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
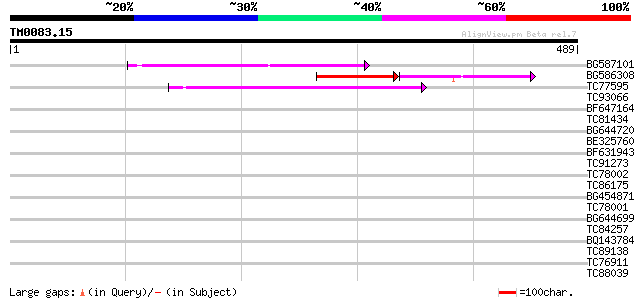
Sequences producing significant alignments: (bits) Value
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 120 1e-27
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 72 3e-24
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 69 5e-12
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 38 0.008
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 37 0.017
TC81434 27 0.10
BG644720 33 0.25
BE325760 similar to PIR|B86423|B8 hypothetical protein AAG12856.... 32 0.72
BF631943 homologue to GP|5902895|dbj type I polyketide synthase ... 31 0.94
TC91273 similar to GP|1435022|dbj|BAA05625.1 DNA-binding protein... 30 2.1
TC78002 homologue to GP|23617209|dbj|BAC20880. contains ESTs C28... 30 2.1
TC86175 similar to GP|23617209|dbj|BAC20880. contains ESTs C2856... 30 2.1
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 30 2.1
TC78001 similar to GP|21618169|gb|AAM67219.1 ARF GAP-like zinc f... 30 2.1
BG644699 similar to PIR|T07863|T078 probable polyprotein - pinea... 30 2.7
TC84257 weakly similar to PIR|G83180|G83180 probable FMN oxidore... 29 3.6
BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antig... 28 6.1
TC89138 homologue to GP|15215736|gb|AAK91413.1 AT5g19690/T29J13_... 28 8.0
TC76911 similar to GP|13543234|gb|AAH05782.1 Unknown (protein fo... 28 8.0
TC88039 homologue to GP|15623873|dbj|BAB67931. hypothetical prot... 28 8.0
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 120 bits (301), Expect = 1e-27
Identities = 66/209 (31%), Positives = 109/209 (51%)
Frame = +2
Query: 102 KLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSLARKVL 161
K VRR W L+K+ ++C+A++ +L H + H K+
Sbjct: 5 KDVRRYLWD---EPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQ 175
Query: 162 RAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLLGPFDTA 221
+AG++WPT+ KDA + +CDPCQR ++ + ++ F WG+D +GPF ++
Sbjct: 176 QAGFWWPTMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSS 355
Query: 222 PG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLITDNGTQFTSK 281
KY++VAVDY +KW+EA T + V + F + RFGVP V+I+D G+ F +K
Sbjct: 356 YN-NKYILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINK 532
Query: 282 GFRDLLDGLHIKQRFTSVEHPQTNGQAES 310
F LL ++ + + HPQ +++S
Sbjct: 533 VFEKLLKKNGVRHKVATAYHPQKAERSKS 619
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 686
Score = 71.6 bits (174), Expect(2) = 3e-24
Identities = 32/71 (45%), Positives = 49/71 (68%)
Frame = -2
Query: 265 GVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLG 324
G+P ++TDNG+ F S FR+ + I+ S +PQ+NGQAE++N++I+ GL+KRL
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 325 DAKGDWAEQLD 335
KG WA++LD
Sbjct: 502 LKKGCWADELD 470
Score = 58.5 bits (140), Expect(2) = 3e-24
Identities = 38/120 (31%), Positives = 54/120 (44%), Gaps = 3/120 (2%)
Frame = -3
Query: 337 VLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLN---QNDQLLGEDLDL 393
VLW++RT P T +P+ + EA+ PAE+ S + N ND+L L+
Sbjct: 465 VLWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNNDRLFNA-LET 289
Query: 394 LAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAAKWEG 453
+ ERR A R + + YNK V + GD+VL + GKL WEG
Sbjct: 288 IEERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNWEG 109
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 68.6 bits (166), Expect = 5e-12
Identities = 54/224 (24%), Positives = 98/224 (43%), Gaps = 2/224 (0%)
Frame = +2
Query: 138 VLAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPAR 197
++ E H+ + H GR+ +++ ++WP + V+ CD C A
Sbjct: 179 LVQESHDSTAAGH-PGRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVCGGIHIWRQAKRGF 355
Query: 198 LSSLVSPWPFHQ-WGMDLLGPFDTAPG*-LKYLVVAVDYYTKWIEAEPLATITSARVQRF 255
L L P H MD + G +YL V VD +K + E + T+ + +
Sbjct: 356 LKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAEACAQR 535
Query: 256 FYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVI 315
F R G+P +++D G+ + + +R+ + Q ++ HPQT+G E N+ I
Sbjct: 536 FLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTERWNQEI 715
Query: 316 LRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFG 359
LR + ++ +W + L V A R +S+ G +P+ + G
Sbjct: 716 QAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHG 847
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 38.1 bits (87), Expect = 0.008
Identities = 42/150 (28%), Positives = 61/150 (40%), Gaps = 15/150 (10%)
Frame = +1
Query: 213 DLLGPFD-TAPG*LKYLVVAVDYYTK--WI-----EAEPLATITSAR--VQRFFYKNVIR 262
DL GP T+ G +Y++ +D + + W+ + E T R V+ KNV +
Sbjct: 115 DLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRYKNETFPTFKKWRILVETQTGKNVKK 294
Query: 263 RFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKR 322
LITDN +F S F + I + T +PQ NG AE R +L R
Sbjct: 295 -------LITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQNGVAERMIRTLLERARCM 453
Query: 323 LGDA-----KGDWAEQLDHVLWAYRTTPHS 347
L +A + W E +PHS
Sbjct: 454 LSNAGL*N*RDLWVEAASTACHLVNRSPHS 543
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 37.0 bits (84), Expect = 0.017
Identities = 29/99 (29%), Positives = 46/99 (46%)
Frame = +1
Query: 390 DLDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAA 449
+L+ L E R A I K+ ++++++ R F G+LVL S GKL +
Sbjct: 103 ELNELEELRLDAYENAKIYKERTKKWHDRRIIRREFREGELVLLFNS--RLKLFPGKLRS 276
Query: 450 KWEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNLRRY 488
W GP++V +GA +E + P T N L+ Y
Sbjct: 277 HWSGPFQVKNVMPSGA--VEVWSESTGPFTVNGQRLKHY 387
>TC81434
Length = 363
Score = 26.9 bits (58), Expect(2) = 0.10
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = +1
Query: 465 AYKLETLAGKEIPRTWNATNLRRY 488
AYKLE A R WNAT+L+ Y
Sbjct: 154 AYKLEGRARNIYLRLWNATHLKLY 225
Score = 26.2 bits (56), Expect(2) = 0.10
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +2
Query: 443 ERGKLAAKWEGPY 455
+ GKLAA W+GPY
Sbjct: 89 KHGKLAASWKGPY 127
>BG644720
Length = 678
Score = 33.1 bits (74), Expect = 0.25
Identities = 30/114 (26%), Positives = 45/114 (39%), Gaps = 4/114 (3%)
Frame = -3
Query: 114 NDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSLARKVLRAGYYWPT--LE 171
N+ L+ F LL CL ++ H CG H L + R GYYW T +
Sbjct: 361 NETLYI*LFEGVLL*CLGEEETIQAFQ*AHSRVCGSHKS--KLYFHIKRMGYYWNT*IML 188
Query: 172 KDA--ADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLLGPFDTAPG 223
K A A+ ++Q PP L ++ F W +++ G F + G
Sbjct: 187 KGALLANFIQQ-------------PPEVLHPTITS*LFESWELNVFGLFPKSSG 65
>BE325760 similar to PIR|B86423|B8 hypothetical protein AAG12856.1 [imported]
- Arabidopsis thaliana, partial (11%)
Length = 669
Score = 31.6 bits (70), Expect = 0.72
Identities = 28/97 (28%), Positives = 41/97 (41%), Gaps = 8/97 (8%)
Frame = +2
Query: 105 RRASWYTIVNDDLFKRG--FSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSL-----A 157
R +W+ + +++ G FST C++ R A L + C HL + A
Sbjct: 95 RTKAWW---GNTIWRNGT*FSTSTWGCISSAR*AAHLF*LSHC*CVFHLWKHAFNNIH*A 265
Query: 158 RKVLRAGYYWPTLEKDAADHVKQCDPCQ-RHADLHHA 193
K +GY W T +K + C P Q RH D HA
Sbjct: 266 SKAASSGYSWDTSKKSSKKSHLSCLPNQ*RHGDSSHA 376
>BF631943 homologue to GP|5902895|dbj type I polyketide synthase AVES 3
{Streptomyces avermitilis}, partial (0%)
Length = 544
Score = 31.2 bits (69), Expect = 0.94
Identities = 14/35 (40%), Positives = 19/35 (54%)
Frame = -1
Query: 40 ARSHPLTTRKHAAGHPRPSGWPAIHRGAHPGAGAH 74
+RS P +R A +GWPA G+HP G+H
Sbjct: 280 SRSSPWYSRVAFATLMTSTGWPATSSGSHPSMGSH 176
>TC91273 similar to GP|1435022|dbj|BAA05625.1 DNA-binding protein {Daucus
carota}, partial (33%)
Length = 722
Score = 30.0 bits (66), Expect = 2.1
Identities = 13/33 (39%), Positives = 19/33 (57%), Gaps = 3/33 (9%)
Frame = +1
Query: 187 HADLHHAPPARLSSLV---SPWPFHQWGMDLLG 216
H D HH PP L+S++ +P +H G+ LG
Sbjct: 313 HQDEHHQPPPSLNSIITSCAPQDYHGGGVSFLG 411
>TC78002 homologue to GP|23617209|dbj|BAC20880. contains ESTs C28562(C61610)
C25917(C11090)~similar to ARF GAP-like zinc
finger-containing protein, partial (26%)
Length = 737
Score = 30.0 bits (66), Expect = 2.1
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +3
Query: 71 AGAHLRLGIGLLAVQASTLSGWLPEEKA 98
+G H LG+ + V+++TL WLPE+ A
Sbjct: 516 SGIHRSLGVHISKVRSATLDTWLPEQVA 599
>TC86175 similar to GP|23617209|dbj|BAC20880. contains ESTs C28562(C61610)
C25917(C11090)~similar to ARF GAP-like zinc
finger-containing protein, partial (47%)
Length = 1922
Score = 30.0 bits (66), Expect = 2.1
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +2
Query: 71 AGAHLRLGIGLLAVQASTLSGWLPEEKA 98
+G H LG+ + V+++TL WLPE+ A
Sbjct: 302 SGIHRSLGVHISKVRSATLDTWLPEQVA 385
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 30.0 bits (66), Expect = 2.1
Identities = 21/95 (22%), Positives = 44/95 (46%), Gaps = 2/95 (2%)
Frame = +2
Query: 297 TSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRL 356
+S HP ++GQ+E+ N+ LR + W++ + Y T+ + + +P++
Sbjct: 83 SSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFPWAEYWYNTSYNISAAMTPFKA 262
Query: 357 TFGTEAVIPAEIREPSARTAGF--DLNQNDQLLGE 389
+G + + + S TA L Q ++LL +
Sbjct: 263 LYGRDLSMLIRSKGSSKDTADLQSQLAQREELLSQ 367
>TC78001 similar to GP|21618169|gb|AAM67219.1 ARF GAP-like zinc
finger-containing protein ZIGA3 {Arabidopsis thaliana},
partial (46%)
Length = 1265
Score = 30.0 bits (66), Expect = 2.1
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +1
Query: 71 AGAHLRLGIGLLAVQASTLSGWLPEEKA 98
+G H LG+ + V+++TL WLPE+ A
Sbjct: 385 SGIHRSLGVHISKVRSATLDTWLPEQVA 468
>BG644699 similar to PIR|T07863|T078 probable polyprotein - pineapple
retrotransposon dea1 (fragment), partial (5%)
Length = 231
Score = 29.6 bits (65), Expect = 2.7
Identities = 15/51 (29%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Frame = +2
Query: 440 RSTERGKLAAKWEGPYRVVEATGTGAYKLETLAG-KEIPRTWNATNLRRYY 489
R +RGKL+ ++ GP+ V++ G AY+L G + ++ + +RY+
Sbjct: 47 RFGKRGKLSLRYIGPFEVIKRIGEVAYELALPPGLSGVHPVFHVSMFKRYH 199
>TC84257 weakly similar to PIR|G83180|G83180 probable FMN oxidoreductase
PA3723 [imported] - Pseudomonas aeruginosa (strain
PAO1), partial (31%)
Length = 777
Score = 29.3 bits (64), Expect = 3.6
Identities = 26/83 (31%), Positives = 34/83 (40%), Gaps = 7/83 (8%)
Frame = +3
Query: 2 RDAQEEIHMLRARLRQLEEQETQSRPCPRFPRKRRRTGARSHPLTTRKHAAGHPRPSGWP 61
R ++ H+ + + L Q + R PR P R AR P+ H GH R WP
Sbjct: 255 RPHHQQPHLGQPHVPVLRRQWPRDRLPPRPPGPVRP--ARRRPV----HGRGHRRRGSWP 416
Query: 62 AIHRGAHPGAG-------AHLRL 77
+ RG AG AH RL
Sbjct: 417 HLPRGCRFVAGLADCAAEAHRRL 485
>BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antigen. [strain
Kaplan PRV] {Pseudorabies virus}, partial (2%)
Length = 1022
Score = 28.5 bits (62), Expect = 6.1
Identities = 18/53 (33%), Positives = 22/53 (40%), Gaps = 3/53 (5%)
Frame = +3
Query: 27 PCPR---FPRKRRRTGARSHPLTTRKHAAGHPRPSGWPAIHRGAHPGAGAHLR 76
P PR PR+ + S P ++ GHP P G P H P AG R
Sbjct: 207 PRPRGGGLPRQPTKKQTPSMPAPKKRRTRGHPPPPGKPH-HAAPPPNAGRERR 362
>TC89138 homologue to GP|15215736|gb|AAK91413.1 AT5g19690/T29J13_110
{Arabidopsis thaliana}, partial (31%)
Length = 909
Score = 28.1 bits (61), Expect = 8.0
Identities = 14/35 (40%), Positives = 22/35 (62%)
Frame = +2
Query: 200 SLVSPWPFHQWGMDLLGPFDTAPG*LKYLVVAVDY 234
+LVSPWP ++W D L FD ++LVV +++
Sbjct: 353 NLVSPWPCNRW--DCLSRFDLDS---RHLVVGIEF 442
>TC76911 similar to GP|13543234|gb|AAH05782.1 Unknown (protein for
MGC:12025) {Mus musculus}, partial (57%)
Length = 1065
Score = 28.1 bits (61), Expect = 8.0
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = +2
Query: 53 GHPRPSGWPAIHRGAHPGAGAHLRLGIGLLAVQASTLSGW 92
G+P P+G+P H G H G+ H +G L A+ + +
Sbjct: 341 GYP-PAGYPGAHAGQHAGSHGHGGMGAMLAGGAAAAAAAY 457
>TC88039 homologue to GP|15623873|dbj|BAB67931. hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (12%)
Length = 903
Score = 28.1 bits (61), Expect = 8.0
Identities = 14/36 (38%), Positives = 20/36 (54%)
Frame = +3
Query: 131 AKDRAAYVLAEIHEGSCGHHLGGRSLARKVLRAGYY 166
AK+ Y +A + + CGH+L G KV RA +Y
Sbjct: 618 AKNTRPYAIAWVLKKLCGHNLDGDIKDPKVRRATFY 725
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.137 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,840,944
Number of Sequences: 36976
Number of extensions: 205516
Number of successful extensions: 1104
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1087
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1100
length of query: 489
length of database: 9,014,727
effective HSP length: 100
effective length of query: 389
effective length of database: 5,317,127
effective search space: 2068362403
effective search space used: 2068362403
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0083.15


