

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
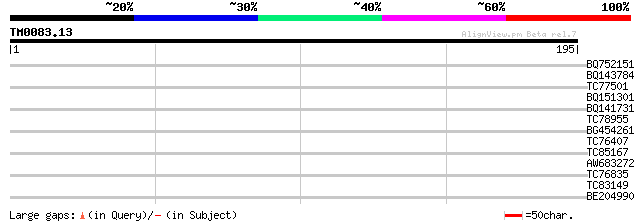
Query= TM0083.13
(195 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BQ752151 similar to GP|1945155|emb MN1 {Homo sapiens}, partial (1%) 31 0.35
BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antig... 30 0.78
TC77501 similar to GP|13560781|gb|AAK30204.1 endoxyloglucan tran... 30 0.78
BQ151301 similar to PIR|S34666|S346 glycine-rich protein - commo... 29 1.3
BQ141731 homologue to PIR|T05444|T054 hypothetical protein F7K2.... 28 3.0
TC78955 weakly similar to GP|22137114|gb|AAM91402.1 At1g24050/T2... 27 3.9
BG454261 weakly similar to GP|22094357|gb| putative receptor-lik... 27 3.9
TC76407 weakly similar to GP|20453227|gb|AAM19852.1 AT3g06540/F5... 27 3.9
TC85167 homologue to PIR|S45788|S45788 probable membrane protein... 27 5.0
AW683272 27 6.6
TC76835 similar to GP|21537194|gb|AAM61535.1 unknown {Arabidopsi... 27 6.6
TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk p... 27 6.6
BE204990 26 8.6
>BQ752151 similar to GP|1945155|emb MN1 {Homo sapiens}, partial (1%)
Length = 650
Score = 30.8 bits (68), Expect = 0.35
Identities = 18/50 (36%), Positives = 22/50 (44%), Gaps = 7/50 (14%)
Frame = +3
Query: 26 RHTPPKARGRGLAKQRDPPGNDTYHPKSQ-------ASHPPQDKMTSPEA 68
R PP +GR A P G+ HP+SQ SHPP K +A
Sbjct: 222 RREPPPPQGREAAWPPGPQGHAMGHPQSQQQPSAWPPSHPPPPKQPQQQA 371
>BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antigen. [strain
Kaplan PRV] {Pseudorabies virus}, partial (2%)
Length = 1022
Score = 29.6 bits (65), Expect = 0.78
Identities = 20/72 (27%), Positives = 28/72 (38%), Gaps = 2/72 (2%)
Frame = +2
Query: 26 RHTPPKARGRGLAKQRDPPGNDTYHPKSQASHP--PQDKMTSPEAYLKALRAEALDCNLY 83
R PP A GRG K + N + + +HP P +P A R +
Sbjct: 194 RSPPPPAPGRGAPKTANQKANPLHASPQKKTHPGTPATPRETPSRRTPAQRGQRTAGPRK 373
Query: 84 YSYQTQVTAPGR 95
+ Q TAPG+
Sbjct: 374 KPQEEQKTAPGK 409
>TC77501 similar to GP|13560781|gb|AAK30204.1 endoxyloglucan transferase
{Daucus carota}, partial (89%)
Length = 1524
Score = 29.6 bits (65), Expect = 0.78
Identities = 10/24 (41%), Positives = 17/24 (70%)
Frame = +2
Query: 10 RPIRPCHVRKEAVTAKRHTPPKAR 33
+ IRPCH+R+ ++ + TPP+ R
Sbjct: 1112 KEIRPCHIRQRQTSSWKTTPPQQR 1183
>BQ151301 similar to PIR|S34666|S346 glycine-rich protein - common tobacco,
partial (22%)
Length = 530
Score = 28.9 bits (63), Expect = 1.3
Identities = 18/51 (35%), Positives = 22/51 (42%), Gaps = 2/51 (3%)
Frame = +3
Query: 11 PIRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTY--HPKSQASHPP 59
P P + +K+ T PP RGR L R PP + HP HPP
Sbjct: 81 PPPPKNKKKKNHTTPPPPPPMRRGR*LCPPRPPPPPLPHGTHPPPPPHHPP 233
>BQ141731 homologue to PIR|T05444|T054 hypothetical protein F7K2.80 -
Arabidopsis thaliana, partial (4%)
Length = 1306
Score = 27.7 bits (60), Expect = 3.0
Identities = 18/56 (32%), Positives = 24/56 (42%)
Frame = +3
Query: 10 RPIRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTS 65
RP R RK+ T +RH R R + PP D + K +PPQ T+
Sbjct: 900 RPKREDRQRKQRHTRQRHADTTKRQRKPDNRCAPPAGDPHRTK----NPPQGPTTT 1055
>TC78955 weakly similar to GP|22137114|gb|AAM91402.1 At1g24050/T23E23_11
{Arabidopsis thaliana}, partial (57%)
Length = 1464
Score = 27.3 bits (59), Expect = 3.9
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = -2
Query: 106 EDLDPQLKAGLLTFLELTEEMWELHRRRL 134
+D+D + K + T ++TEE+WE RL
Sbjct: 371 DDIDRECKESVETLKKITEELWEEKEGRL 285
>BG454261 weakly similar to GP|22094357|gb| putative receptor-like protein
kinase {Oryza sativa (japonica cultivar-group)}, partial
(16%)
Length = 650
Score = 27.3 bits (59), Expect = 3.9
Identities = 14/46 (30%), Positives = 25/46 (53%)
Frame = -3
Query: 116 LLTFLELTEEMWELHRRRLEVNMLIWGLEVSATEHKGEETGTMGEI 161
++ FL T + W+LH R + + ++ L+ ++ GEE M EI
Sbjct: 480 IIWFLNFT*QSWKLHERGMHLELVDKALD--PNDYDGEEVKKMIEI 349
>TC76407 weakly similar to GP|20453227|gb|AAM19852.1 AT3g06540/F5E6_13
{Arabidopsis thaliana}, partial (62%)
Length = 2233
Score = 27.3 bits (59), Expect = 3.9
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +2
Query: 9 HRPIRPCHVRKEAVTAKRHTPPKAR 33
HRP R H+R+ K H PP+++
Sbjct: 146 HRPFRIHHLRRRIRRRKNHPPPRSQ 220
>TC85167 homologue to PIR|S45788|S45788 probable membrane protein YBL053w -
yeast (Saccharomyces cerevisiae), partial (9%)
Length = 988
Score = 26.9 bits (58), Expect = 5.0
Identities = 20/66 (30%), Positives = 26/66 (39%)
Frame = -1
Query: 28 TPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSPEAYLKALRAEALDCNLYYSYQ 87
T P A G+ + P P H P TSP A AL A A+ ++ +S
Sbjct: 706 TSPTAFFAGVQNCQT*PYGSVTFPTRPPPHAPSHSATSPSATAPALTAFAIAASM-FSDS 530
Query: 88 TQVTAP 93
Q T P
Sbjct: 529 RQKTTP 512
>AW683272
Length = 367
Score = 26.6 bits (57), Expect = 6.6
Identities = 13/43 (30%), Positives = 22/43 (50%)
Frame = +3
Query: 62 KMTSPEAYLKALRAEALDCNLYYSYQTQVTAPGRIRSLVSGLL 104
+ +S + YL+ + A+DC+ YY + R SL+ LL
Sbjct: 6 RSSSLQVYLQYAESPAMDCDYYYYHYKWQCRDKR*TSLIGNLL 134
>TC76835 similar to GP|21537194|gb|AAM61535.1 unknown {Arabidopsis thaliana},
partial (40%)
Length = 1427
Score = 26.6 bits (57), Expect = 6.6
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = +1
Query: 40 QRDPPGNDTYHPKSQASHPPQDKMTSP 66
QRD ++++HP+ A PP+ M SP
Sbjct: 1132 QRDLVYHNSFHPRQSAFQPPR*PMFSP 1212
>TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk protein
{Argiope trifasciata}, partial (3%)
Length = 1177
Score = 26.6 bits (57), Expect = 6.6
Identities = 19/60 (31%), Positives = 23/60 (37%), Gaps = 4/60 (6%)
Frame = +3
Query: 11 PIRPCHVRKEAV-TAKRHTP---PKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSP 66
P RP A T+ R TP P R R PP HPK PP+ + + P
Sbjct: 66 PTRPSSPTPSAPRTSSRSTPSTSPSRRPRSKTPLPPPPPPLPQHPKPPQKSPPRMRRSKP 245
>BE204990
Length = 629
Score = 26.2 bits (56), Expect = 8.6
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +2
Query: 65 SPEAYLKALRAEALDCNLYYSYQTQ 89
SP +Y A +DC+LY+ +Q Q
Sbjct: 416 SPTSYKLAFLCSKVDCSLYFLHQNQ 490
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,930,968
Number of Sequences: 36976
Number of extensions: 81131
Number of successful extensions: 504
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 489
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 501
length of query: 195
length of database: 9,014,727
effective HSP length: 91
effective length of query: 104
effective length of database: 5,649,911
effective search space: 587590744
effective search space used: 587590744
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0083.13


