

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
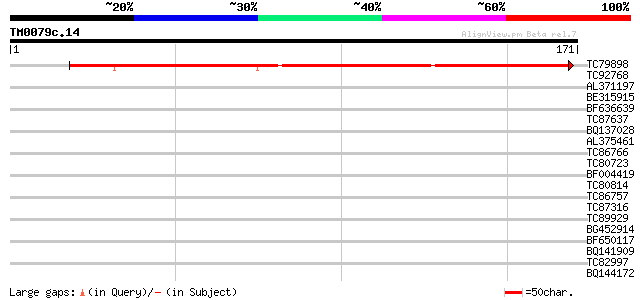
Query= TM0079c.14
(171 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC79898 similar to GP|15450862|gb|AAK96702.1 Unknown protein {Ar... 172 6e-44
TC92768 weakly similar to PIR|C96593|C96593 unknown protein 999... 31 0.21
AL371197 homologue to GP|6273331|gb| glycine-rich RNA binding pr... 31 0.28
BE315915 similar to GP|21618095|gb| unknown {Arabidopsis thalian... 30 0.62
BF636639 similar to GP|20198018|gb putative eukaryotic translati... 29 0.81
TC87637 similar to PIR|T02462|T02462 probable AT-hook DNA-bindin... 29 1.1
BQ137028 similar to GP|4914334|gb|A F14N23.20 {Arabidopsis thali... 29 1.1
AL375461 weakly similar to GP|7300610|gb|A CG17203 gene product ... 26 1.1
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 28 1.4
TC80723 similar to PIR|T48524|T48524 lysophospholipase-like prot... 28 1.4
BF004419 similar to GP|20197706|gb| unknown protein {Arabidopsis... 28 1.4
TC80814 similar to GP|3413719|gb|AAC31242.1| unknown protein {Ar... 28 1.8
TC86757 auxin influx carrier protein [Medicago truncatula] 28 1.8
TC87316 weakly similar to PIR|G84675|G84675 probable cytochrome ... 28 2.4
TC89929 similar to GP|10187185|emb|CAC09066. cDNA~Lemon acyl tra... 28 2.4
BG452914 similar to GP|13374061|emb ZF-HD homeobox protein {Flav... 23 3.0
BF650117 similar to PIR|C86187|C861 YUP8H12.12 [imported] - Arab... 27 3.1
BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox ... 27 4.0
TC82997 homologue to GP|7293564|gb|AAF48937.1| CG7884 gene produ... 27 4.0
BQ144172 weakly similar to PIR|D86416|D864 probable beta-1 3 glu... 27 4.0
>TC79898 similar to GP|15450862|gb|AAK96702.1 Unknown protein {Arabidopsis
thaliana}, partial (59%)
Length = 1632
Score = 172 bits (436), Expect = 6e-44
Identities = 83/155 (53%), Positives = 115/155 (73%), Gaps = 3/155 (1%)
Frame = +3
Query: 19 GGNIYWGRKHAT--DFRGIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCYAHYLSA-FR 75
GG YWGR+ GIVV+FAW+S + L +V+LYSSLGWNSL+C++ +L+ F
Sbjct: 90 GGRYYWGRRKVDCEKANGIVVVFAWMSSEEKHLMRYVDLYSSLGWNSLICHSQFLNMFFP 269
Query: 76 DESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRCETPHCLHNYH 135
D++T+P A +++EL+E LK + CP+VFA+FS G+KAC+ KVLQ+I G CET H + +Y
Sbjct: 270 DKATIP-AVDILNELVEVLKIRRCPIVFASFSGGAKACMLKVLQIIGGECET-HNMDDYQ 443
Query: 136 LLRNCVSGHIYDSGPLDVTSDFGFRFALHPSMAKV 170
L+R+C+SG+IYDS P+D TSD G RF LHPS+ KV
Sbjct: 444 LVRDCISGYIYDSSPVDFTSDLGVRFVLHPSVLKV 548
>TC92768 weakly similar to PIR|C96593|C96593 unknown protein 99945-98618
[imported] - Arabidopsis thaliana, partial (27%)
Length = 668
Score = 31.2 bits (69), Expect = 0.21
Identities = 22/68 (32%), Positives = 31/68 (45%), Gaps = 12/68 (17%)
Frame = -2
Query: 1 MRGAESEGGGGGGG--CAVRGG-----NIYWGRK-HATDFRGIVVIFAW----VSVPQTL 48
M+G+ GGGGGG + GG + W R + DF G V+I W +S T
Sbjct: 256 MKGSLGSGGGGGGAV*VPIDGGEENSFDFCWPRNCFSCDFFGFVII*FWRRSMISTTTTF 77
Query: 49 LQDFVNLY 56
F++ Y
Sbjct: 76 THSFLHFY 53
>AL371197 homologue to GP|6273331|gb| glycine-rich RNA binding protein
{Medicago sativa}, complete
Length = 437
Score = 30.8 bits (68), Expect = 0.28
Identities = 12/21 (57%), Positives = 13/21 (61%)
Frame = +1
Query: 2 RGAESEGGGGGGGCAVRGGNI 22
RG GGGG GGC RGG +
Sbjct: 295 RGIGGGGGGGPGGCGYRGGGV 357
>BE315915 similar to GP|21618095|gb| unknown {Arabidopsis thaliana}, partial
(10%)
Length = 492
Score = 29.6 bits (65), Expect = 0.62
Identities = 21/62 (33%), Positives = 29/62 (45%), Gaps = 5/62 (8%)
Frame = +3
Query: 52 FVNLYSSLGWNSLVCYAH-----YLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAF 106
F+N++ S WN+L+ Y H YL + V L FC D EL +F +F
Sbjct: 78 FLNIFISQSWNTLI-YLH*SKILYLLQAAQQQEVTLTFC--DNYACELSLSLPSNLFTSF 248
Query: 107 SA 108
SA
Sbjct: 249 SA 254
>BF636639 similar to GP|20198018|gb putative eukaryotic translation
initiation factor 2 alpha subunit eIF2 {Arabidopsis
thaliana}, partial (50%)
Length = 591
Score = 29.3 bits (64), Expect = 0.81
Identities = 18/52 (34%), Positives = 25/52 (47%)
Frame = +2
Query: 44 VPQTLLQDFVNLYSSLGWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELK 95
+P+TL D +LY +GW Y H AF+ P A V+ L E+K
Sbjct: 59 LPETLNLDLEDLYVHIGWPLYRKYGHAFEAFKRVLADPDA--VLSALTREIK 208
>TC87637 similar to PIR|T02462|T02462 probable AT-hook DNA-binding protein
[imported] - Arabidopsis thaliana, partial (53%)
Length = 1659
Score = 28.9 bits (63), Expect = 1.1
Identities = 16/41 (39%), Positives = 19/41 (46%)
Frame = -1
Query: 120 LIDGRCETPHCLHNYHLLRNCVSGHIYDSGPLDVTSDFGFR 160
L+ G TPH NYHL R C + D+ D T G R
Sbjct: 1230 LVLGFWSTPHKAPNYHLNRTCSNEQSTDASTNDTTVRTGKR 1108
>BQ137028 similar to GP|4914334|gb|A F14N23.20 {Arabidopsis thaliana},
partial (7%)
Length = 687
Score = 28.9 bits (63), Expect = 1.1
Identities = 12/22 (54%), Positives = 12/22 (54%)
Frame = -2
Query: 4 AESEGGGGGGGCAVRGGNIYWG 25
A GGGGGGGC G WG
Sbjct: 494 APGGGGGGGGGCGCGGEGGDWG 429
>AL375461 weakly similar to GP|7300610|gb|A CG17203 gene product {Drosophila
melanogaster}, partial (15%)
Length = 524
Score = 25.8 bits (55), Expect(2) = 1.1
Identities = 13/30 (43%), Positives = 15/30 (49%)
Frame = -3
Query: 9 GGGGGGCAVRGGNIYWGRKHATDFRGIVVI 38
GGGGGG G R D RG+VV+
Sbjct: 213 GGGGGGGDDAAGVFRVPRPRREDVRGVVVV 124
Score = 21.6 bits (44), Expect(2) = 1.1
Identities = 8/11 (72%), Positives = 9/11 (81%)
Frame = -3
Query: 4 AESEGGGGGGG 14
A + GGGGGGG
Sbjct: 366 AAAVGGGGGGG 334
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 28.5 bits (62), Expect = 1.4
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = -1
Query: 3 GAESEGGGGGGGCAVRGG 20
G ++GGGGGGG + +GG
Sbjct: 336 GGSTQGGGGGGGGSTQGG 283
>TC80723 similar to PIR|T48524|T48524 lysophospholipase-like protein -
Arabidopsis thaliana, partial (27%)
Length = 669
Score = 28.5 bits (62), Expect = 1.4
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = -2
Query: 6 SEGGGGGGGCAVRGGNIYWGRK 27
+ GGGGGGGC GG W +K
Sbjct: 113 ANGGGGGGGCG--GGKWRWRKK 54
>BF004419 similar to GP|20197706|gb| unknown protein {Arabidopsis thaliana},
partial (20%)
Length = 484
Score = 28.5 bits (62), Expect = 1.4
Identities = 11/13 (84%), Positives = 13/13 (99%)
Frame = +3
Query: 159 FRFALHPSMAKVP 171
FRF+LHPS+AKVP
Sbjct: 3 FRFSLHPSIAKVP 41
>TC80814 similar to GP|3413719|gb|AAC31242.1| unknown protein {Arabidopsis
thaliana}, partial (11%)
Length = 1436
Score = 28.1 bits (61), Expect = 1.8
Identities = 15/35 (42%), Positives = 19/35 (53%), Gaps = 3/35 (8%)
Frame = -1
Query: 36 VVIFAWVSVPQTLLQDFVNLYS---SLGWNSLVCY 67
VVIF +S + ++S SLGW SLVCY
Sbjct: 1286 VVIFV*ISTMYRTASPSIMIFSAIDSLGWTSLVCY 1182
>TC86757 auxin influx carrier protein [Medicago truncatula]
Length = 1590
Score = 28.1 bits (61), Expect = 1.8
Identities = 14/39 (35%), Positives = 21/39 (52%), Gaps = 2/39 (5%)
Frame = -1
Query: 128 PHCLHNYHLLRNCVSG--HIYDSGPLDVTSDFGFRFALH 164
PHC+HN+H G ++ SG +V + FG+ LH
Sbjct: 858 PHCMHNFHCNSMSTKGIENVGSSG--EVKNQFGWTRVLH 748
>TC87316 weakly similar to PIR|G84675|G84675 probable cytochrome P450
[imported] - Arabidopsis thaliana, partial (77%)
Length = 1678
Score = 27.7 bits (60), Expect = 2.4
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = +2
Query: 99 CPVVFAAFSAGSKACLYKVLQLID 122
CP + F AGS+ CL K L +++
Sbjct: 1379 CPFKYPVFQAGSRVCLGKELAIVE 1450
>TC89929 similar to GP|10187185|emb|CAC09066. cDNA~Lemon acyl transferase
{Citrus limon}, partial (22%)
Length = 1361
Score = 27.7 bits (60), Expect = 2.4
Identities = 24/87 (27%), Positives = 37/87 (41%), Gaps = 17/87 (19%)
Frame = +1
Query: 89 ELIEELKTKSCPVVFAAF---SAGSKACLYKVLQLIDGRCET--------------PHCL 131
EL++ + S P ++F A S CL K + ++ E P
Sbjct: 760 ELLDTNELTSKPPTLSSFVLTCAYSLVCLAKAIHGVEKEKEKFGFAFTVDCRARLEPPLP 939
Query: 132 HNYHLLRNCVSGHIYDSGPLDVTSDFG 158
+NY NCV GH+ D+ PLD ++ G
Sbjct: 940 NNY--FGNCVWGHLVDTKPLDYINEDG 1014
>BG452914 similar to GP|13374061|emb ZF-HD homeobox protein {Flaveria
bidentis}, partial (29%)
Length = 454
Score = 23.5 bits (49), Expect(2) = 3.0
Identities = 9/14 (64%), Positives = 9/14 (64%)
Frame = +2
Query: 1 MRGAESEGGGGGGG 14
M S GGGGGGG
Sbjct: 224 MSNPSSSGGGGGGG 265
Score = 22.3 bits (46), Expect(2) = 3.0
Identities = 11/33 (33%), Positives = 16/33 (48%)
Frame = +2
Query: 8 GGGGGGGCAVRGGNIYWGRKHATDFRGIVVIFA 40
GGGGGGG G + K + + ++ FA
Sbjct: 251 GGGGGGGSGGSGSRKRFRTKFTQEQKEKLLAFA 349
>BF650117 similar to PIR|C86187|C861 YUP8H12.12 [imported] - Arabidopsis
thaliana, partial (25%)
Length = 604
Score = 27.3 bits (59), Expect = 3.1
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = -1
Query: 7 EGGGGGGGCAVRGGNIYWGRKH 28
+GGGGGG GG I+W R H
Sbjct: 157 DGGGGGG-----GGGIWWMRVH 107
>BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox carteri f.
nagariensis}, partial (13%)
Length = 1223
Score = 26.9 bits (58), Expect = 4.0
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = +1
Query: 2 RGAESEGGGGGGGCAVRGGNIYWGRK 27
+G + E GGGGGG GG + G++
Sbjct: 166 KG*KGEAGGGGGGGGGSGGGVGEGKR 243
>TC82997 homologue to GP|7293564|gb|AAF48937.1| CG7884 gene product
{Drosophila melanogaster}, partial (2%)
Length = 792
Score = 26.9 bits (58), Expect = 4.0
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = -2
Query: 7 EGGGGGGGCAVRGGNIYWGRKHATDFRG 34
EGGGGG G + R G I G+++ RG
Sbjct: 371 EGGGGGIGPS*RNGGIRLGKRNKRGERG 288
>BQ144172 weakly similar to PIR|D86416|D864 probable beta-1 3 glucanase
26636-27432 [imported] - Arabidopsis thaliana, partial
(15%)
Length = 1333
Score = 26.9 bits (58), Expect = 4.0
Identities = 15/29 (51%), Positives = 15/29 (51%), Gaps = 2/29 (6%)
Frame = +2
Query: 8 GGGGGGGC--AVRGGNIYWGRKHATDFRG 34
GGGGGGGC RGG R HA G
Sbjct: 14 GGGGGGGCGRGTRGGR----RGHAPSCEG 88
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.141 0.461
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,519,697
Number of Sequences: 36976
Number of extensions: 103612
Number of successful extensions: 2709
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 1374
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2267
length of query: 171
length of database: 9,014,727
effective HSP length: 89
effective length of query: 82
effective length of database: 5,723,863
effective search space: 469356766
effective search space used: 469356766
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0079c.14


