

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079a.1
(139 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
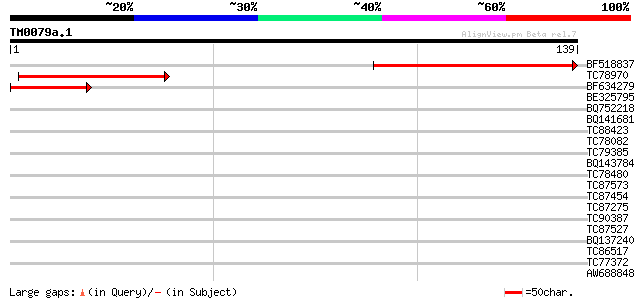
Score E
Sequences producing significant alignments: (bits) Value
BF518837 similar to GP|15010772|gb| At1g49590/F14J22_9 {Arabidop... 84 1e-17
TC78970 similar to GP|21703117|gb|AAM74500.1 AT5g09390/T5E8_190 ... 48 1e-06
BF634279 43 5e-05
BE325795 similar to GP|22136848|gb| unknown protein {Arabidopsis... 35 0.013
BQ752218 similar to GP|19170914|emb hypothetical protein {Enceph... 31 0.14
BQ141681 weakly similar to GP|15145793|gb| basic proline-rich pr... 29 0.53
TC88423 similar to GP|9758777|dbj|BAB09075.1 contains similarity... 28 0.91
TC78082 similar to GP|21618292|gb|AAM67342.1 unknown {Arabidopsi... 28 0.91
TC79385 similar to GP|9279590|dbj|BAB01048.1 gene_id:MPE11.2~unk... 28 0.91
BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antig... 28 1.2
TC78480 homologue to PIR|S42368|S42368 guanine nucleotide releas... 28 1.2
TC87573 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 27 2.0
TC87454 similar to SP|P49730|RIR2_TOBAC Ribonucleoside-diphospha... 27 2.0
TC87275 homologue to GP|16974682|gb|AAL32441.1 Fe-superoxide dis... 27 2.7
TC90387 similar to PIR|H86378|H86378 protein F21J9.16 [imported]... 27 2.7
TC87527 similar to GP|21689681|gb|AAM67462.1 unknown protein {Ar... 27 3.5
BQ137240 similar to PIR|F75311|F753 ABC transporter ATP-binding... 27 3.5
TC86517 homologue to PIR|T09640|T09640 protein phosphatase 2C - ... 27 3.5
TC77372 similar to GP|8745402|gb|AAF78903.1| proline-rich protei... 26 4.5
AW688848 homologue to PIR|T46027|T4 hypothetical protein T10K17.... 26 4.5
>BF518837 similar to GP|15010772|gb| At1g49590/F14J22_9 {Arabidopsis
thaliana}, partial (39%)
Length = 324
Score = 84.3 bits (207), Expect = 1e-17
Identities = 40/50 (80%), Positives = 47/50 (94%)
Frame = +2
Query: 90 SSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLLGLYSKP 139
SSL++GKRKRP EKPKV+S+EEK ALKAREAARKRV++REK LLGLY+KP
Sbjct: 5 SSLAVGKRKRPNEKPKVISQEEKEALKAREAARKRVEQREKPLLGLYNKP 154
>TC78970 similar to GP|21703117|gb|AAM74500.1 AT5g09390/T5E8_190 {Arabidopsis
thaliana}, partial (45%)
Length = 1600
Score = 48.1 bits (113), Expect = 1e-06
Identities = 19/37 (51%), Positives = 25/37 (67%)
Frame = +1
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEE 39
++ SSGYYY + G+YYDP +G Y S A GKW + E
Sbjct: 967 YDESSGYYYSSSLGYYYDPNTGLYCSAASGKWYSFNE 1077
>BF634279
Length = 464
Score = 42.7 bits (99), Expect = 5e-05
Identities = 16/20 (80%), Positives = 18/20 (90%)
Frame = +1
Query: 1 WDFESSSGYYYHKTNGFYYD 20
W+F+SSSGYYYHKTNG YD
Sbjct: 400 WEFDSSSGYYYHKTNGSCYD 459
>BE325795 similar to GP|22136848|gb| unknown protein {Arabidopsis thaliana},
partial (6%)
Length = 676
Score = 34.7 bits (78), Expect = 0.013
Identities = 23/73 (31%), Positives = 36/73 (48%)
Frame = +2
Query: 37 QEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGK 96
QEE Y P KK+SST K KP+N +S N T+ +K+ ++LS
Sbjct: 353 QEEEYEQP----LKKKSSSTKNQTKLIKPNNN-------LSSKNQTKTIKSNTNNLSSKN 499
Query: 97 RKRPGEKPKVVSE 109
+ +P + K+ S+
Sbjct: 500 QTKPSKDSKLTSK 538
>BQ752218 similar to GP|19170914|emb hypothetical protein {Encephalitozoon
cuniculi}, partial (5%)
Length = 637
Score = 31.2 bits (69), Expect = 0.14
Identities = 18/37 (48%), Positives = 23/37 (61%), Gaps = 2/37 (5%)
Frame = -2
Query: 96 KRKRPGEKPKVVSEEE--KSALKAREAARKRVQEREK 130
K K+ EK K + EE K A KAREAA K+ +E +K
Sbjct: 606 KAKKEAEKKKKKAREEAAKKAKKAREAAYKKAKEEKK 496
>BQ141681 weakly similar to GP|15145793|gb| basic proline-rich protein {Sus
scrofa}, partial (9%)
Length = 516
Score = 29.3 bits (64), Expect = 0.53
Identities = 16/59 (27%), Positives = 26/59 (43%)
Frame = +3
Query: 66 HNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKR 124
+ GP + T + NP +A ++RP EK K + ++ A R+ RKR
Sbjct: 132 NQGPTQPKARTQTQNPRAQQRAQTKGQKQPTKERPAEKEKNRQKAKQRARATRKTGRKR 308
>TC88423 similar to GP|9758777|dbj|BAB09075.1 contains similarity to unknown
protein~dbj|BAA90625.1~gene_id:MNJ7.8 {Arabidopsis
thaliana}, partial (15%)
Length = 1640
Score = 28.5 bits (62), Expect = 0.91
Identities = 28/103 (27%), Positives = 42/103 (40%)
Frame = +1
Query: 22 KSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNP 81
K+ FYY + + +WV +E A A+ +T+ G+ +N L + T L P
Sbjct: 712 KNKFYYDEKLKRWV-EEGAEVPAEEAALPPPPPTTAAFQNGSADYN--LKSALKTEGLTP 882
Query: 82 TRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKR 124
SS L PG P S + SA ++R R R
Sbjct: 883 NEFSSTRTSSPELS----PGMPPIPPSSNQFSA-RSRLGVRSR 996
>TC78082 similar to GP|21618292|gb|AAM67342.1 unknown {Arabidopsis
thaliana}, partial (60%)
Length = 1626
Score = 28.5 bits (62), Expect = 0.91
Identities = 18/54 (33%), Positives = 28/54 (51%)
Frame = +2
Query: 76 TTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQERE 129
T L+ T + AAP SL +GK R ++PK E KS ++ + R ++E
Sbjct: 689 TRELHTTDALPAAPVSLEVGKPPRRAKRPK----ESKSDSPNKKTPKSRKVKKE 838
>TC79385 similar to GP|9279590|dbj|BAB01048.1 gene_id:MPE11.2~unknown
protein {Arabidopsis thaliana}, partial (16%)
Length = 912
Score = 28.5 bits (62), Expect = 0.91
Identities = 18/48 (37%), Positives = 23/48 (47%)
Frame = -1
Query: 54 SSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPG 101
SS S N P P P + V +S NPT+ VK+ S S +PG
Sbjct: 636 SSAPPSLSDNNPVP*PNPSKPVKSSSNPTKPVKSTLSGSSEPGSFKPG 493
>BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antigen. [strain
Kaplan PRV] {Pseudorabies virus}, partial (2%)
Length = 1022
Score = 28.1 bits (61), Expect = 1.2
Identities = 19/83 (22%), Positives = 32/83 (37%), Gaps = 4/83 (4%)
Frame = +2
Query: 52 KASSTSQSNKGNKPHNGPL----PGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVV 107
+ + + + K N H P PG T P+R A + G RK+P E+ K
Sbjct: 221 RGAPKTANQKANPLHASPQKKTHPGTPATPRETPSRRTPAQRGQRTAGPRKKPQEEQKTA 400
Query: 108 SEEEKSALKAREAARKRVQEREK 130
+ + +K + + EK
Sbjct: 401 PGKNAAPPPP*HPPQKELDQLEK 469
Score = 26.6 bits (57), Expect = 3.5
Identities = 17/59 (28%), Positives = 22/59 (36%)
Frame = +1
Query: 71 PGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQERE 129
PG +T PTR P + G+ K K + SA K RKR + E
Sbjct: 310 PGNPITPHPRPTRAENGGPQEKATGRAKNRPRKKRSPPTTIASAPKRTGPTRKRKGDSE 486
>TC78480 homologue to PIR|S42368|S42368 guanine nucleotide releasing factor
homolog - Caenorhabditis elegans, partial (1%)
Length = 1033
Score = 28.1 bits (61), Expect = 1.2
Identities = 12/53 (22%), Positives = 25/53 (46%)
Frame = -3
Query: 64 KPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALK 116
+PH LPG + ++S+ PT + ++P + E ++ +AL+
Sbjct: 1025 QPHQHKLPGSLFSSSITPTTSDTSSPDGFGSSMQSSSSEDVNILDFTAAAALR 867
>TC87573 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, partial (50%)
Length = 836
Score = 27.3 bits (59), Expect = 2.0
Identities = 15/36 (41%), Positives = 22/36 (60%), Gaps = 2/36 (5%)
Frame = +1
Query: 80 NPTRNVKAAPSSLSLGKR--KRPGEKPKVVSEEEKS 113
NPT+ ++ P+S K+ K EKPKVV EE++
Sbjct: 37 NPTQPKESKPTSKDAPKKVAKSEPEKPKVVEAEEEA 144
>TC87454 similar to SP|P49730|RIR2_TOBAC Ribonucleoside-diphosphate
reductase small chain (EC 1.17.4.1) (Ribonucleotide
reductase), partial (96%)
Length = 1675
Score = 27.3 bits (59), Expect = 2.0
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 17/64 (26%)
Frame = +2
Query: 46 FASTTKKASSTSQSN--------KGNKPHNGPLPGRV---------VTTSLNPTRNVKAA 88
F T+ S+ SQSN K KPH+GPL + + + N T ++ ++
Sbjct: 110 FLPQTQTDSACSQSNTQKSGKCTKKPKPHSGPLKKSIFPPISSTGKTSPTANATSSLMSS 289
Query: 89 PSSL 92
PSSL
Sbjct: 290 PSSL 301
>TC87275 homologue to GP|16974682|gb|AAL32441.1 Fe-superoxide dismutase
precursor {Medicago sativa}, complete
Length = 1111
Score = 26.9 bits (58), Expect = 2.7
Identities = 17/51 (33%), Positives = 22/51 (42%), Gaps = 3/51 (5%)
Frame = -1
Query: 66 HNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPG---EKPKVVSEEEKS 113
H P P V SLN + N P SLS+ R P P + ++KS
Sbjct: 595 HAHPEPNCVEAASLNCSTNFSNEPKSLSISFRSSPDGFPPPPGFIDSQKKS 443
>TC90387 similar to PIR|H86378|H86378 protein F21J9.16 [imported] -
Arabidopsis thaliana, partial (53%)
Length = 1308
Score = 26.9 bits (58), Expect = 2.7
Identities = 17/38 (44%), Positives = 21/38 (54%)
Frame = -2
Query: 90 SSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQE 127
+S S G K+PG KPK V+E RE RKR+ E
Sbjct: 296 NSASSGSGKKPG*KPKRVTE--------REPRRKRLVE 207
>TC87527 similar to GP|21689681|gb|AAM67462.1 unknown protein {Arabidopsis
thaliana}, partial (96%)
Length = 1582
Score = 26.6 bits (57), Expect = 3.5
Identities = 16/49 (32%), Positives = 23/49 (46%), Gaps = 6/49 (12%)
Frame = +2
Query: 97 RKRPGEKPKVVSEEEKSALKAREAARKRVQER------EKSLLGLYSKP 139
R+ KP+ + E+ A K EAA +R + R EK L + KP
Sbjct: 812 REAESSKPRAAKKREEEADKKAEAAARRAEVRRLAELEEKELEKMIKKP 958
>BQ137240 similar to PIR|F75311|F753 ABC transporter ATP-binding protein -
Deinococcus radiodurans (strain R1), partial (4%)
Length = 1094
Score = 26.6 bits (57), Expect = 3.5
Identities = 32/126 (25%), Positives = 50/126 (39%), Gaps = 30/126 (23%)
Frame = +2
Query: 36 TQEEAYASPHFASTTKKASS---TSQSNKGNKPHNGPLPGRVVTTSLNPTRNVK----AA 88
TQ A A+ + ++ A+ TS S++ H+ NPTRN K A
Sbjct: 437 TQRAAQAASPTSPPSRPATQHIDTSHSSRTRSTHD-----------TNPTRNAKGRVAAT 583
Query: 89 PSSLSLGKRKRPGE-------------------KPKVVSEEEKSALKAREAARKR----V 125
S+GK ++ E K +E + +K+RE ++R +
Sbjct: 584 TDDESIGKEEQEQERYEQRTRV*EAEDEG*QTTKRSKRAERRRVNIKSRERTKERRIERI 763
Query: 126 QEREKS 131
QEREKS
Sbjct: 764 QEREKS 781
>TC86517 homologue to PIR|T09640|T09640 protein phosphatase 2C - alfalfa,
complete
Length = 1616
Score = 26.6 bits (57), Expect = 3.5
Identities = 30/125 (24%), Positives = 51/125 (40%)
Frame = +2
Query: 14 TNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGR 73
+N + P S + + + + +T ++ P ++TT +SS+S S + P R
Sbjct: 152 SNSPVFSPSSSLFCNKS--ETLTLSLSHLKPS-STTTTTSSSSSSSPSPCSSPSSPFCLR 322
Query: 74 VVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLL 133
+ L T N + + ++ KRKRP VSE A + V E E
Sbjct: 323 LPKLPLVFTSNKDSGSQNDAVLKRKRPTRLNIPVSEHAFCVPATPSAVARDVVEAEGDGY 502
Query: 134 GLYSK 138
+Y K
Sbjct: 503 SVYCK 517
>TC77372 similar to GP|8745402|gb|AAF78903.1| proline-rich protein {Glycine
max}, partial (91%)
Length = 831
Score = 26.2 bits (56), Expect = 4.5
Identities = 17/50 (34%), Positives = 25/50 (50%), Gaps = 2/50 (4%)
Frame = -3
Query: 48 STTKKASSTSQSNKG--NKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLG 95
S ++ S+ S+S K N H P G +TT NP R+ + +SLG
Sbjct: 310 SAVQRQSAASKSAKPLINAQHLLPEGGFPITTLTNPKRSAQTPSFKVSLG 161
>AW688848 homologue to PIR|T46027|T4 hypothetical protein T10K17.260 -
Arabidopsis thaliana, partial (10%)
Length = 661
Score = 26.2 bits (56), Expect = 4.5
Identities = 14/32 (43%), Positives = 20/32 (61%), Gaps = 1/32 (3%)
Frame = +1
Query: 102 EKPKVVSEEEKSALKAREAA-RKRVQEREKSL 132
++ K++ EEEK + E RKR +EREK L
Sbjct: 241 KQTKLLEEEEKEKREEEERKERKRAKEREKKL 336
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.306 0.125 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,166,143
Number of Sequences: 36976
Number of extensions: 52931
Number of successful extensions: 366
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 356
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 363
length of query: 139
length of database: 9,014,727
effective HSP length: 86
effective length of query: 53
effective length of database: 5,834,791
effective search space: 309243923
effective search space used: 309243923
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0079a.1


